
Chinese Journal OF Rice Science ›› 2019, Vol. 33 ›› Issue (6): 513-522.DOI: 10.16819/j.1001-7216.2019. 9090
• Orginal Article • Previous Articles Next Articles
Zhuanzhuan CHEN1,2, Xianfeng LI1, Min ZHONG1, Jiaqi GE1, Xiaolei FAN1,2, Changquan ZHANG1,2, Qiaoquan LIU1,2,*( )
)
Received:2019-08-07
Revised:2019-09-13
Online:2019-11-10
Published:2019-11-10
Contact:
Qiaoquan LIU
陈专专1,2, 李先锋1, 仲敏1, 葛家奇1, 范晓磊1,2, 张昌泉1,2, 刘巧泉1,2,*( )
)
通讯作者:
刘巧泉
基金资助:CLC Number:
Zhuanzhuan CHEN, Xianfeng LI, Min ZHONG, Jiaqi GE, Xiaolei FAN, Changquan ZHANG, Qiaoquan LIU. Grain Quality as Affected by Down-regulation of Expression of Different ALK Alleles in indica Rice (Oryza sativa L.)[J]. Chinese Journal OF Rice Science, 2019, 33(6): 513-522.
陈专专, 李先锋, 仲敏, 葛家奇, 范晓磊, 张昌泉, 刘巧泉. 籼稻背景下抑制不同ALK等位基因表达对稻米品质的影响[J]. 中国水稻科学, 2019, 33(6): 513-522.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2019. 9090
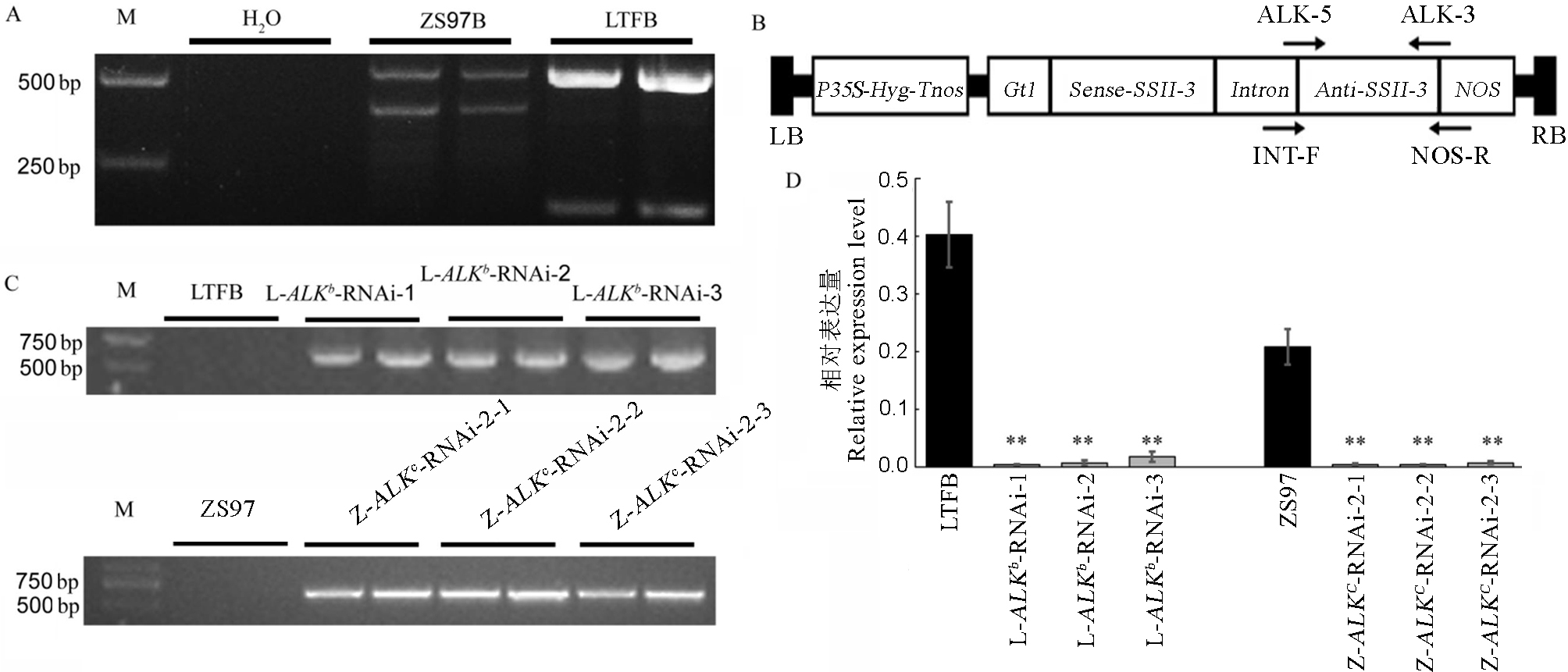
Fig. 1. ALK-RNAi construction and idendification of transgenic rice. A, ALK genotype in Longtefu B(LTFB) and Zhenshan 97B(ZS97B) on sites 4342 and 4343 based on primer 4342(GC/TT). B, The T-DNA structure of ALK-RNAi construct. P35S and Tnos represent the promoter and terminator region of the CaMV 35S gene respectively; NOS represents agrobacterium nopaline synthetase gene terminator; Hyg represents hygromycin resistance gene; LB and RB represent the left and right boundary sequences of the T-DNA region respectively; the anti-sense and the sense represent the reverse and forward structure of the target gene segment respectively; and the arrow in the figure indicates the direction of the designed primer. C, PCR analysis of ALK-RNAi transgenic rice plants. D, Transcription level expression of ALK Gene in RNAi Line. L-ALKb-RNAi-1, L-ALKb-RNAi-2 and L-ALKb-RNAi-3 are ALKb-RNAi transgenic homozygous lines under the background of Longtefu and Z-ALKc-RNAi-2-1, Z-ALKc-RNAi-2-2 and Z-ALKc-RNAi-2-3 are homozygous transgenic lines carrying both ALKb-RNAi interference structure and ALKc allele. Data were subjected to one-way analysis of variance (ANOVA) depending on the experiment, followed by a comparison of the means according to a Duncan’s multiple range test at P< 0.05 or P < 0.01. Double asterisks denote a highly significant difference between transgenic line and its wild type (P<0.01), single asterisk denotes a significant difference between transgenic line and it’s wild type (0.01≤ P<0.05). (n=3).
