
中国水稻科学 ›› 2020, Vol. 34 ›› Issue (1): 17-27.DOI: 10.16819/j.1001-7216.2020.9052
王小雷, 刘杨, 孙晓棠, 欧阳林娟, 潘锦龙, 彭小松, 陈小荣, 贺晓鹏, 傅军如, 边建民, 胡丽芳, 徐杰, 贺浩华*( ), 朱昌兰*(
), 朱昌兰*( )
)
收稿日期:2019-04-30
修回日期:2019-06-13
出版日期:2020-01-10
发布日期:2020-01-10
通讯作者:
贺浩华,朱昌兰
作者简介:作者简介:#共同第一作者
基金资助:
Xiaolei WANG, Yang LIU, Xiaotang SUN, Linjuan OUYANG, Jinlong PAN, Xiaosong PENG, Xiaorong CHEN, Xiaopeng HE, Junru FU, Jianmin BIAN, Lifang HU, Jie XU, Haohua HE*( ), Changlan ZHU*(
), Changlan ZHU*( )
)
Received:2019-04-30
Revised:2019-06-13
Online:2020-01-10
Published:2020-01-10
Contact:
Haohua HE, Changlan ZHU
About author:About author:#These authors contributed equally to this work
摘要:
【目的】挖掘新的稻米品质性状QTL并利用分子育种技术改良稻米品质。【方法】利用构建的一套以籼稻香型恢复系昌恢121为背景亲本,以优质粳稻越光为供体亲本的染色体片段代换系(CSSLs)为材料,在4个环境下对稻米品质性状进行QTL检测及稳定性分析。【结果】在4个环境下共检测到44个QTL,其中6个QTL能在多个环境下被检测到;第2、3、5、6和10染色体上存在多效QTL簇,对稻米品质性状具有明显的调控作用;第1、6和12染色体上7个QTL能在不同环境下稳定表达。【结论】第1染色体RM3143-RM1117区间qPGWC1和第12染色体RM3331-RM5479区间qPaT12是两个新的稳定表达QTL。
中图分类号:
王小雷, 刘杨, 孙晓棠, 欧阳林娟, 潘锦龙, 彭小松, 陈小荣, 贺晓鹏, 傅军如, 边建民, 胡丽芳, 徐杰, 贺浩华, 朱昌兰. 不同环境下稻米品质性状QTL的检测及稳定性分析[J]. 中国水稻科学, 2020, 34(1): 17-27.
Xiaolei WANG, Yang LIU, Xiaotang SUN, Linjuan OUYANG, Jinlong PAN, Xiaosong PENG, Xiaorong CHEN, Xiaopeng HE, Junru FU, Jianmin BIAN, Lifang HU, Jie XU, Haohua HE, Changlan ZHU. Identification and Stability Analysis of QTL for Grain Quality Traits Under Multiple Environments in Rice[J]. Chinese Journal OF Rice Science, 2020, 34(1): 17-27.
