
中国水稻科学 ›› 2026, Vol. 40 ›› Issue (2): 145-154.DOI: 10.16819/j.1001-7216.2026.250706
• 综述与专论 • 下一篇
杨大兵, 杜雪树, 李进波, 夏明元, 胡亮, 石桓, 万丙良*( )
)
收稿日期:2025-07-17
修回日期:2025-10-21
出版日期:2026-03-10
发布日期:2026-03-16
通讯作者:
* email:ricewanbl@126.com基金资助:
YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang*( )
)
Received:2025-07-17
Revised:2025-10-21
Online:2026-03-10
Published:2026-03-16
Contact:
* email:ricewanbl@126.com摘要:
抽穗期是决定水稻生产季节与地域适应性的关键农艺性状,了解水稻抽穗期调控分子机理对于培育适应特定生态环境的高产水稻品种具有重要意义。近年来,大量水稻抽穗期相关基因相继被克隆,其功能逐渐明晰,抽穗期调控的分子网络也日趋完善。本文系统综述了以光周期调控网络为核心的水稻抽穗期调控分子机理,及其在育种实践中的应用进展,以期为水稻抽穗期遗传改良和生态适应性育种提供理论依据。
杨大兵, 杜雪树, 李进波, 夏明元, 胡亮, 石桓, 万丙良. 水稻抽穗期调控的分子机理及育种应用进展[J]. 中国水稻科学, 2026, 40(2): 145-154.
YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang. Advances in Molecular Mechanism and Breeding Application of Heading Date Regulation in Rice[J]. Chinese Journal OF Rice Science, 2026, 40(2): 145-154.
| 基因符号 Gene symbol | RAP位点 RAP locus | 功能 Function | 参考文献 Reference |
|---|---|---|---|
| LHD3/OsphyC/PHYC | Os03g0752100 | 光敏色素基因,与PHYB与PHYA协同感应红光和远红光。 | [ |
| PHYB | Os03g0309200 | 光敏色素基因,红光受体。调节Hd1介导的水稻成花素Hd3a的表达和临界日长。 | [ |
| PHYA | Os03g0719800 | 光敏色素基因,远红光受体。与PHYB 、PHYC协同调控水稻抽穗期。 | [ |
| OsCRY2 | Os02g0625000 | 隐花色素基因,蓝光受体。 | [ |
| OsPRR73/OsCCT11 | Os03g0284100 | 生物钟核心转录因子,通过结合Ehd1启动子,参与水稻昼夜节律钟的反馈回路,并连接光周期开花途径。 | [ |
| PCL1/OsLUX | Os01g0971800 | 生物钟核心元件,通过招募OsELF3-1和OsELF4s组成三元抑制蛋白复合物OsEC1,抑制Hd1和Ghd7基因表达。 | [ |
| OsCCA1/OsLHY/Nhd1 | Os08g0157600 | 生物钟核心元件,通过OsGI-Hd1路径精准调控抽穗期的日长临界点,也是氮素介导的抽穗开花因子。 | [ |
| Hd17/OsELF3/ OsELF3-1 | Os06g0142600 | 生物钟元件,与OsLUX和OsELF4s形成OsEC1复合物,通过结合Hd1和Ghd7启动子,抑制基因表达。 | [ |
| ELF4A | Os11g0621500 | 生物钟元件,与OsELF3-1和OsLUX组成三元蛋白复合物OsEC1,抑制Hd1和Ghd7的表达。 | [ |
| OsGI | Os01g0182600 | 生物钟元件,长日照条件下抑制抽穗,受三元抑制蛋白复合物OsEC1的直接调控。 | [ |
| Hd1 | Os06g0275000 | 抽穗期调控核心基因,在长日照条件下,与Ghd7形成复合体共同抑制Ehd1基因的表达,从而延迟抽穗;在短日照条件下促进抽穗。 | [ |
| Ehd1 | Os10g0463400 | 抽穗期调控核心基因,短日照条件下正向调控Hd3a/RFT1,促进抽穗。 | [ |
| Ghd7 | Os07g0261200 | 抽穗期调控核心基因,每穗粒数、株高和抽穗期多效性控制基因。长日照条件下,增强表达能推迟抽穗、增加株高和每穗粒数。 | [ |
| Ghd8/DTH8/Hd5/ OsHAP3H/LHD1 | Os08g0174500 | 籽粒产量、株高和抽穗期多效性控制基因,开花抑制因子。长日照条件下,DTH8协同Ghd7抑制Ehd1、Hd3a/RFT1表达,从而延迟开花时间。 | [ |
| Hd3a/FTL2 | Os06g0157700 | 成花素基因,将上游光周期开花信号传递给下游开花基因。主要在短日照条件下起作用。 | [ |
| RFT1/FTL3 | Os06g0157500 | 成花素基因,将上游光周期开花信号传递给下游开花基因。长日照条件下的主要开花激活因子。 | [ |
| OsC2DP5/OsFTIP9 | Os01g0587300 | 成花素互作蛋白基因,介导Hd3a/RFT1从叶片向茎端分生组织(SAM)的运输。 | [ |
| OsFTIP1 | Os06g0614000 | 成花素互作蛋白基因,负责将成花素RFT1从叶片向茎尖分生组织(SAM)的转运。 | [ |
| RIFLA/OsMADS56 | Os10g0536150 | MADS盒基因,与OsMADS50相互作用形成复合物,调控下游基因OsLFL1-Ehd1的表达。 | [ |
| OsMADS51 | Os01g0922800 | MADS盒基因,开花促进因子,在短日照下,作用于OsGI 下游,参与将OsGI的信号传递至Ehd1。 | [ |
| OsMADS14 | Os03g0752800 | MADS盒基因,受成花素激活复合体FAC调控,促进抽穗。 | [ |
| OsMADS50/OsSOC1/ DTH3 | Os03g0122600 | MADS盒基因,与其他MADS-box基因如OsMADS56相互作用,形成复合物,协同调控下游基因OsLFL1-Ehd1。 | [ |
| Ehd3 | Os08g0105000 | 早抽穗基因,长日照下,Ehd3通过抑制Ghd7诱导水稻抽穗,同时,又能通过不依赖Ghd7的方式上调Ehd1的表达促进抽穗。 | [ |
| OsTOC1/ OsPRR1 | Os11g0157600 | 转录抑制因子,作用于生物钟钟核心元件,如OsLHY和OsGI,抑制其表达。 | [ |
| OsRR1 | Os04g0442300 | A型反应调节因子,与Ehd1结合形成异二聚体,抑制Ehd1的活性,延迟开花。 | [ |
| OsSGI1/OsHBP1 | Os04g0489600 | bHLH转录因子,激活Hd1表达,促进抽穗。 | [ |
| OsFD1/OsbZIP77 | Os09g0540800 | b-ZIP转录因子,与成花素Hd3a/RFT1与14-3-3蛋白形成成花素复合物,诱导OsMADS15/OsMADS14转录,促进抽穗。 | [ |
| OsbZIP69/OsFD4 | Os08g0549600 | bZIP转录因子,与Gf14s、RFT1互作,形成成花素激活复合物,促进成花转变。 | [ |
| OsbZIP1/OsRE1 | Os01g0174000 | bZIP转录因子,抑制Ehd1表达,微调抽穗期。 | [ |
| OsbZIP42/HBF1 | Os05g0489700 | 开花抑制因子,抑制Ehd1表达,延迟抽穗。 | [ |
| Ehd2/RID1/OsId1/ Ghd10 | Os10g0419200 | 锌指转录因子,通过上调Ehd1的表达来促进抽穗。 | [ |
| Ghd2 | Os02g0731700 | 控制抽穗期、株高和每穗颖花数的多效性基因,通过抑制Ehd1途径延迟抽穗。 | [ |
| Ehd4 | Os03g0112700 | CCH类锌指蛋白基因,通过上调Ehd1的表达来促进Hd3a和RFT1的表达。 | [ |
| OsCO3 | Os09g0240200 | CONSTANS基因,通过抑制Ehd1、Hd3a和RFT1的转录来负向调节开花。 | [ |
| DHD4 | Os02g0110100 | CONSTANS转录因子,与OsFD1相互作用,影响Hd3a-14-3-3-OsFD1三蛋白FAC复合体的形成,延迟抽穗。 | [ |
| OsCOL4 | Os02g0610500 | CONSTANS转录因子,组成型的开花抑制因子,作用于Ehd1上游。 | [ |
| OsCOL15 | Os08g0536300 | CONSTANS转录因子,通过上调Ghd7和下调RID1来抑制抽穗。 | [ |
| HD6/CK2α | Os03g0762000 | 酪蛋白激酶CK2α亚基,通过调控Hd1基因,在长日照条件下延迟抽穗。 | [ |
| Hd16/CKI/EL1 | Os03g0793500 | 酪蛋白激酶CKI,长日条件下磷酸化Ghd7,增强其功能,延迟抽穗。 | [ |
| OsFTL12 | Os06g0552900 | FT家族蛋白,与GF14b和OsFD1相互作用,形成成花素抑制复合体FRC,调控OsMADS14和OsMADS15,延迟抽穗。 | [ |
| IDD4/SID1 | Os02g0672100 | IDD转录因子,作用于Hd3a和RFT1的启动子区域,启动水稻开花转变。 | [ |