| 正向引物 Forward primer | 引物序列 Primer sequence(5′-3′) | 反向引物 Reverse primer | 引物序列 Primer sequence(5′-3′) |
|---|---|---|---|
| ALK-5 | GTCCATGGTCGACCTCAAGTAAGAACGGAG | ALK-3 | GTACTAGTCGCCTTTGGCTTC |
| 4342(GC/TT)-F1 | CAAGGAGAGCTGGAGGGGGC | 4342(GC/TT)-R1 | CATGCCGCGCACCTGGAAA |
| 4342(GC/TT)-F | TCGGCGGGCTGAGGGACAC | 4342(GC/TT)-R | TCGCATCAATGGACATAACAAACAC |
| INT-F | CCTCGTAATCAATTTTAGAT | NOS-R | GACCGCACAGGATTCAAT |
| ALK-RT-F | TGATCTGAACGAACCGGACG | ALK-RT-R | TGGGTAAAGCACCTGCAACA |
Table 1 Primers used in this study.
| 正向引物 Forward primer | 引物序列 Primer sequence(5′-3′) | 反向引物 Reverse primer | 引物序列 Primer sequence(5′-3′) |
|---|---|---|---|
| ALK-5 | GTCCATGGTCGACCTCAAGTAAGAACGGAG | ALK-3 | GTACTAGTCGCCTTTGGCTTC |
| 4342(GC/TT)-F1 | CAAGGAGAGCTGGAGGGGGC | 4342(GC/TT)-R1 | CATGCCGCGCACCTGGAAA |
| 4342(GC/TT)-F | TCGGCGGGCTGAGGGACAC | 4342(GC/TT)-R | TCGCATCAATGGACATAACAAACAC |
| INT-F | CCTCGTAATCAATTTTAGAT | NOS-R | GACCGCACAGGATTCAAT |
| ALK-RT-F | TGATCTGAACGAACCGGACG | ALK-RT-R | TGGGTAAAGCACCTGCAACA |
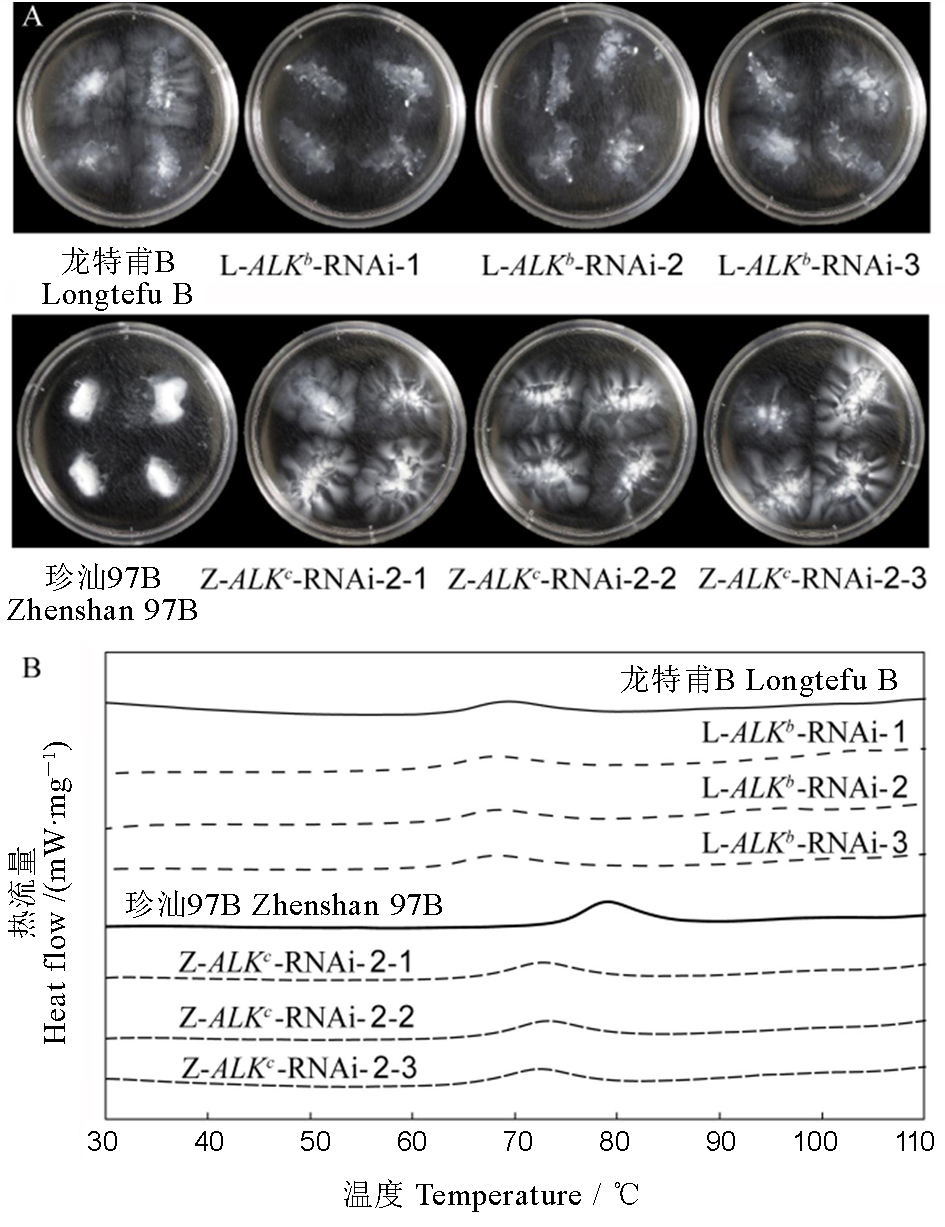
Fig 2. Gelatinization properties of ALK-RNAi transgenic rice. A, Alkali spreading value of milled rice in 1.7% KOH; B, DSC curve of rice flours of ALK-RNAi transgenic rice.