| 性状 Trait | 环境 E | 亲本 Parent | CSSL群体 CSSL population | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 昌恢121 Changhui 121 | 越光 Koshihikari | 变异范围 Range | 平均±标准差 Mean±SD | 峰度 Kurtosis | 偏度 Skewness | ||||||||
| 垩白粒率PGWC/% | E1 | 4.5±0.5 | 5.0±0.0 | 4.0~42.0 | 14.9±9.6 | 1.4 | 1.4 | ||||||
| E2 | 2.5±0.5 | 3.0±1.0 | 0.5~49.0 | 6.2±8.1 | 13.8 | 3.5 | |||||||
| E3 | 3.5±1.5 | 33.0±2.0 | 2.0~56.0 | 16.0±9.8 | 3.4 | 1.3 | |||||||
| E4 | 7.0±0.5 | 6.0±1.0 | 4.0~41.0 | 15.8±7.8 | 1.2 | 1.1 | |||||||
| 胶稠度 GC/mm | E1 | 30.0±2.0 | 51.5±0.5 | 24.5~66.0 | 36.0±9.8 | 2.4 | 1.7 | ||||||
| E2 | 30.5±0.5 | 56.5±0.5 | 30.5~92.5 | 43.5±15.9 | 2.4 | 1.7 | |||||||
| E3 | 55.5±1.5 | 82.5±1.5 | 34.0~87.5 | 57.4±13.3 | -0.7 | 0.3 | |||||||
| E4 | 32.5±1.5 | 67.5±1.5 | 31.5~100.0 | 59.3±19.0 | -0.6 | 0.4 | |||||||
| 蛋白质含量PC/% | E1 | 8.6±0.2 | 8.11±0.1 | 5.9~10.6 | 7.7±0.8 | 1.6 | 0.8 | ||||||
| E2 | 8.9±0.1 | 8.98±0.1 | 6.4~10.1 | 8.5±0.8 | 0.8 | 0.7 | |||||||
| E3 | 10.2±0.0 | 9.04±0.0 | 70~11.3 | 8.6±1.0 | 0.1 | 0.6 | |||||||
| E4 | 7.1±0.0 | 7.44±0.0 | 5.6~7.9 | 6.5±0.5 | 0.2 | 0.7 | |||||||
| 最高黏度PKV/cP | E1 | 4936.0±91.0 | 5718.0±29.0 | 3850.5~5922.0 | 4962.1±434.3 | 0.6 | -0.3 | ||||||
| E2 | 4972.0±24.0 | 4557.5±77.0 | 3193.5~6027.0 | 4387.6±536.1 | 0.7 | 0.7 | |||||||
| E3 | 3900.0±22.0 | 4448.0±73.0 | 2567.0~4949.5 | 4152.1±336.9 | 8.0 | -1.6 | |||||||
| E4 | 4878.0±82.0 | 4944.5±48.5 | 4015.0~5194.5 | 4462.5±228.8 | 0.7 | 0.6 | |||||||
| 热浆黏度HPV/cP | E1 | 3412.5±100.5 | 2321.5±56.5 | 2163.5~4098.5 | 3298.6±456.0 | 0.3 | -1.1 | ||||||
| E2 | 4136.5±54.5 | 2965.5±50.5 | 2637.5~4645.0 | 3485.9±431.3 | 0.1 | 0.2 | |||||||
| E3 | 2784.0±86.0 | 1948.0±52.0 | 1592.5~3709.5 | 2807.3±428.7 | 1.6 | -1.2 | |||||||
| E4 | 3404.5±48.5 | 2818.5±55.5 | 2029.0~3589.0 | 3097.8±340.4 | 2.2 | -1.5 | |||||||
| 崩解值BDV/cP | E1 | 1523.5±103.5 | 3396.5±98.5 | 1037.0~3713.5 | 1663.5±620.4 | 3.5 | 2.1 | ||||||
| E2 | 835.5±30.5 | 1592.0±27.0 | 298.0~3000.0 | 901.7±490.8 | 5.5 | 2.2 | |||||||