| OsRIP1 | Os04g0540200 | OsRE1相互作用蛋白,在依赖OsRE1的方式下抑制Ehd1的转录表达调控抽穗期。 | [ |
| OsVIL1 | Os12g0533500 | 与OsVIL2互作,形成PRC2复合体,短日照条件下通过抑制OsLFL1表达诱导抽穗,而长日照条件下通过提高Ghd7表达延迟抽穗。 | [ |
| Ehd5 | Os08g0493900 | WD40结构域蛋白,与Roc4和Ghd8相互作用,影响Ghd7-Ghd8复合物的形成,促进开花相关基因的表达。 | [ |
| DTH7/Ghd7.1/ OsPRR37/Hd2 | Os07g0695100 | PRR蛋白,长日条件下,抑制Ehd1表达延迟抽穗;短日条件下,根据互作基因的不同,既可抑制也可促进抽穗。 | [ |
| SDG724/OsSET34 | Os09g0307800 | 组蛋白H3K36甲基转移酶,介导MADS50和RFT1的H3K36me2/3沉积,促进水稻开花。 | [ |
| Hd18 | Os08g0143400 | 组蛋白乙酰转移酶基因,通过提高Ehd1的转录水平促进抽穗。 | [ |
| OsSUF4 | Os09g0560900 | 锌指转录因子,促进H3K36me3对RFT1和Hd3a的修饰,从而促进抽穗。 | [ |
| OsHUB2/ FRRP1 | Os10g0565600 | E3泛素连接酶,对Ehd1的组蛋白H2B进行泛素化,抑制抽穗。 | [ |
| HAF1 | Os04g0648800 | E3泛素连接酶,作用于Hd1并将其降解,精细调控Hd1昼夜节律的积累。 | [ |
表1 水稻抽穗期调控网络的主要基因
Table 1. Key genes in the regulatory network controlling heading date in rice
| 基因符号 Gene symbol | RAP位点 RAP locus | 功能 Function | 参考文献 Reference |
|---|---|---|---|
| LHD3/OsphyC/PHYC | Os03g0752100 | 光敏色素基因,与PHYB与PHYA协同感应红光和远红光。 | [ |
| PHYB | Os03g0309200 | 光敏色素基因,红光受体。调节Hd1介导的水稻成花素Hd3a的表达和临界日长。 | [ |
| PHYA | Os03g0719800 | 光敏色素基因,远红光受体。与PHYB 、PHYC协同调控水稻抽穗期。 | [ |
| OsCRY2 | Os02g0625000 | 隐花色素基因,蓝光受体。 | [ |
| OsPRR73/OsCCT11 | Os03g0284100 | 生物钟核心转录因子,通过结合Ehd1启动子,参与水稻昼夜节律钟的反馈回路,并连接光周期开花途径。 | [ |
| PCL1/OsLUX | Os01g0971800 | 生物钟核心元件,通过招募OsELF3-1和OsELF4s组成三元抑制蛋白复合物OsEC1,抑制Hd1和Ghd7基因表达。 | [ |
| OsCCA1/OsLHY/Nhd1 | Os08g0157600 | 生物钟核心元件,通过OsGI-Hd1路径精准调控抽穗期的日长临界点,也是氮素介导的抽穗开花因子。 | [ |
| Hd17/OsELF3/ OsELF3-1 | Os06g0142600 | 生物钟元件,与OsLUX和OsELF4s形成OsEC1复合物,通过结合Hd1和Ghd7启动子,抑制基因表达。 | [ |
| ELF4A | Os11g0621500 | 生物钟元件,与OsELF3-1和OsLUX组成三元蛋白复合物OsEC1,抑制Hd1和Ghd7的表达。 | [ |
| OsGI | Os01g0182600 | 生物钟元件,长日照条件下抑制抽穗,受三元抑制蛋白复合物OsEC1的直接调控。 | [ |
| Hd1 | Os06g0275000 | 抽穗期调控核心基因,在长日照条件下,与Ghd7形成复合体共同抑制Ehd1基因的表达,从而延迟抽穗;在短日照条件下促进抽穗。 | [ |
| Ehd1 | Os10g0463400 | 抽穗期调控核心基因,短日照条件下正向调控Hd3a/RFT1,促进抽穗。 | [ |
| Ghd7 | Os07g0261200 | 抽穗期调控核心基因,每穗粒数、株高和抽穗期多效性控制基因。长日照条件下,增强表达能推迟抽穗、增加株高和每穗粒数。 | [ |
| Ghd8/DTH8/Hd5/ OsHAP3H/LHD1 | Os08g0174500 | 籽粒产量、株高和抽穗期多效性控制基因,开花抑制因子。长日照条件下,DTH8协同Ghd7抑制Ehd1、Hd3a/RFT1表达,从而延迟开花时间。 | [ |
| Hd3a/FTL2 | Os06g0157700 | 成花素基因,将上游光周期开花信号传递给下游开花基因。主要在短日照条件下起作用。 | [ |
| RFT1/FTL3 | Os06g0157500 | 成花素基因,将上游光周期开花信号传递给下游开花基因。长日照条件下的主要开花激活因子。 | [ |
| OsC2DP5/OsFTIP9 | Os01g0587300 | 成花素互作蛋白基因,介导Hd3a/RFT1从叶片向茎端分生组织(SAM)的运输。 | [ |
| OsFTIP1 | Os06g0614000 | 成花素互作蛋白基因,负责将成花素RFT1从叶片向茎尖分生组织(SAM)的转运。 | [ |
| RIFLA/OsMADS56 | Os10g0536150 | MADS盒基因,与OsMADS50相互作用形成复合物,调控下游基因OsLFL1-Ehd1的表达。 | [ |
| OsMADS51 | Os01g0922800 | MADS盒基因,开花促进因子,在短日照下,作用于OsGI 下游,参与将OsGI的信号传递至Ehd1。 | [ |
| OsMADS14 | Os03g0752800 | MADS盒基因,受成花素激活复合体FAC调控,促进抽穗。 | [ |
| OsMADS50/OsSOC1/ DTH3 | Os03g0122600 | MADS盒基因,与其他MADS-box基因如OsMADS56相互作用,形成复合物,协同调控下游基因OsLFL1-Ehd1。 | [ |
| Ehd3 | Os08g0105000 | 早抽穗基因,长日照下,Ehd3通过抑制Ghd7诱导水稻抽穗,同时,又能通过不依赖Ghd7的方式上调Ehd1的表达促进抽穗。 | [ |
| OsTOC1/ OsPRR1 | Os11g0157600 | 转录抑制因子,作用于生物钟钟核心元件,如OsLHY和OsGI,抑制其表达。 | [ |
| OsRR1 | Os04g0442300 | A型反应调节因子,与Ehd1结合形成异二聚体,抑制Ehd1的活性,延迟开花。 | [ |
| OsSGI1/OsHBP1 | Os04g0489600 | bHLH转录因子,激活Hd1表达,促进抽穗。 | [ |
| OsFD1/OsbZIP77 | Os09g0540800 | b-ZIP转录因子,与成花素Hd3a/RFT1与14-3-3蛋白形成成花素复合物,诱导OsMADS15/OsMADS14转录,促进抽穗。 | [ |
| OsbZIP69/OsFD4 | Os08g0549600 | bZIP转录因子,与Gf14s、RFT1互作,形成成花素激活复合物,促进成花转变。 | [ |
| OsbZIP1/OsRE1 | Os01g0174000 | bZIP转录因子,抑制Ehd1表达,微调抽穗期。 | [ |
| OsbZIP42/HBF1 | Os05g0489700 | 开花抑制因子,抑制Ehd1表达,延迟抽穗。 | [ |
| Ehd2/RID1/OsId1/ Ghd10 | Os10g0419200 | 锌指转录因子,通过上调Ehd1的表达来促进抽穗。 | [ |
| Ghd2 | Os02g0731700 | 控制抽穗期、株高和每穗颖花数的多效性基因,通过抑制Ehd1途径延迟抽穗。 | [ |
| Ehd4 | Os03g0112700 | CCH类锌指蛋白基因,通过上调Ehd1的表达来促进Hd3a和RFT1的表达。 | [ |
| OsCO3 | Os09g0240200 | CONSTANS基因,通过抑制Ehd1、Hd3a和RFT1的转录来负向调节开花。 | [ |
| DHD4 | Os02g0110100 | CONSTANS转录因子,与OsFD1相互作用,影响Hd3a-14-3-3-OsFD1三蛋白FAC复合体的形成,延迟抽穗。 | [ |
| OsCOL4 | Os02g0610500 | CONSTANS转录因子,组成型的开花抑制因子,作用于Ehd1上游。 | [ |
| OsCOL15 | Os08g0536300 | CONSTANS转录因子,通过上调Ghd7和下调RID1来抑制抽穗。 | [ |
| HD6/CK2α | Os03g0762000 | 酪蛋白激酶CK2α亚基,通过调控Hd1基因,在长日照条件下延迟抽穗。 | [ |
| Hd16/CKI/EL1 | Os03g0793500 | 酪蛋白激酶CKI,长日条件下磷酸化Ghd7,增强其功能,延迟抽穗。 | [ |
| OsFTL12 | Os06g0552900 | FT家族蛋白,与GF14b和OsFD1相互作用,形成成花素抑制复合体FRC,调控OsMADS14和OsMADS15,延迟抽穗。 | [ |
| IDD4/SID1 | Os02g0672100 | IDD转录因子,作用于Hd3a和RFT1的启动子区域,启动水稻开花转变。 | [ |
| OsRIP1 | Os04g0540200 | OsRE1相互作用蛋白,在依赖OsRE1的方式下抑制Ehd1的转录表达调控抽穗期。 | [ |