| 转基因系 | 起始温度 | 峰值温度 | 终止温度 | 热焓值 | 直链淀粉含量 | 胶稠度 | 粗蛋白含量 | |
|---|---|---|---|---|---|---|---|---|
| Transgenic line | To /℃ | Tp /℃ | Tc /℃ | △H /(J∙g-1) | AAC/% | GC/mm | Protein content/% | |
| 龙特甫B LTFB | 63.6±0.1 | 69.5±0.1 | 77.2±0.3 | 6.30±0.41 | 23.16±1.55 | 19.0±0.0 | 10.47±0.12 | |
| L-ALKb-RNAi-1 | 61.8±0.4* | 68.2±0.0** | 76.2±0.5* | 6.40±0.49 | 24.42±0.13 | 18.0±0.0 | 9.80±0.17 | |
| L-ALKb-RNAi-2 | 62.2±0.3* | 68.3±0.1** | 76.0±0.5* | 6.30±0.82 | 24.01±0.15 | 18.0±0.1 | 9.73±0.19 | |
| L-ALKb-RNAi-3 | 62.2±0.5* | 68.6±0.1** | 76.0±0.0* | 6.00±0.31 | 23.66±0.26 | 17.7±0.1 | 10.11±0.49 | |
| 珍汕97B ZS97B | 74.6±0.2 | 79.2±0.2 | 85.2±0.1 | 9.43±0.16 | 25.11±0.04 | 58.5±0.4 | 10.61±0.04 | |
| Z-ALKc-RNAi-2-1 | 67.1±0.3** | 73.4±0.1** | 80.2±0.1** | 8.20±0.08** | 27.73±0.24** | 25.5±0.1** | 9.21±0.05* | |
| Z-ALKc-RNAi-2-2 | 66.3±0.0** | 72.9±0.1** | 79.4±0.2** | 7.99±0.12** | 27.85±0.13** | 28.0±0.3** | 9.65±0.42* | |
| Z-ALKc-RNAi-2-3 | 65.6±0.1** | 72.4±0.1** | 79.0±0.0** | 7.29±0.07** | 26.69±0.01** | 45.0±0.0** | 9.03±0.46* | |
Table 2 Gelatinization properties and physical-chemical qualities of ALK-RNAi transgenic rice.