| E3 | 1116.0±60.7 | 2500.0±17.0 | 700.5~2556.0 | 1344.8±354.2 | 2.7 | 1.4 | |||||||
| E4 | 1473.5±75.5 | 2126.0±81.0 | 903.0~3165.5 | 1364.8±436.2 | 6.2 | 2.4 | |||||||
| 冷浆黏度CPV/cP | E1 | 6664.0±34.5 | 3723.5±18.5 | 3351.0~7426.0 | 6008.0±1028.7 | 1.4 | -1.5 | ||||||
| E2 | 7332.5±4.5 | 4775.5±40.5 | 3963.0~8000.0 | 6459.8±951.6 | 0.4 | -1.0 | |||||||
| E3 | 5643.5±47.5 | 2901.5±6.5 | 2790.5~7454.5 | 5350.3±870.2 | 2.7 | -1.5 | |||||||
| E4 | 6222.0±35.0 | 4224.5±37.5 | 3254.0~6477.5 | 5641.7±773.1 | 3.1 | -2.0 | |||||||
| 消减值SBV/cP | E1 | 1728.0±36.0 | -1994.5±97.5 | -2505.0~2160.5 | 1045.9±1177.2 | 3.1 | -2.1 | ||||||
| E2 | 2360.5±28.5 | 218.0±37.0 | -1765.0~3005.5 | 2072.3±982.7 | 4.6 | -2.3 | |||||||
| E3 | 1743.5±31.5 | -1546.5±12.5 | -1434.5~2505.0 | 1198.3±760.4 | 3.9 | -2.1 | |||||||
| E4 | 1344.0±42.0 | -720.0±31.0 | -1940.5~2189.5 | 1179.2±865.0 | 4.6 | -2.3 | |||||||
| 回复值CSV/cP | E1 | 3251.5±125.5 | 1402.0±20.0 | 1187.5~3546.5 | 2709.4±602.8 | 1.9 | -1.7 | ||||||
| E2 | 3196.0±59.0 | 1810.0±10.0 | 1235.0~3757.5 | 2973.9±584.8 | 1.5 | -1.5 | |||||||
| E3 | 2859.5±27.5 | 953.5±5.5 | 1121.5~3745.0 | 2543.1±545.3 | 1.0 | -1.1 | |||||||
| E4 | 2817.5±55.5 | 1406.0±28.0 | 1138.5~3460.0 | 2544.0±520.0 | 1.2 | -1.2 | |||||||
| 成糊温度PaT/℃ | E1 | 72.4±0.3 | 74.6±0.7 | 68.1~76.0 | 71.8±1.5 | 1.3 | 0.3 | ||||||
| E2 | 78.4±0.8 | 80.7±0.1 | 74.1~87.0 | 81.1±3.3 | -0.6 | -0.6 | |||||||
| E3 | 70.6±0.1 | 76.8±0.3 | 70.0~76.4 | 72.1±1.2 | 1.2 | 0.9 | |||||||
| E4 | 72.1±0.1 | 73.7±0.3 | 70.9~74.8 | 72.1±1.5 | 1.6 | 1.1 | |||||||
| 成糊时间PeT/min | E1 | 5.9±0.1 | 5.7±0.1 | 5.5~6.2 | 5.9±0.1 | 1.4 | -1.0 | ||||||
| E2 | 6.1±0.0 | 6.0±0.0 | 5.7~6.4 | 6.0±0.1 | 0.4 | 0.2 | |||||||
| E3 | 6.0±0.1 | 5.7±0.3 | 5.7~6.7 | 6.0±0.2 | 0.3 | 0.6 | |||||||
| E4 | 6.0±0.2 | 6.2±0.1 | 5.6~6.4 | 6.1±0.2 | -0.2 | -0.2 | |||||||
表1 亲本及CSSLs群体在4个环境中的稻米品质性状
Table 1 Phenotypic performance of CSSLs population and its parents in 4 environments.