| OsVIL1 | Os12g0533500 | 与OsVIL2互作,形成PRC2复合体,短日照条件下通过抑制OsLFL1表达诱导抽穗,而长日照条件下通过提高Ghd7表达延迟抽穗。 | [ |
| Ehd5 | Os08g0493900 | WD40结构域蛋白,与Roc4和Ghd8相互作用,影响Ghd7-Ghd8复合物的形成,促进开花相关基因的表达。 | [ |
| DTH7/Ghd7.1/ OsPRR37/Hd2 | Os07g0695100 | PRR蛋白,长日条件下,抑制Ehd1表达延迟抽穗;短日条件下,根据互作基因的不同,既可抑制也可促进抽穗。 | [ |
| SDG724/OsSET34 | Os09g0307800 | 组蛋白H3K36甲基转移酶,介导MADS50和RFT1的H3K36me2/3沉积,促进水稻开花。 | [ |
| Hd18 | Os08g0143400 | 组蛋白乙酰转移酶基因,通过提高Ehd1的转录水平促进抽穗。 | [ |
| OsSUF4 | Os09g0560900 | 锌指转录因子,促进H3K36me3对RFT1和Hd3a的修饰,从而促进抽穗。 | [ |
| OsHUB2/ FRRP1 | Os10g0565600 | E3泛素连接酶,对Ehd1的组蛋白H2B进行泛素化,抑制抽穗。 | [ |
| HAF1 | Os04g0648800 | E3泛素连接酶,作用于Hd1并将其降解,精细调控Hd1昼夜节律的积累。 | [ |
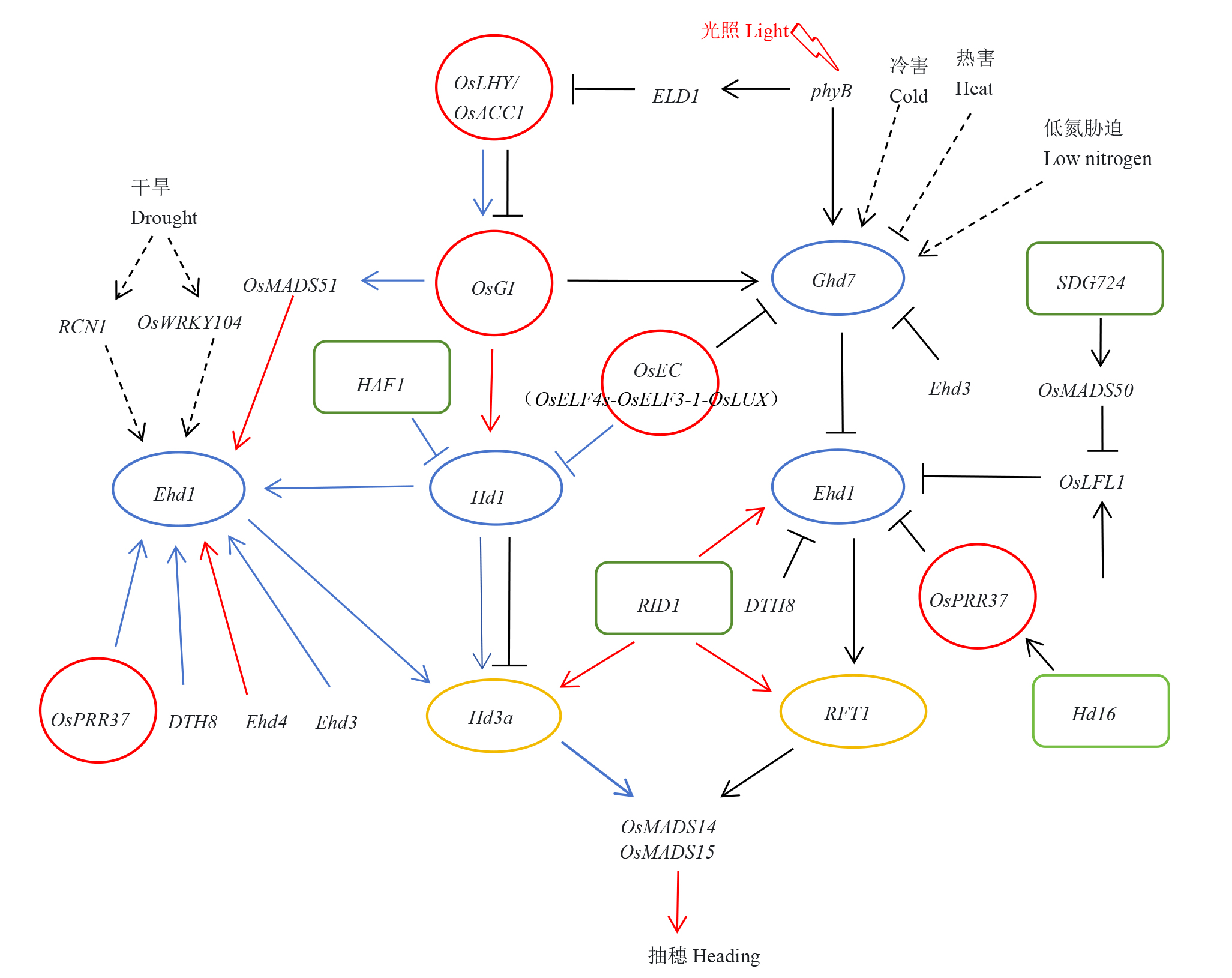
图1 水稻抽穗期调控基因网络 水稻抽穗开花主要受光周期调控,即短日照条件下的Hd1-Hd3a/RFT1途径和长日照条件下的Hd1/Ghd7/DTH8-Ehd1-Hd3a/RFT1途径。另外,温度、干旱、氮营养等环境因素也参与水稻抽穗期的调控。图中红色圆形表示生物钟相关基因,蓝色椭圆表示光周期调控网络的核心基因,黄色椭圆表示成花素基因,绿色圆角矩形表示表观遗传与翻译后修饰相关基因;实心箭头表示光周期调控途径中促进基因表达,实心T型表示光周期调控中抑制基因表达,其中蓝色表示短日照条件,黑色表示长日照条件,红色表示短日照和长日照两种条件;虚线箭头和虚线T型分别表示环境因子(温度、干旱、氮营养等)促进和抑制抽穗期调控基因的表达。
Fig. 1. Gene networks regulating heading date in rice Rice heading is primarily governed by photoperiod pathways: the Hd1-Hd3a/RFT1 module under short-day (SD) conditions and the Hd1/Ghd7/DTH8-Ehd1-Hd3a/RFT1 cascade under long-day (LD) conditions. Additionally, environmental factors such as temperature, drought, and nitrogen availability modulate reproductive transition. In the figure, red circles represent circadian clock-related genes; blue ellipses represent core genes of the photoperiod regulatory network; yellow ellipses represent florigen genes; green rounded rectangles represent genes related to epigenetics and post-translational modification; solid arrows indicate the activation of gene expression in photoperiod regulation; solid T-bars indicate the repression of gene expression in photoperiod regulation, where blue denotes SD conditions, black denotes LD conditions, and red denotes both SD and LD conditions; dashed arrows and dashed T-bars represent the activation and repression of gene expression by environmental factors (e.g., temperature, drought, nitrogen nutrition), respectively.
| [1] | Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS[J]. The Plant Cell, 2000, 12(12): 2473-2484. |
| [2] | Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1[J]. Genes & Development, 2004, 18(8): 926-936. |
| [3] | Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice[J]. Nature Genetics, 2008, 40(6): 761-767. |
| [4] | Sun K, Huang M, Zong W, Xiao D, Lei C, Luo Y, Song Y, Li S, Hao Y, Luo W, Xu B, Guo X, Wei G, Chen L, Liu Y G, Guo J. Hd1, Ghd7, and DTH8 synergistically determine the rice heading date and yield-related agronomic traits[J]. Journal of Genetics and Genomics, 2022, 49(5): 437-447. |
| [5] | Cao S, Luo X, Xu D, Tian X, Song J, Xia X, Chu C, He Z. Genetic architecture underlying light and temperature mediated flowering in Arabidopsis, rice, and temperate cereals[J]. The New Phytologist, 2021, 230(5): 1731-1745. |
| [6] | Lin X, Huang Y, Rao Y, Ouyang L, Zhou D, Zhu C, Fu J, Chen C, Yin J, Bian J, He H, Zou G, Xu J. A base substitution in OsphyC disturbs its Interaction with OsphyB and affects flowering time and chlorophyll synthesis in rice[J]. BMC Plant Biology, 2022, 22(1): 612. |
| [7] | Takano M, Inagaki N, Xie X, Kiyota S, Baba-Kasai A, Tanabata T, Shinomura T. Phytochromes are the sole photoreceptors for perceiving red/far-red light in rice[J]. Proceedings of the National Academy of Sciences of the United States of America, 2009, 106(34): 14705-14710. |
| [8] | Singh S, Vergish S, Jain N, Sharma A K, Khurana P, Khurana J P. OsCRY2 and OsFBO10 co-regulate photomorphogenesis and photoperiodic flowering in indica rice[J]. Plant Science, 2023, 330: 111631. |
| [9] | Li C, Liu X J, Yan Y, Alam M S, Liu Z, Yang Z K, Tao R F, Yue E K, Duan M H, Xu J H. OsLHY is involved in regulating flowering through the Hd1- and Ehd1- mediated pathways in rice (Oryza sativa L.)[J]. Plant Science, 2022, 315: 111145. |
| [10] | Lee S J, Kang K, Lim J H, Paek N C. Natural alleles of CIRCADIAN CLOCK ASSOCIATED1 contribute to rice cultivation by fine-tuning flowering time[J]. Plant Physiology, 2022, 190(1): 640-656. |