| 转基因系 | 起始温度 | 峰值温度 | 终止温度 | 热焓值 | 直链淀粉含量 | 胶稠度 | 粗蛋白含量 | |
|---|---|---|---|---|---|---|---|---|
| Transgenic line | To /℃ | Tp /℃ | Tc /℃ | △H /(J∙g-1) | AAC/% | GC/mm | Protein content/% | |
| 龙特甫B LTFB | 63.6±0.1 | 69.5±0.1 | 77.2±0.3 | 6.30±0.41 | 23.16±1.55 | 19.0±0.0 | 10.47±0.12 | |
| L-ALKb-RNAi-1 | 61.8±0.4* | 68.2±0.0** | 76.2±0.5* | 6.40±0.49 | 24.42±0.13 | 18.0±0.0 | 9.80±0.17 | |
| L-ALKb-RNAi-2 | 62.2±0.3* | 68.3±0.1** | 76.0±0.5* | 6.30±0.82 | 24.01±0.15 | 18.0±0.1 | 9.73±0.19 | |
| L-ALKb-RNAi-3 | 62.2±0.5* | 68.6±0.1** | 76.0±0.0* | 6.00±0.31 | 23.66±0.26 | 17.7±0.1 | 10.11±0.49 | |
| 珍汕97B ZS97B | 74.6±0.2 | 79.2±0.2 | 85.2±0.1 | 9.43±0.16 | 25.11±0.04 | 58.5±0.4 | 10.61±0.04 | |
| Z-ALKc-RNAi-2-1 | 67.1±0.3** | 73.4±0.1** | 80.2±0.1** | 8.20±0.08** | 27.73±0.24** | 25.5±0.1** | 9.21±0.05* | |
| Z-ALKc-RNAi-2-2 | 66.3±0.0** | 72.9±0.1** | 79.4±0.2** | 7.99±0.12** | 27.85±0.13** | 28.0±0.3** | 9.65±0.42* | |
| Z-ALKc-RNAi-2-3 | 65.6±0.1** | 72.4±0.1** | 79.0±0.0** | 7.29±0.07** | 26.69±0.01** | 45.0±0.0** | 9.03±0.46* | |
| 转基因系 Transgenic line | 峰值黏度 Peak viscosity /cP | 热浆黏度 Hot paste viscosity/cP | 崩解值Breakdown /cP | 冷胶黏度 Cool paste viscosity /cP | 消减值 Setback /cP | 峰值时间 Peak time /min | 起浆温度 Pasting temperature /℃ |
|---|---|---|---|---|---|---|---|
| 龙特甫B LTFB | 3699±11 | 3217±18 | 482±7 | 6385±18 | 2686±5 | 6.5 | 87.9 |
| L-ALKb-RNAi-1 | 3923±7** | 3390±10** | 533±13** | 6390±11 | 2467±8** | 6.5 | 87.0* |
| L-ALKb-RNAi-2 | 3810±13** | 3312±25** | 498±8* | 6055±7** | 2245±6** | 6.5 | 86.9* |
| L-ALKb-RNAi-3 | 3843±8** | 3137±17* | 706±10** | 6345±14* | 2502±7** | 6.5 | 87.8* |
| 珍汕97B ZS97B | 2885±12 | 2510±16 | 375±9 | 4974±15 | 2089±9 | 6.5 | 88.6 |
| Z-ALKc-RNAi-2-1 | 3608±9** | 2945±17** | 663±12** | 4749±17** | 1141±11** | 6.5 | 76.5** |
| Z-ALKc-RNAi-2-2 | 3930±11** | 3249±13** | 681±11** | 4590±16** | 660±12** | 6.6 | 77.4** |
| Z-ALKc-RNAi-2-3 | 3976±6** | 3337±14** | 639±8** | 4956±19 | 980±14** | 6.5 | 76.5** |
Table 3 RVA parameters of mature rice flour in ALK-RNAi transgenic rice lines.
| 转基因系 Transgenic line | 峰值黏度 Peak viscosity /cP | 热浆黏度 Hot paste viscosity/cP | 崩解值Breakdown /cP | 冷胶黏度 Cool paste viscosity /cP | 消减值 Setback /cP | 峰值时间 Peak time /min | 起浆温度 Pasting temperature /℃ |
|---|---|---|---|---|---|---|---|
| 龙特甫B LTFB | 3699±11 | 3217±18 | 482±7 | 6385±18 | 2686±5 | 6.5 | 87.9 |
| L-ALKb-RNAi-1 | 3923±7** | 3390±10** | 533±13** | 6390±11 | 2467±8** | 6.5 | 87.0* |
| L-ALKb-RNAi-2 | 3810±13** | 3312±25** | 498±8* | 6055±7** | 2245±6** | 6.5 | 86.9* |
| L-ALKb-RNAi-3 | 3843±8** | 3137±17* | 706±10** | 6345±14* | 2502±7** | 6.5 | 87.8* |
| 珍汕97B ZS97B | 2885±12 | 2510±16 | 375±9 | 4974±15 | 2089±9 | 6.5 | 88.6 |
| Z-ALKc-RNAi-2-1 | 3608±9** | 2945±17** | 663±12** | 4749±17** | 1141±11** | 6.5 | 76.5** |
| Z-ALKc-RNAi-2-2 | 3930±11** | 3249±13** | 681±11** | 4590±16** | 660±12** | 6.6 | 77.4** |
| Z-ALKc-RNAi-2-3 | 3976±6** | 3337±14** | 639±8** | 4956±19 | 980±14** | 6.5 | 76.5** |
| 样品 Sample | 转基因系 Transgenic line | 综合 Integrated score | 外观 Appearance | 口感 Taste |
|---|---|---|---|---|
| 热饭 Cooked rice | 龙特甫B LTFB | 53.3±1.3 | 5.0±0.1 | 4.5±0.2 |
| L-ALKb-RNAi-1 | 48.3±1.3** | 4.3±0.1** | 3.9±0.2* | |
| L-ALKb-RNAi-2 | 51.0±0.8* | 4.7±0.1* | 4.2±0.1* | |
| L-ALKb-RNAi-3 | 50.5±1.7** | 4.6±0.2* | 4.1±0.2* | |
| 珍汕97B ZS97B | 55.0±2.6 | 5.1±0.3 | 4.8±0.3 | |
| Z-ALKc-RNAi-2-1 | 43.8±1.9** | 3.6±0.3** | 3.3±0.3** | |
| Z-ALKc-RNAi-2-2 | 42.0±0.8** | 3.5±0.0** | 3.1±0.1** | |