| 性状 Trait | 环境 E | 亲本 Parent | CSSL群体 CSSL population | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 昌恢121 Changhui 121 | 越光 Koshihikari | 变异范围 Range | 平均±标准差 Mean±SD | 峰度 Kurtosis | 偏度 Skewness | ||||||||
| 垩白粒率PGWC/% | E1 | 4.5±0.5 | 5.0±0.0 | 4.0~42.0 | 14.9±9.6 | 1.4 | 1.4 | ||||||
| E2 | 2.5±0.5 | 3.0±1.0 | 0.5~49.0 | 6.2±8.1 | 13.8 | 3.5 | |||||||
| E3 | 3.5±1.5 | 33.0±2.0 | 2.0~56.0 | 16.0±9.8 | 3.4 | 1.3 | |||||||
| E4 | 7.0±0.5 | 6.0±1.0 | 4.0~41.0 | 15.8±7.8 | 1.2 | 1.1 | |||||||
| 胶稠度 GC/mm | E1 | 30.0±2.0 | 51.5±0.5 | 24.5~66.0 | 36.0±9.8 | 2.4 | 1.7 | ||||||
| E2 | 30.5±0.5 | 56.5±0.5 | 30.5~92.5 | 43.5±15.9 | 2.4 | 1.7 | |||||||
| E3 | 55.5±1.5 | 82.5±1.5 | 34.0~87.5 | 57.4±13.3 | -0.7 | 0.3 | |||||||
| E4 | 32.5±1.5 | 67.5±1.5 | 31.5~100.0 | 59.3±19.0 | -0.6 | 0.4 | |||||||
| 蛋白质含量PC/% | E1 | 8.6±0.2 | 8.11±0.1 | 5.9~10.6 | 7.7±0.8 | 1.6 | 0.8 | ||||||
| E2 | 8.9±0.1 | 8.98±0.1 | 6.4~10.1 | 8.5±0.8 | 0.8 | 0.7 | |||||||
| E3 | 10.2±0.0 | 9.04±0.0 | 70~11.3 | 8.6±1.0 | 0.1 | 0.6 | |||||||
| E4 | 7.1±0.0 | 7.44±0.0 | 5.6~7.9 | 6.5±0.5 | 0.2 | 0.7 | |||||||
| 最高黏度PKV/cP | E1 | 4936.0±91.0 | 5718.0±29.0 | 3850.5~5922.0 | 4962.1±434.3 | 0.6 | -0.3 | ||||||
| E2 | 4972.0±24.0 | 4557.5±77.0 | 3193.5~6027.0 | 4387.6±536.1 | 0.7 | 0.7 | |||||||
| E3 | 3900.0±22.0 | 4448.0±73.0 | 2567.0~4949.5 | 4152.1±336.9 | 8.0 | -1.6 | |||||||
| E4 | 4878.0±82.0 | 4944.5±48.5 | 4015.0~5194.5 | 4462.5±228.8 | 0.7 | 0.6 | |||||||
| 热浆黏度HPV/cP | E1 | 3412.5±100.5 | 2321.5±56.5 | 2163.5~4098.5 | 3298.6±456.0 | 0.3 | -1.1 | ||||||
| E2 | 4136.5±54.5 | 2965.5±50.5 | 2637.5~4645.0 | 3485.9±431.3 | 0.1 | 0.2 | |||||||
| E3 | 2784.0±86.0 | 1948.0±52.0 | 1592.5~3709.5 | 2807.3±428.7 | 1.6 | -1.2 | |||||||
| E4 | 3404.5±48.5 | 2818.5±55.5 | 2029.0~3589.0 | 3097.8±340.4 | 2.2 | -1.5 | |||||||
| 崩解值BDV/cP | E1 | 1523.5±103.5 | 3396.5±98.5 | 1037.0~3713.5 | 1663.5±620.4 | 3.5 | 2.1 | ||||||
| E2 | 835.5±30.5 | 1592.0±27.0 | 298.0~3000.0 | 901.7±490.8 | 5.5 | 2.2 | |||||||
| E3 | 1116.0±60.7 | 2500.0±17.0 | 700.5~2556.0 | 1344.8±354.2 | 2.7 | 1.4 | |||||||
| E4 | 1473.5±75.5 | 2126.0±81.0 | 903.0~3165.5 | 1364.8±436.2 | 6.2 | 2.4 | |||||||
| 冷浆黏度CPV/cP | E1 | 6664.0±34.5 | 3723.5±18.5 | 3351.0~7426.0 | 6008.0±1028.7 | 1.4 | -1.5 | ||||||