| [11] | Murakami M, Ashikari M, Miura K, Yamashino T, Mizuno T. The evolutionarily conserved OsPRR quintet: Rice pseudo-response regulators implicated in circadian rhythm[J]. Plant & Cell Physiology, 2003, 44(11): 1229-1236. |
| [12] | Lee Y S, An G. OsGI controls flowering time by modulating rhythmic flowering time regulators preferentially under short day in rice[J]. Journal of Plant Biology, 2015, 58(2): 137-145. |
| [13] | Xu P, Zhang Y, Wen X, Yang Q, Liu L, Hao S, Li J, Wu Z, Shah L, Sohail A, Liu Q, Sun L, Hong Y, Chen D, Shen X, Zhan X, Cheng S, Cao L, Wu W. The clock component OsLUX regulates rice heading through recruiting OsELF3-1 and OsELF4s to repress Hd1 and Ghd7[J]. Journal of Advanced Research, 2023, 48: 17-31. |
| [14] | Cai L, Hao B, Xu Z, Cui S, Wu Q, Lee J, Hou H, Hu Y, Zhu L, Wang J, Li W, Chang K, Shao W, Zhu S, Gan X, Li C, Jiang L, Tian Y, Liu X, Liu S, Chen L, Wang H, Zhou S, Wan J. ELD1 mediates photoperiodic flowering via OsCCA1 alternative splicing and interacts with phytochrome signaling in rice[J]. Nature Communications, 2025, 16(1): 5329. |
| [15] | Zong W, Ren D, Huang M, Sun K, Feng J, Zhao J, Xiao D, Xie W, Liu S, Zhang H, Qiu R, Tang W, Yang R, Chen H, Xie X, Chen L, Liu Y G, Guo J. Strong photoperiod sensitivity is controlled by cooperation and competition among Hd1, Ghd7 and DTH8 in rice heading[J]. The New Phytologist, 2021, 229(3): 1635-1649. |
| [16] | Park S J, Kim S L, Lee S, Je B I, Piao H L, Park S H, Kim C M, Ryu C H, Park S H, Xuan Y H, Colasanti J, An G, Han C D. OsId1) is necessary for the expression of Ehd1 (Early heading date 1) regardless of photoperiod[J]. The Plant Journal, 2008, 56(6): 1018-1029. |
| [17] | Gao H, Zheng X M, Fei G, Chen J, Jin M, Ren Y, Wu W, Zhou K, Sheng P, Zhou F, Jiang L, Wang J, Zhang X, Guo X, Wang J L, Cheng Z, Wu C, Wang H, Wan J M. Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice[J]. PLoS Genetics, 2013, 9(2): e1003281. |
| [18] | Kim S L, Lee S, Kim H J, Nam H G, An G. OsMADS51is a short-day flowering promoter that functions upstream of Ehd1, OsMADS14, and Hd3a[J]. Plant Physiology, 2007, 145(4): 1484-1494. |
| [19] | Tamaki S, Matsuo S, Wong H L, Yokoi S, Shimamoto K. Hd3a protein is a mobile flowering signal in rice[J]. Science, 2007, 316(5827): 1033-1036. |
| [20] | Komiya R, Ikegami A, Tamaki S, Yokoi S, Shimamoto K. Hd3a and RFT1 are essential for flowering in rice[J]. Development, 2008, 135(4): 767-774. |
| [21] | Komiya R, Yokoi S, Shimamoto K. A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice[J]. Development, 2009, 136(20): 3443-3450. |
| [22] | Taoka K I, Ohki I, Tsuji H, Furuita K, Hayashi K, Yanase T, Yamaguchi M, Nakashima C, Purwestri Y A, Tamaki S, Ogaki Y, Shimada C, Nakagawa A, Kojima C, Shimamoto K. 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen[J]. Nature, 2011, 476(7360): 332-335. |
| [23] | Pasriga R, Yoon J, Cho L H, An G. Overexpression of RICE FLOWERING LOCUS T 1 (RFT1) induces extremely early flowering in rice[J]. Molecules and Cells, 2019, 42(5): 406-417. |
| [24] | Peng Q, Zhu C, Liu T, Zhang S, Feng S, Wu C. Phosphorylation of OsFD1 by OsCIPK3 promotes the formation of RFT1-containing florigen activation complex for long-day flowering in rice[J]. Molecular Plant, 2021, 14(7): 1135-1148. |
| [25] | Song S, Chen Y, Liu L, Wang Y, Bao S, Zhou X, Teo Z W N, Mao C, Gan Y, Yu H. OsFTIP1-mediated regulation of florigen transport in rice is negatively regulated by the ubiquitin-like domain kinase OsUbDKγ4[J]. The Plant Cell, 2017, 29(3): 491-507. |
| [26] | Zheng R, Meng X, Hu Q, Yang B, Cui G, Li Y, Zhang S, Zhang Y, Ma X, Song X, Liang S, Li Y, Li J, Yu H, Luan W. OsFTL12, a member of FT-like family, modulates the heading date and plant architecture by florigen repression complex in rice[J]. Plant Biotechnology Journal, 2023, 21(7): 1343-1360. |
| [27] | Chai J, Zhu S, Li C, Wang C, Cai M, Zheng X, Zhou L, Zhang H, Sheng P, Wu M, Jin X, Cheng Z, Zhang X, Lei C, Ren Y, Lin Q, Zhou S, Guo X, Wang J, Zhao Z, Wan J. OsRE1 interacts with OsRIP1 to regulate rice heading date by finely modulating Ehd1 expression[J]. Plant Biotechnology Journal, 2021, 19(2): 300-310. |