| Z-ALKc-RNAi-2-3 | 44.5±0.6** | 3.8±0.0** | 3.4±0.0** | |
| 冷饭 Cooled rice | 龙特甫B LTFB | 49.8±0.5 | 4.6±0.0 | 3.9±0.1 |
| L-ALKb-RNAi-1 | 44.8±0.5** | 4.1±0.1** | 3.6±0.1** | |
| L-ALKb-RNAi-2 | 48.5±1.0* | 4.3±0.1* | 3.7±0.1* | |
| L-ALKb-RNAi-3 | 46.5±0.6** | 4.3±0.1* | 3.7±0.1* | |
| 珍汕97B ZS97B | 54.0±0.8 | 5.1±0.0 | 4.7±0.1 | |
| Z-ALKc-RNAi-2-1 | 45.8±0.5** | 3.0±1.4** | 3.4±0.0** | |
| Z-ALKc-RNAi-2-2 | 44.8±1.7** | 3.4±0.2* | 3.1±0.4** | |
| Z-ALKc-RNAi-2-3 | 47.8±2.1** | 4.0±0.2 | 3.6±0.3** |
Table 4 Sensory evaluation of milled rice in ALK-RNAi transgenic rice lines.
| 样品 Sample | 转基因系 Transgenic line | 综合 Integrated score | 外观 Appearance | 口感 Taste |
|---|---|---|---|---|
| 热饭 Cooked rice | 龙特甫B LTFB | 53.3±1.3 | 5.0±0.1 | 4.5±0.2 |
| L-ALKb-RNAi-1 | 48.3±1.3** | 4.3±0.1** | 3.9±0.2* | |
| L-ALKb-RNAi-2 | 51.0±0.8* | 4.7±0.1* | 4.2±0.1* | |
| L-ALKb-RNAi-3 | 50.5±1.7** | 4.6±0.2* | 4.1±0.2* | |
| 珍汕97B ZS97B | 55.0±2.6 | 5.1±0.3 | 4.8±0.3 | |
| Z-ALKc-RNAi-2-1 | 43.8±1.9** | 3.6±0.3** | 3.3±0.3** | |
| Z-ALKc-RNAi-2-2 | 42.0±0.8** | 3.5±0.0** | 3.1±0.1** | |
| Z-ALKc-RNAi-2-3 | 44.5±0.6** | 3.8±0.0** | 3.4±0.0** | |
| 冷饭 Cooled rice | 龙特甫B LTFB | 49.8±0.5 | 4.6±0.0 | 3.9±0.1 |
| L-ALKb-RNAi-1 | 44.8±0.5** | 4.1±0.1** | 3.6±0.1** | |
| L-ALKb-RNAi-2 | 48.5±1.0* | 4.3±0.1* | 3.7±0.1* | |
| L-ALKb-RNAi-3 | 46.5±0.6** | 4.3±0.1* | 3.7±0.1* | |
| 珍汕97B ZS97B | 54.0±0.8 | 5.1±0.0 | 4.7±0.1 | |
| Z-ALKc-RNAi-2-1 | 45.8±0.5** | 3.0±1.4** | 3.4±0.0** | |
| Z-ALKc-RNAi-2-2 | 44.8±1.7** | 3.4±0.2* | 3.1±0.4** | |
| Z-ALKc-RNAi-2-3 | 47.8±2.1** | 4.0±0.2 | 3.6±0.3** |
| [1] | 陈静. 江苏省水稻食味改良育种研究进展. 江苏农业科学, 2015, 43(12): 77-80. |
| Chen J.A review of the research on the improvement of rice taste in Jiangsu Province.Jiangsu Agric Sci, 2015, 43(12): 77-80. (in Chinese) | |
| [2] | Tian Z X, Qian Q, Liu Q Q, Yan M X, Liu X F, Yan C J, Liu G F, Gao Z Y, Tang S Z, Zeng D L, Wang Y H, Yu J M, Gu M H, Li J Y.Allelic diversities in rice starch biosynthesis lead to diverse array of rice eating and cooking qualities.Proc Natl Acad Sci USA, 2009, 106(51): 21 760-21 765. |
| [3] | Xu C W, Mo H D.Qualitative-quantitative analysis for inheritance of gelatinization temperature in indica rice(O. sativa subsp. indica). Acta Agron Sin, 1996, 22(4): 386-389. |
| [4] | Khush G S, Paule C M, De La Cruz, N M. Rice grain quality evaluation and improvement at IRRI//The Workshop on Chemical Aspects of Rice Grain Quality, 1978. |
| [5] | Chun A, Lee H J, Hamaker B R, Janaswamy S.Effects of ripening temperature on starch structure and gelatinization, pasting, and cooking properties in rice (Oryza sativa). J Agric Food Chem, 2015, 63(12): 3085-3093. |
| [6] | Nakamura Y, Francisco J P, Hosaka Y, Sato A, Sawada T, Kubo A, Fujita N.Essential amino acids of starch synthase IIa differentiate amylopectin structure and starch quality between japonica and indica rice varieties. Plant Mol Biol, 2005, 58(2): 213-227. |
| [7] | Umemoto T, Yano M, Satoh H, Shomura A, Nakamura Y.Mapping of a gene responsible for the difference in amylopectin structure betweenjaponica-type and indica-type rice varieties. Theor Appl Genet, 2002, 104(1): 1-8. |
| [8] | Gao Z Y, Zeng D L, Cui X, Zhou Y H, Yan M X, Huang D N, Li J Y, Qian Q.Map-based cloning of theALK gene, which controls the gelatinization temperature of rice. Sci China C Life Sci, 2003, 46(6): 661-668. |