| E2 | 7332.5±4.5 | 4775.5±40.5 | 3963.0~8000.0 | 6459.8±951.6 | 0.4 | -1.0 | |||||||
| E3 | 5643.5±47.5 | 2901.5±6.5 | 2790.5~7454.5 | 5350.3±870.2 | 2.7 | -1.5 | |||||||
| E4 | 6222.0±35.0 | 4224.5±37.5 | 3254.0~6477.5 | 5641.7±773.1 | 3.1 | -2.0 | |||||||
| 消减值SBV/cP | E1 | 1728.0±36.0 | -1994.5±97.5 | -2505.0~2160.5 | 1045.9±1177.2 | 3.1 | -2.1 | ||||||
| E2 | 2360.5±28.5 | 218.0±37.0 | -1765.0~3005.5 | 2072.3±982.7 | 4.6 | -2.3 | |||||||
| E3 | 1743.5±31.5 | -1546.5±12.5 | -1434.5~2505.0 | 1198.3±760.4 | 3.9 | -2.1 | |||||||
| E4 | 1344.0±42.0 | -720.0±31.0 | -1940.5~2189.5 | 1179.2±865.0 | 4.6 | -2.3 | |||||||
| 回复值CSV/cP | E1 | 3251.5±125.5 | 1402.0±20.0 | 1187.5~3546.5 | 2709.4±602.8 | 1.9 | -1.7 | ||||||
| E2 | 3196.0±59.0 | 1810.0±10.0 | 1235.0~3757.5 | 2973.9±584.8 | 1.5 | -1.5 | |||||||
| E3 | 2859.5±27.5 | 953.5±5.5 | 1121.5~3745.0 | 2543.1±545.3 | 1.0 | -1.1 | |||||||
| E4 | 2817.5±55.5 | 1406.0±28.0 | 1138.5~3460.0 | 2544.0±520.0 | 1.2 | -1.2 | |||||||
| 成糊温度PaT/℃ | E1 | 72.4±0.3 | 74.6±0.7 | 68.1~76.0 | 71.8±1.5 | 1.3 | 0.3 | ||||||
| E2 | 78.4±0.8 | 80.7±0.1 | 74.1~87.0 | 81.1±3.3 | -0.6 | -0.6 | |||||||
| E3 | 70.6±0.1 | 76.8±0.3 | 70.0~76.4 | 72.1±1.2 | 1.2 | 0.9 | |||||||
| E4 | 72.1±0.1 | 73.7±0.3 | 70.9~74.8 | 72.1±1.5 | 1.6 | 1.1 | |||||||
| 成糊时间PeT/min | E1 | 5.9±0.1 | 5.7±0.1 | 5.5~6.2 | 5.9±0.1 | 1.4 | -1.0 | ||||||
| E2 | 6.1±0.0 | 6.0±0.0 | 5.7~6.4 | 6.0±0.1 | 0.4 | 0.2 | |||||||
| E3 | 6.0±0.1 | 5.7±0.3 | 5.7~6.7 | 6.0±0.2 | 0.3 | 0.6 | |||||||
| E4 | 6.0±0.2 | 6.2±0.1 | 5.6~6.4 | 6.1±0.2 | -0.2 | -0.2 | |||||||
| 性状Trait | 环境E | 性状Trait | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PGWC | GC | PC | PKV | HPV | BDV | CPV | SBV | CSV | PaT | |||
| GC | E1 | 0.08 | ||||||||||
| E2 | 0.11 | |||||||||||
| E3 | 0.05 | |||||||||||
| E4 | –0.14 | |||||||||||
| PC | E1 | –0.08 | –0.13 | |||||||||
| E2 | –0.31* | –0.00 | ||||||||||
| E3 | –0.42** | –0.29* | ||||||||||
| E4 | –0.18 | –0.05 | ||||||||||
| PKV | E1 | –0.18 | 0.27* | –0.21 | ||||||||
| E2 | 0.23 | 0.25 | –0.76** | |||||||||
| E3 | 0.02 | 0.19 | –0.31* | |||||||||
| E4 | –0.07 | 0.37** | –0.03 | |||||||||
| HPV | E1 | –0.04 | –0.48** | –0.08 | 0.03 | |||||||