| [28] | Zhang H, Zhu S, Liu T, Wang C, Cheng Z, Zhang X, Chen L, Sheng P, Cai M, Li C, Wang J, Zhang Z, Chai J, Zhou L, Lei C, Guo X, Wang J, Wang J, Jiang L, Wu C, Wan J. DELAYED HEADING DATE1 interacts with OsHAP5C/D, delays flowering time and enhances yield in rice[J]. Plant Biotechnology Journal, 2019, 17(2): 531-539. |
| [29] | Liu T, Zhang H, Zhou L, Zhang X, Zhou C, Li S, Cheng Z, Guo X, Zhu S, Wan J. DELAYED HEADING DATE3, encoding a heat shock transcription factor, delays flowering time and improves yield in rice (Oryza sativa L.)[J]. Agriculture, 2022, 12(7): 1022. |
| [30] | Cai M, Zhu S, Wu M, Zheng X, Wang J, Zhou L, Zheng T, Cui S, Zhou S, Li C, Zhang H, Chai J, Zhang X, Jin X, Cheng Z, Zhang X, Lei C, Ren Y, Lin Q, Guo X, Zhao L, Wang J, Zhao Z, Jiang L, Wang H, Wan J. DHD4, a CONSTANS-like family transcription factor, delays heading date by affecting the formation of the FAC complex in rice[J]. Molecular Plant, 2021, 14(2): 330-343. |
| [31] | Matsubara K, Yamanouchi U, Nonoue Y, Sugimoto K, Wang Z X, Minobe Y, Yano M. Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering[J]. The Plant Journal, 2011, 66(4): 603-612. |
| [32] | Ryu C H, Lee S, Cho L H, Kim S L, Lee Y S, Choi S C, Jeong H J, Yi J, Park S J, Han C D, An G. OsMADS50 and OsMADS56 function antagonistically in regulating long day (LD)-dependent flowering in rice[J]. Plant, Cell & Environment, 2009, 32(10): 1412-1427. |
| [33] | Han S H, Yoo S C, Lee B D, An G, Paek N C. Rice FLAVIN-BINDING, KELCH REPEAT, F-BOX 1 (OsFKF1) promotes flowering independent of photoperiod[J]. Plant, Cell & Environment, 2015, 38(12): 2527-2540. |
| [34] | Kim S K, Yun C H, Lee J H, Jang Y H, Park H Y, Kim J K. OsCO3, a CONSTANS-LIKE gene, controls flowering by negatively regulating the expression of FT-like genes under SD conditions in rice[J]. Planta, 2008, 228(2): 355-365. |
| [35] | Li D, Yang C, Li X, Gan Q, Zhao X, Zhu L. Functional characterization of rice OsDof12[J]. Planta, 2009, 229(6): 1159-1169. |
| [36] | Wu W, Zheng X M, Lu G, Zhong Z, Gao H, Chen L, Wu C, Wang H J, Wang Q, Zhou K, Wang J L, Wu F, Zhang X, Guo X, Cheng Z, Lei C, Lin Q, Jiang L, Wang H, Ge S, Wan J. Association of functional nucleotide polymorphisms at DTH2 with the northward expansion of rice cultivation in Asia[J]. Proceedings of the National Academy of Sciences of the United States of America, 2013, 110(8): 2775-2780. |
| [37] | Pan T, He M, Liu H, Tian X, Wang Z, Yu X, Miao X, Li X. Transcription factor bZIP65 delays flowering via suppressing Ehd1 expression in rice[J]. Molecular Breeding, 2022, 42(10): 63. |
| [38] | Li X, Tian X, He M, Liu X, Li Z, Tang J, Mei E, Xu M, Liu Y, Wang Z, Guan Q, Meng W, Fang J, Zhang J, Bu Q. bZIP71 delays flowering by suppressing Ehd1 expression in rice[J]. Journal of Integrative Plant Biology, 2022, 64(7): 1352-1363. |
| [39] | Jiang P, Wang S, Zheng H, Li H, Zhang F, Su Y, Xu Z, Lin H, Qian Q, Ding Y. SIP1 participates in regulation of flowering time in rice by recruiting OsTrx1 to Ehd1[J]. The New Phytologist, 2018, 219(1): 422-435. |
| [40] | Xuan H, Shi N, Chen J, Jiang Y, Zhang H, Chu C, Li S, Chen X, Yang H. Physical coupling of H3K4me3 demethylases and Polycomb repressive complex 2 to accelerate flowering in rice[J]. Plant Physiology, 2024, 195(3): 1802-1806. |
| [41] | Hori K, Ogiso-Tanaka E, Matsubara K, Yamanouchi U, Ebana K, Yano M. Hd16, a gene for casein kinase I, is involved in the control of rice flowering time by modulating the day-length response[J]. The Plant Journal, 2013, 76(1): 36-46. |
| [42] | Kwon C T, Koo B H, Kim D, Yoo S C, Paek N C. Casein kinases I and 2α phosphorylate Oryza sativa pseudo-response regulator 37 (OsPRR37) in photoperiodic flowering in rice[J]. Molecules and Cells, 2015, 38(1): 81-88. |
| [43] | Luan W, Chen H, Fu Y, Si H, Peng W, Song S, Liu W, Hu G, Sun Z, Xie D, Sun C. The effect of the crosstalk between photoperiod and temperature on the heading-date in rice[J]. PLoS One, 2009, 4(6): e5891. |
| [44] | Song Y, Gao Z, Luan W. Interaction between temperature and photoperiod in regulation of flowering time in rice[J]. Science China Life Sciences, 2012, 55(3): 241-249. |
| [45] | Weng X, Wang L, Wang J, Hu Y, Du H, Xu C, Xing Y, Li X, Xiao J, Zhang Q. Grain number, plant height, and heading date 7 is a central regulator of growth, development, and stress response[J]. Plant Physiology, 2014, 164(2): 735-747. |