| [9] | Bao J S, Sun M, Zhu L H, Corke H.Analysis of quantitative trait loci for some starch properties of rice (Oryza sativa L.): Thermal properties, gel texture and swelling volume. J Cereal Sci, 2004, 39(3): 379-385. |
| [10] | Fan C C, Yu X Q, Xing Y Z, Xu C G, Luo L J, Zhang Q F.The main effects, epistatic effects and environmental interactions of QTLs on the cooking and eating quality of rice in a doubled-haploid line population.Theor Appl Genet, 2005, 110(8): 1445-1452. |
| [11] | Ebadi A A, Farshadfar E, Rabiei B.Mapping QTLs controlling cooking and eating quality indicators of Iranian rice using RILs across three years.Aust J Crop Sci, 2013, 7(10): 1494-1502. |
| [12] | Zhao W G, Chung J W, Kwon S W, Lee J H, Ma K H, Park Y J.Association analysis of physicochemical traits on eating quality in rice (Oryza sativa L.). Euphytica, 2013, 191(1): 9-21. |
| [13] | Gao Z Y, Zeng D L, Cheng F M, Tian Z X, Guo L B, Su Y, Yan M X, Jiang H, Dong G J, Huang Y C, Han B, Li J Y, Qian Q.ALK, the key gene for gelatinization temperature,is a modifier gene for gel consistency in rice. J Integr Plant Biol, 2011, 53(9): 756-765. |
| [14] | Nakamura Y, Francisco P Y, Sato A, Sawada T, Kubo A, Fujita N.Essential amino acids of starch synthase IIa differentiate amylopectin structure and starch quality betweenjaponica and indica rice varieties. Plant Mol Biol, 2005, 58(2): 213-227. |
| [15] | Cuevas R P, Daygon V D, Corpuz H, Nora L, Reinke R, Waters D, Fitzgerald M.Melting the secrets of gelatinisation temperature in rice.Funct Plant Biol, 2010, 37(5): 439-447. |
| [16] | Bao J S, Peng X, Hiratsuka M, Sun M, Umemoto T.Granule-bound SSIIa protein content and its relationship with amylopectin structure and gelatinization temperature of rice starch.Starch-Stärke, 2010, 61(8): 431-437. |
| [17] | Kharabianmasouleh A, Waters D L E, Reinke R F, Ward R, Henry R J. SNP in starch biosynthesis genes associated with nutritional and functional properties of rice.Sci Rep, 2012, 2: 557. |
| [18] | Waters D L E, Henry R J, Reinke R F, Fitzgerald M A. Gelatinization temperature of rice explained by polymorphisms in starch synthase.Plant Biotechnol J, 2010, 4(1): 115-122. |
| [19] | Chen M H, Bergman C J, Fjellstrom R G.SSSIIA locus genetic variation associated with alkali spreading value in international rice germplasm. Proceedings of the Plant and Animal Genome Conference, 2003, 153. |
| [20] | Bao J S, Corke H, Sun M.Nucleotide diversity in starch synthase IIa and validation of single nucleotide polymorphisms in relation to starch gelatinization temperature and other physicochemical properties in rice (Oryza sativa L.). Theor Appl Genet, 2006, 113(7): 1171-1183. |
| [21] | Zhou Y, Zheng H Y, Wei G C, Zhou H, Han Y N, Bai X F, Xing Y Z, Han Y P.Nucleotide Diversity and Molecular Evolution of the ALK Gene in Cultivated Rice and its Wild Relatives. Plant Mol Biol Rep, 2016, 34(5): 1-8. |
| [22] | 陈秀花, 刘巧泉, 王宗阳, 王兴稳, 蔡秀玲, 张景六, 顾铭洪. 反义Wx基因导入我国籼型杂交稻重点亲本. 科学通报, 2002, 47(9): 684-688. |
| Chen X H, Liu Q Q, Wang Z Y, Wang X W, Cai X L, Zhang J L, Gu M H.Introduction of antisense Wx Gene into key parents of indica hybrid rice in China. Chin Sci Bull, 2002, 47(9): 684-688. (in Chinese) | |
| [23] | Murray M G, Thompson W F.Rapid isolation of high molecular weight plant DNA.Nucl Acids Res, 1980, 8(19): 4321-4325. |
| [24] | Zhang C Q, Zhou L H, Zhu Z B, Lu H W, Zhou X Z, Qian Y T, Li Q F, Lu Y, Gu M H, Liu Q Q.Characterization of grain quality and starch fine structure of two japonica rice(Oryza sativa) cultivars with good sensory properties at different temperatures during the filling stage. J Agric Food Chem, 2016, 64(20): 4048-4057. |