| E2 | –0.23 | –0.48** | –0.46** | 0.50** | ||||||||
| E3 | 0.25 | –0.27* | –0.28* | 0.55** | ||||||||
| E4 | 0.16 | –0.35** | –0.08 | –0.14 | ||||||||
| BDV | E1 | –0.09 | 0.54** | –0.09 | 0.68** | –0.71** | ||||||
| E2 | 0.45** | 0.70** | –0.43** | 0.65** | –0.33* | |||||||
| E3 | –0.27* | 0.48** | 0.04 | 0.26 | –0.66** | |||||||
| E4 | –0.16 | 0.47** | 0.05 | 0.64** | –0.86** | |||||||
| CPV | E1 | –0.04 | –0.50** | –0.05 | –0.16 | 0.96** | –0.82** | |||||
| E2 | –0.34** | –0.62** | –0.33** | 0.22 | 0.91** | –0.56** | ||||||
| E3 | 0.17 | –0.41** | –0.15 | 0.44** | 0.85** | –0.59** | ||||||
| E4 | –0.03 | –0.47** | –0.05 | –0.28* | 0.84** | –0.80** | ||||||
| SBV | E1 | 0.03 | –0.54** | 0.03 | –0.51** | 0.83** | –0.96** | 0.93** | ||||
| E2 | –0.45** | –0.74** | 0.10 | –0.33** | 0.61** | –0.90** | 0.85** | |||||
| E3 | 0.19 | –0.54** | –0.04 | 0.06 | 0.71** | –0.77** | 0.92** | |||||
| E4 | –0.01 | –0.51** | –0.04 | –0.51** | 0.79** | –0.89** | 0.97** | |||||
| CSV | E1 | –0.04 | –0.49** | –0.03 | –0.29* | 0.89** | –0.85** | 0.98** | 0.96** | |||
| E2 | –0.38** | –0.65** | –0.19 | –0.01 | 0.75** | –0.67** | 0.95** | 0.93** | ||||
| E3 | 0.08 | –0.43** | –0.03 | 0.26 | 0.55** | –0.41** | 0.91** | 0.90** | ||||
| E4 | –0.15 | –0.46** | –0.03 | –0.32* | 0.60** | –0.63** | 0.94** | 0.92** | ||||
| PaT | E1 | –0.18 | 0.16 | 0.48** | 0.16 | –0.54** | 0.51** | –0.57** | –0.55** | –0.56** | ||
| E2 | –0.29* | –0.11 | 0.76** | –0.89** | –0.45** | –0.58** | –0.26* | 0.23 | –0.09 | |||
| E3 | –0.05 | –0.18 | 0.27* | –0.42** | –0.40** | 0.09 | –0.43** | –0.30* | –0.36** | |||
| E4 | –0.19 | –0.01 | 0.44** | –0.14 | –0.32** | 0.18 | –0.23 | –0.17 | –0.14 | |||
| PeT | E1 | 0.18 | –0.37** | 0.14 | –0.38** | 0.67** | –0.76** | 0.66** | 0.71** | 0.61** | –0.30* | |
| E2 | –0.18 | –0.51** | 0.48** | –0.44** | 0.21 | –0.67** | 0.24 | 0.47** | 0.24 | 0.49** | ||
| E3 | 0.21 | –0.10 | –0.13 | –0.06 | 0.55** | –0.69** | 0.17 | 0.21 | –0.17 | 0.05 | ||
| E4 | 0.37** | –0.18 | –0.04 | –0.32* | 0.64** | –0.67** | 0.23 | 0.29* | –0.07 | –0.05 | ||
表2 昌恢121/越光CSSLs群体在4个环境中稻米各品质性状间的相关系数
Table 2 Coefficients of pairwise correlations among rice quality traits in the Changhui 121/Koshihikari CSSLs population.