| [46] | Guo T, Mu Q, Wang J, Vanous A E, Onogi A, Iwata H, Li X, Yu J. Dynamic effects of interacting genes underlying rice flowering-time phenotypic plasticity and global adaptation[J]. Genome Research, 2020, 30(5): 673-683. |
| [47] | Zhang C, Liu J, Zhao T, Gomez A, Li C, Yu C, Li H, Lin J, Yang Y, Liu B, Lin C. A drought-inducible transcription factor delays reproductive timing in rice[J]. Plant Physiology, 2016, 171(1): 334-343. |
| [48] | Wang Y, Lu Y, Guo Z, Ding Y, Ding C. RICE CENTRORADIALIS 1, a TFL1-like gene, responses to drought stress and regulates rice flowering transition[J]. Rice (N Y), 2020, 13(1): 70. |
| [49] | Castro Marín I, Loef I, Bartetzko L, Searle I, Coupland G, Stitt M, Osuna D. Nitrate regulates floral induction in Arabidopsis, acting independently of light, gibberellin and autonomous pathways[J]. Planta, 2011, 233(3): 539-552. |
| [50] | Ye T, Li Y, Zhang J, Hou W, Zhou W, Lu J, Xing Y, Li X. Nitrogen, phosphorus, and potassium fertilization affects the flowering time of rice (Oryza sativa L.)[J]. Global Ecology and Conservation, 2019, 20: e00753. |
| [51] | Zhang S, Zhang Y, Li K, Yan M, Zhang J, Yu M, Tang S, Wang L, Qu H, Luo L, Xuan W, Xu G. Nitrogen mediates flowering time and nitrogen use efficiency via floral regulators in rice[J]. Current Biology, 2021, 31(4): 671-683.e5. |
| [52] | Wang Q, Su Q, Nian J, Zhang J, Guo M, Dong G, Hu J, Wang R, Wei C, Li G, Wang W, Guo H S, Lin S, Qian W, Xie X, Qian Q, Chen F, Zuo J. The Ghd7 transcription factor represses ARE1 expression to enhance nitrogen utilization and grain yield in rice[J]. Molecular Plant, 2021, 14(6): 1012-1023. |
| [53] | Liang L, Zhang Z, Cheng N, Liu H, Song S, Hu Y, Zhou X, Zhang J, Xing Y. The transcriptional repressor OsPRR73 links circadian clock and photoperiod pathway to control heading date in rice[J]. Plant, Cell & Environment, 2021, 44(3): 842-855. |
| [54] | Sun C, Zhang K, Zhou Y, Xiang L, He C, Zhong C, Li K, Wang Q, Yang C, Wang Q, Chen C, Chen D, Wang Y, Liu C, Yang B, Wu H, Chen X, Li W, Wang J, Xu P, Wang P, Fang J, Chu C, Deng X. Dual function of clock component OsLHY sets critical day length for photoperiodic flowering in rice[J]. Plant Biotechnology Journal, 2021, 19(8): 1644-1657. |
| [55] | Matsubara K, Ogiso-Tanaka E, Hori K, Ebana K, Ando T, Yano M. Natural variation in Hd17, a homolog of Arabidopsis ELF3 that is involved in rice photoperiodic flowering[J]. Plant & Cell Physiology, 2012, 53(4): 709-716. |
| [56] | Wang X, He Y, Wei H, Wang L. A clock regulatory module is required for salt tolerance and control of heading date in rice[J]. Plant, Cell & Environment, 2021, 44(10): 3283-3301. |
| [57] | Zhang L, Zhang F, Zhou X, Poh T X, Xie L, Shen J, Yang L, Song S, Yu H, Chen Y. The tetratricopeptide repeat protein OsTPR075 promotes heading by regulating florigen transport in rice[J]. The Plant Cell, 2022, 34(10): 3632-3646. |
| [58] | Lee S, Kim J, Han J J, Han M J, An G. Functional analyses of the flowering time gene OsMADS50, the putative SUPPRESSOR OF OVEREXPRESSION OF CO 1/AGAMOUS-LIKE 20 (SOC1/AGL20) ortholog in rice[J]. The Plant Journal, 2004, 38(5): 754-764. |
| [59] | Li J, Qiu J X, Zeng Q H, Zhuang Y, Zhang N, Xu S X, Jin J, Dong Z C, Chen L, Huang W. OsTOC1 plays dual roles in the regulation of plant circadian clock by functioning as a direct transcription activator or repressor[J]. Cell Reports, 2023, 42(7): 112765. |
| [60] | Cho LH, Yoon J, Pasriga R, An G. Homodimerization of Ehd1 is required to induce flowering in rice[J]. Plant Physiology, 2016, 170(4): 2159-2171. |
| [61] | Yin Y, Yan Z, Guan J, Huo Y, Wang T, Li T, Cui Z, Ma W, Wang X, Chen W. Two interacting basic helix-loop-helix transcription factors control flowering time in rice[J]. Plant Physiology, 2023, 192(1): 205-221. |
| [62] | Cerise M, Giaume F, Galli M, Khahani B, Lucas J, Podico F, Tavakol E, Parcy F, Gallavotti A, Brambilla V, Fornara F. OsFD4 promotes the rice floral transition via florigen activation complex formation in the shoot apical meristem[J]. The New Phytologist, 2021, 229(1): 429-443. |