| [25] | Zhu L J, Meng X L, Shi Y C, Gu M H, Liu Q Q.High-amylose rice improves indices of animal health in normal and diabetic rats.Plant Biotechnol J, 2012, 10(3): 353-362. |
| [26] | Umemoto T, Horibata T, Aoki N, Hiratsuka M, Yano M, Inouchi N.Effects of variations in starch synthase on starch properties and eating quality of rice.Plant Prod Sci, 2008, 11(4): 472-480. |
| [27] | Miura S, Crofts N, Saito Y, Hosaka Y, Oitome N F, Watanabe T, Kumamaru T, Fujita N.Starch Synthase IIa-deficient mutant rice line produces endosperm starch with lower gelatinization temperature than japonica rice cultivars. Front Plant Sci, 2018, 9: 1-10. |
| [28] | Umemoto T, Aoki N, Lin H, Nakamura Y, Inouchi N, Sato Y.Natural variation in rice starch synthase IIa affects enzyme and starch properties.Funct Plant Biol, 2004, 31(7): 671-684. |
| [29] | Kaw R N, Cruz N M D L. Genetic analysis of amylose content, gelatinization temperature and gel consistency in rice.J Genet Breed, 1990: 103-111. |
| [30] | 李晓方, 肖昕, 刘彦卓, 刘志霞, 卢东柏, 毛兴学, 杨俊, 方正武, 李志新. 籼稻稻米品质性状遗传特点新解析. 分子植物育种, 2009, 7(6): 1077-1083. |
| Li X F, Xiao X, Liu Y Z, Liu Z X, Lu D B, Mao X X, Yang J, Fang Z W, Li Z X.Novel anlysis on genetic characters of quality traits in indica rice.Mol Plant Breed, 2009, 7(6): 1077-1083. (in Chinese with English abstract) | |
| [31] | 隋炯明, 李欣, 严松, 严长杰, 张蓉, 汤述翥, 陆驹飞, 陈宗祥, 顾铭洪. 稻米淀粉RVA谱特征与品质性状相关性研究. 中国农业科学, 2005, 38(4): 657-663. |
| Sui J M, Li X, Yan S, Yan C J, Zhang R, Tang S Z, Lu J F, Chen Z X, Gu M H.Studies on the rice RVA profile characteristics and its correlation with the quality.Sci Agric Sin, 2005, 38(4): 657-663. (in Chinese with English abstract) | |
| [32] | 曹清明, 钟海雁, 李忠海, 孙昌波. 蕨根淀粉糊化温度测定及影响因素研究. 食品与机械, 2007, 23(3): 16-19. |
| Cao Q M, Zhong H Y, Li Z H, Sun C B.Studies on the rice RVA profile characteristics and its correlation with the quality. Food & Mach, 2007, 23(3): 16-19. (in Chinese with English abstract) | |
| [33] | 张昌泉, 潘立旭, 周兴忠, 李钱峰, 刘巧泉. 不同Wx等位组合对杂交稻天优3611品质的影响 . 中国科技论文在线, 2007. |
| Zhang C Q, Pan L X, Zhou X Z, Li Q F, Liu Q Q.The combined effects of different Wx allele on grain quality of hybrid rice Tianyou 3611. Chin Sci Paper,2007. | |
| [34] | 刘鑫燕, 朱孔志, 张昌泉, 洪燃, 孙鹏, 汤述翥, 顾铭洪, 刘巧泉. 利用9311来源的粳型染色体片段代换系定位控制稻米糊化温度的微效QTL. 作物学报, 2014, 40(10): 1740-1747. |
| Liu X Y, Zhu K Z, Zhang C Q, Hong R, Sun P, Tang S Z, Gu M H, Liu Q Q.Mapping of minor QTLs for rice gelatinization temperature using chromosome segment substitution lines from indica 9311 in the japonica background. Acta Agron Sin, 2014, 40(10): 1740-1747. (in Chinese with English abstract) |
| [1] | GUO Zhan, ZHANG Yunbo. Research Progress in Physiological,Biochemical Responses of Rice to Drought Stress and Its Molecular Regulation [J]. Chinese Journal OF Rice Science, 2024, 38(4): 335-349. |
| [2] | WEI Huanhe, MA Weiyi, ZUO Boyuan, WANG Lulu, ZHU Wang, GENG Xiaoyu, ZHANG Xiang, MENG Tianyao, CHEN Yinglong, GAO Pinglei, XU Ke, HUO Zhongyang, DAI Qigen. Research Progress in the Effect of Salinity, Drought, and Their Combined Stresses on Rice Yield and Quality Formation [J]. Chinese Journal OF Rice Science, 2024, 38(4): 350-363. |
| [3] | XU Danjie, LIN Qiaoxia, LI Zhengkang, ZHUANG Xiaoqian, LING Yu, LAI Meiling, CHEN Xiaoting, LU Guodong. OsOPR10 Positively Regulates Rice Blast and Bacterial Blight Resistance [J]. Chinese Journal OF Rice Science, 2024, 38(4): 364-374. |
| [4] | CHEN Mingliang, ZENG Xihua, SHEN Yumin, LUO Shiyou, HU Lanxiang, XIONG Wentao, XIONG Huanjin, WU Xiaoyan, XIAO Yeqing. Typing of Inter-subspecific Fertility Loci and Fertility Locus Pattern of indica-japonica Hybrid Rice [J]. Chinese Journal OF Rice Science, 2024, 38(4): 386-396. |
| [5] | DING Zhengquan, PAN Yueyun, SHI Yang, HUANG Haixiang. Comprehensive Evaluation and Comparative Analysis of Jiahe Series Long-Grain japonica Rice with High Eating Quality Based on Gene Chip Technology [J]. Chinese Journal OF Rice Science, 2024, 38(4): 397-408. |
| [6] | HOU Xiaoqin, WANG Ying, YU Bei, FU Weimeng, FENG Baohua, SHEN Yichao, XIE Hangjun, WANG Huanran, XU Yongqiang, WU Zhihai, WANG Jianjun, TAO Longxing, FU Guanfu. Mechanisms Behind the Role of Potassium Fulvic Acid in Enhancing Salt Tolerance in Rice Seedlings [J]. Chinese Journal OF Rice Science, 2024, 38(4): 409-421. |
| [7] | LÜ Zhou, YI Binghuai, CHEN Pingping, ZHOU Wenxin, TANG Wenbang, YI Zhenxie. Effects of Nitrogen Application Rate and Transplanting Density on Yield Formation of Small Seed Hybrid Rice [J]. Chinese Journal OF Rice Science, 2024, 38(4): 422-436. |
| [8] | HU Jijie, HU Zhihua, ZHANG Junhua, CAO Xiaochuang, JIN Qianyu, ZHANG Zhiyuan, ZHU Lianfeng. Effects of Rhizosphere Saturated Dissolved Oxygen on Photosynthetic and Growth Characteristics of Rice at Tillering Stage [J]. Chinese Journal OF Rice Science, 2024, 38(4): 437-446. |
| [9] | WU Yue, LIANG Chengwei, ZHAO Chenfei, SUN Jian, MA Dianrong. Occurrence of Weedy Rice Disaster and Ecotype Evolution in Direct-Seeded Rice Fields [J]. Chinese Journal OF Rice Science, 2024, 38(4): 447-455. |
| [10] | LIU Fuxiang, ZHEN Haoyang, PENG Huan, ZHENG Liuchun, PENG Deliang, WEN Yanhua. Investigation and Species Identification of Cyst Nematode Disease on Rice in Guangdong Province [J]. Chinese Journal OF Rice Science, 2024, 38(4): 456-461. |
| [11] | CHEN Haotian, QIN Yuan, ZHONG Xiaohan, LIN Chenyu, QIN Jinghang, YANG Jianchang, ZHANG Weiyang. Research Progress on the Relationship Between Rice Root, Soil Properties and Methane Emissions in Paddy Fields [J]. Chinese Journal OF Rice Science, 2024, 38(3): 233-245. |
| [12] | MIAO Jun, RAN Jinhui, XU Mengbin, BO Liubing, WANG Ping, LIANG Guohua, ZHOU Yong. Overexpression of RGG2, a Heterotrimeric G Protein γ Subunit-Encoding Gene, Improves Drought Tolerance in Rice [J]. Chinese Journal OF Rice Science, 2024, 38(3): 246-255. |
| [13] | YIN Xiaoxiao, ZHANG Zhihan, YAN Xiulian, LIAO Rong, YANG Sijia, Beenish HASSAN, GUO Daiming, FAN Jing, ZHAO Zhixue, WANG Wenming. Signal Peptide Validation and Expression Analysis of Multiple Effectors from Ustilaginoidea virens [J]. Chinese Journal OF Rice Science, 2024, 38(3): 256-265. |
| [14] | ZHU Yujing, GUI Jinxin, GONG Chengyun, LUO Xinyang, SHI Jubin, ZHANG Haiqing, HE Jiwai. QTL Mapping for Tiller Angle in Rice by Genome-wide Association Analysis [J]. Chinese Journal OF Rice Science, 2024, 38(3): 266-276. |
| [15] | WEI Qianqian, WANG Yulei, KONG Haimin, XU Qingshan, YAN Yulian, PAN Lin, CHI Chunxin, KONG Yali, TIAN Wenhao, ZHU Lianfeng, CAO Xiaochuang, ZHANG Junhua, ZHU Chunqun. Mechanism of Hydrogen Sulfide, a Signaling Molecule Involved in Reducing the Inhibitory Effect of Aluminum Toxicity on Rice Growth Together with Sulfur Fertilizer [J]. Chinese Journal OF Rice Science, 2024, 38(3): 290-302. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||