| [63] | Brambilla V, Martignago D, Goretti D, Cerise M, Somssich M, de Rosa M, Galbiati F, Shrestha R, Lazzaro F, Simon R, Fornara F. Antagonistic transcription factor complexes modulate the floral transition in rice[J]. The Plant Cell, 2017, 29(11): 2801-2816. |
| [64] | Matsubara K, Yamanouchi U, Wang Z X, Minobe Y, Izawa T, Yano M. Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1[J]. Plant Physiology, 2008, 148(3): 1425-1435. |
| [65] | Fan X, Wang P, Qi F, Hu Y, Li S, Zhang J, Liang L, Zhang Z, Liu J, Xiong L, Xing Y. The CCT transcriptional activator Ghd2 constantly delays the heading date by upregulating CO3 in rice[J]. Journal of Genetics and Genomics, 2023, 50(10): 755-764 |
| [66] | Lee Y S, Jeong D H, Lee D Y, Yi J, Ryu C H, Kim S L, Jeong H J, Choi S C, Jin P, Yang J, Cho L H, Choi H, An G. OsCOL4 is a constitutive flowering repressor upstream of Ehd1 and downstream of OsphyB[J]. The Plant Journal, 2010, 63(1): 18-30. |
| [67] | Wu W, Zhang Y, Zhang M, Zhan X, Shen X, Yu P, Chen D, Liu Q, Sinumporn S, Hussain K, Cheng S, Cao L. The rice CONSTANS-like protein OsCOL15 suppresses flowering by promoting Ghd7 and repressing RID1[J]. Biochemical and Biophysical Research Communications, 2018, 495(1): 1349-1355. |
| [68] | Ogiso E, Takahashi Y, Sasaki T, Yano M, Izawa T. The role of casein kinase II in flowering time regulation has diversified during evolution[J]. Plant Physiology, 2010, 152(2): 808-820. |
| [69] | Deng L, Li L, Zhang S, Shen J, Li S, Hu S, Peng Q, Xiao J, Wu C. Suppressor of rid1 (SID1) shares common targets with RID1 on florigen genes to initiate floral transition in rice[J]. PLoS Genetics, 2017, 13(2): 1006642. |
| [70] | Jeong H J, Yang J, Cho L H, An G. OsVIL1 controls flowering time in rice by suppressing OsLF under short days and by inducing Ghd7 under long days[J]. Plant Cell Reports, 2016, 35(4): 905-920. |
| [71] | Zhang X, Feng Q, Miao J, Zhu J, Zhou C, Fan D, Lu Y, Tian Q, Wang Y, Zhan Q, Wang Z Q, Wang A, Zhang L, Shangguan Y, Li W, Chen J, Weng Q, Huang T, Tang S, Si L, Huang X, Wang Z X, Han B. The WD40 domain-containing protein Ehd5 positively regulates flowering in rice (Oryza sativa)[J]. The Plant Cell, 2023, 35(11): 4002-4019. |
| [72] | Zhang B, Liu H, Qi F, Zhang Z, Li Q, Han Z, Xing Y. Genetic interactions among Ghd7, Ghd8, OsPRR37 and Hd1 contribute to large variation in heading date in rice[J]. Rice (N Y), 2019, 12(1): 48. |
| [73] | Sun C, Fang J, Zhao T, Xu B, Zhang F, Liu L, Tang J, Zhang G, Deng X, Chen F, Qian Q, Cao X, Chu C. The histone methyltransferase SDG724 mediates H3K36me2/3 deposition at MADS50 and RFT1and promotes flowering in rice[J]. The Plant Cell, 2012, 24(8): 3235-3247. |
| [74] | Shibaya T, Hori K, Ogiso-Tanaka E, Yamanouchi U, Shu K, Kitazawa N, Shomura A, Ando T, Ebana K, Wu J, Yamazaki T, Yano M. Hd18, encoding histone acetylase related to Arabidopsis FLOWERING LOCUS D, is involved in the control of flowering time in rice[J]. Plant & Cell Physiology, 2016, 57(9): 1828-1838. |
| [75] | Liu B, Liu Y, Wang B, Luo Q, Shi J, Gan J, Shen W H, Yu Y, Dong A. The transcription factor OsSUF4 interacts with SDG725 in promoting H3K36me3 establishment[J]. Nature Communications, 2019, 10(1): 2999. |
| [76] | Xu Z, Li E, Xue G, Zhang C, Yang Y, Ding Y. OsHUB2 inhibits function of OsTrx1 in heading date in rice[J]. The Plant Journal, 2022, 110(6): 1670-1680. |
| [77] | Yang Y, Fu D, Zhu C, He Y, Zhang H, Liu T, Li X, Wu C. The RING-finger ubiquitin ligase HAF1 mediates Heading date 1 degradation during photoperiodic flowering in rice[J]. The Plant Cell, 2015, 27(9): 2455-2468. |
| [78] | Zhang J, Zhou X, Yan W, Zhang Z, Lu L, Han Z, Zhao H, Liu H, Song P, Hu Y, Shen G, He Q, Guo S, Gao G, Wang G, Xing Y. Combinations of the Ghd7, Ghd8 and Hd1 genes largely define the ecogeographical adaptation and yield potential of cultivated rice[J]. The New Phytologist, 2015, 208(4): 1056-1066. |
| [79] | Zhou X, Nong C, Wu B, Zhou T, Zhang B, Liu X, Gao G, Mi J, Zhang Q, Liu H, Liu S, Li Z, He Y, Mou T, Guo S, Li S, Yang Y, Zhang Q, Xing Y. Combinations of Ghd7, Ghd8, and Hd1 determine strong heterosis of commercial rice hybrids in diverse ecological regions[J]. Journal of Experimental Botany, 2021, 72(20): 6963-6976. |
| [80] | Koo B H, Yoo S C, Park J W, Kwon C T, Lee B D, An G, Zhang Z, Li J, Li Z, Paek N C. Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes[J]. Molecular Plant, 2013, 6(6): 1877-1888. |
| [81] | Su Q, Wang R, Feng X, Zhao M, Zhu G, Wang Q, Zhang F, Lin S, Zhang Y, Zhu L, Qian Q, Chen F. A new strategy of molecular breeding for optimal heading date and grain yield in rice by modulating elite allelic combinations of Ghd7, Hd3a, RFT1 and Gn1a[J]. The Crop Journal, 2025, 13(2): 490-500. |
| [82] | Sun K, Zong W, Xiao D, Wu Z, Guo X, Li F, Song Y, Li S, Wei G, Hao Y, Xu B, Li W, Lin Z, Xie W, Liu Y G, Guo J. Effects of the core heading date genes Hd1, Ghd7, DTH8 and PRR37 on yield-related traits in rice[J]. Theoretical and Applied Genetics, 2023, 136(11): 227. |
| [83] | Li X, Liu H, Wang M, Liu H, Tian X, Zhou W, Lü T, Wang Z, Chu C, Fang J, Bu Q. Combinations of Hd2 and Hd4 genes determine rice adaptability to Heilongjiang Province, northern limit of China[J]. Journal of Integrative Plant Biology, 2015, 57(8): 698-707. |
| [84] | Wang X, Zhou T, Li G, Yao W, Hu W, Wei X, Che J, Yang H, Shao L, Hua J, Li X, Xiao J, Xing Y, Ouyang Y, Zhang Q. A Ghd7-centered regulatory network provides a mechanistic approximation to optimal heterosis in an elite rice hybrid[J]. The Plant Journal, 2022, 112(1): 68-83. |
| [85] | Liu J, Yi Q, Dong G, Chen Y, Guo L, Gao Z, Zhu L, Ren D, Zhang Q, Li Q, Li J, Liu Q, Zhang G, Qian Q, Shen L. Improving rice quality by regulating the heading dates of rice varieties without yield penalties[J]. Plants (Basel), 2024, 13(16): 2221. |
| [86] | Leng Y, Gao Y, Chen L, Yang Y, Huang L, Dai L, Ren D, Xu Q, Zhang Y, Ponce K, Hu J, Shen L, Zhang G, Chen G, Dong G, Gao Z, Guo L, Ye G, Qian Q, Zhu L, Zeng D. Using Heading date 1 preponderant alleles from indica cultivars to breed high-yield, high-quality japonica rice varieties for cultivation in South China[J]. Plant Biotechnology Journal, 2020, 18(1): 119-128. |
| [87] | Guo X, Sun K, Wu Z, Xiao D, Song Y, Li S, Wei G, Li W, Hao Y, Xu B, Zhang K, Liao N, Hu D, Liu Y G, Zong W, Guo J. Improving yield-related traits by editing the promoter and distal regulatory region of heading date genes Ghd7 and PRR37 in elite rice variety Mei Xiang Zhan 2[J]. Theoretical and Applied Genetics, 2025, 138(4): 92. |
| [88] | Zhou S, Cai L, Wu H, Wang B, Gu B, Cui S, Huang X, Xu Z, Hao B, Hou H, Hu Y, Li C, Tian Y, Liu X, Chen L, Liu S, Jiang L, Wan J. Fine-tuning rice heading date through multiplex editing of the regulatory regions of key genes by CRISPR-Cas9[J]. Plant Biotechnology Journal, 2024, 22(3): 751-758. |
| [1] | 薛炮, 王友霜, 何弯弯, 黄晨博, 张涵, 丁震乾, 陈秋丽, 范运新, 丁成伟, 孙廉平, 胡婷婷. 水稻颖壳不闭合基因SG5的鉴定与克隆[J]. 中国水稻科学, 2026, 40(2): 210-222. |
| [2] | 倪晨, 张家豪, 朱昌进, 徐继伟, 胡秋倩, 霍中洋, 戴其根, 许轲, 李国辉. 水稻源流库形成与调控及其影响因素研究进展[J]. 中国水稻科学, 2026, 40(2): 155-170. |
| [3] | 王梦宁, 谢可冉, 高逖, 王真梅, 熊栋梁, 崔克辉. 高温对水稻粒重形成的影响及其栽培调控研究进展[J]. 中国水稻科学, 2026, 40(2): 171-180. |
| [4] | 罗肖郧, 郑兴飞, 彭宣国, 余启芝, 董华林, 殷得所, 王红波, 胡建林, 薛莲, 胡鹏, 徐得泽. 水稻抗倒伏研究:现状、挑战与未来方向[J]. 中国水稻科学, 2026, 40(2): 181-195. |
| [5] | 段敏, 谢留杰, 岳雅妮, 黄善军. 基于CRISPR/Cas9技术创制优质香味粳稻品系[J]. 中国水稻科学, 2026, 40(2): 235-243. |
| [6] | 张梦柯, 陆佳雨, 何金, 许学, 吴爽, 王沛然, 陈若凡, 金青, 汪秀峰. 水稻野败型三系杂交种纯度检测功能标记的开发与应用[J]. 中国水稻科学, 2026, 40(2): 244-252. |
| [7] | 李兴沂, 陈玲, 邵建韬, 肖素勤, 李金璐, 付惠仙, 殷富有, 张建红, 程在全, 刘丽. 水稻产量与淀粉品质协同调控的分子遗传研究进展[J]. 中国水稻科学, 2026, 40(1): 1-17. |
| [8] | 苗哲宁, 陈积金, 李自明, 刘毅, 罗利军. 华南稻区抗白叶枯病育种研究现状与展望[J]. 中国水稻科学, 2026, 40(1): 18-26. |
| [9] | 岳轩宇, 谢文亚, 冯志明, 陈宗祥, 胡珂鸣, 左示敏. OsERF93参与调控水稻纹枯病抗性的研究[J]. 中国水稻科学, 2026, 40(1): 37-50. |
| [10] | 王轶欣, 林参, 马刘洋, 陈龙, 奉保华, 倪深, 魏祥进, 贺记外, 陈天晓. 谷丙转氨酶基因OsAlaAT4调控水稻氮素吸收和产量[J]. 中国水稻科学, 2026, 40(1): 51-60. |
| [11] | 黄奇娜, 姜鸿瑞, 杨婕, 于坤宇, 杨长登, 梁燕. 种子休眠基因Sdr4的生物信息学分析与分子标记开发和应用[J]. 中国水稻科学, 2026, 40(1): 61-71. |
| [12] | 程朝平, 何旎清, 白康呈, 林少俊, 黄凤凰, 刘军化, 程祖锌, 黄成志, 杨德卫. 聚合稻瘟病抗性基因Pigm-1和Pid2的水稻三系不育系福梦A的选育与利用[J]. 中国水稻科学, 2026, 40(1): 72-84. |
| [13] | 刘亚萍, 董译词, 郑君妍, 邱绚, 刘鹏程, 叶亚峰, 刘斌美, 陈析丰, 马伯军. 水稻类病变早衰突变体lmes7的鉴定与基因精细定位[J]. 中国水稻科学, 2026, 40(1): 85-94. |
| [14] | 谢世民, 周誉株, 薛晓迪, 朱广飞, 孙良, 陈建能. 水稻钵苗取栽协同作业式移栽机构设计与试验[J]. 中国水稻科学, 2026, 40(1): 131-144. |
| [15] | 宋安琪, 吴松权, 马秋月, 班宛宁, 刘相国, 金永梅. 植物引导编辑技术——作物育种的新方向[J]. 中国水稻科学, 2025, 39(6): 711-730. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||