| 性状Trait | 环境E | 性状Trait | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PGWC | GC | PC | PKV | HPV | BDV | CPV | SBV | CSV | PaT | |||
| GC | E1 | 0.08 | ||||||||||
| E2 | 0.11 | |||||||||||
| E3 | 0.05 | |||||||||||
| E4 | –0.14 | |||||||||||
| PC | E1 | –0.08 | –0.13 | |||||||||
| E2 | –0.31* | –0.00 | ||||||||||
| E3 | –0.42** | –0.29* | ||||||||||
| E4 | –0.18 | –0.05 | ||||||||||
| PKV | E1 | –0.18 | 0.27* | –0.21 | ||||||||
| E2 | 0.23 | 0.25 | –0.76** | |||||||||
| E3 | 0.02 | 0.19 | –0.31* | |||||||||
| E4 | –0.07 | 0.37** | –0.03 | |||||||||
| HPV | E1 | –0.04 | –0.48** | –0.08 | 0.03 | |||||||
| E2 | –0.23 | –0.48** | –0.46** | 0.50** | ||||||||
| E3 | 0.25 | –0.27* | –0.28* | 0.55** | ||||||||
| E4 | 0.16 | –0.35** | –0.08 | –0.14 | ||||||||
| BDV | E1 | –0.09 | 0.54** | –0.09 | 0.68** | –0.71** | ||||||
| E2 | 0.45** | 0.70** | –0.43** | 0.65** | –0.33* | |||||||
| E3 | –0.27* | 0.48** | 0.04 | 0.26 | –0.66** | |||||||
| E4 | –0.16 | 0.47** | 0.05 | 0.64** | –0.86** | |||||||
| CPV | E1 | –0.04 | –0.50** | –0.05 | –0.16 | 0.96** | –0.82** | |||||
| E2 | –0.34** | –0.62** | –0.33** | 0.22 | 0.91** | –0.56** | ||||||
| E3 | 0.17 | –0.41** | –0.15 | 0.44** | 0.85** | –0.59** | ||||||
| E4 | –0.03 | –0.47** | –0.05 | –0.28* | 0.84** | –0.80** | ||||||
| SBV | E1 | 0.03 | –0.54** | 0.03 | –0.51** | 0.83** | –0.96** | 0.93** | ||||
| E2 | –0.45** | –0.74** | 0.10 | –0.33** | 0.61** | –0.90** | 0.85** | |||||
| E3 | 0.19 | –0.54** | –0.04 | 0.06 | 0.71** | –0.77** | 0.92** | |||||
| E4 | –0.01 | –0.51** | –0.04 | –0.51** | 0.79** | –0.89** | 0.97** | |||||
| CSV | E1 | –0.04 | –0.49** | –0.03 | –0.29* | 0.89** | –0.85** | 0.98** | 0.96** | |||
| E2 | –0.38** | –0.65** | –0.19 | –0.01 | 0.75** | –0.67** | 0.95** | 0.93** | ||||
| E3 | 0.08 | –0.43** | –0.03 | 0.26 | 0.55** | –0.41** | 0.91** | 0.90** | ||||
| E4 | –0.15 | –0.46** | –0.03 | –0.32* | 0.60** | –0.63** | 0.94** | 0.92** | ||||
| PaT | E1 | –0.18 | 0.16 | 0.48** | 0.16 | –0.54** | 0.51** | –0.57** | –0.55** | –0.56** | ||
| E2 | –0.29* | –0.11 | 0.76** | –0.89** | –0.45** | –0.58** | –0.26* | 0.23 | –0.09 | |||
| E3 | –0.05 | –0.18 | 0.27* | –0.42** | –0.40** | 0.09 | –0.43** | –0.30* | –0.36** | |||
| E4 | –0.19 | –0.01 | 0.44** | –0.14 | –0.32** | 0.18 | –0.23 | –0.17 | –0.14 | |||
| PeT | E1 | 0.18 | –0.37** | 0.14 | –0.38** | 0.67** | –0.76** | 0.66** | 0.71** | 0.61** | –0.30* | |
| E2 | –0.18 | –0.51** | 0.48** | –0.44** | 0.21 | –0.67** | 0.24 | 0.47** | 0.24 | 0.49** | ||
| E3 | 0.21 | –0.10 | –0.13 | –0.06 | 0.55** | –0.69** | 0.17 | 0.21 | –0.17 | 0.05 | ||
| E4 | 0.37** | –0.18 | –0.04 | –0.32* | 0.64** | –0.67** | 0.23 | 0.29* | –0.07 | –0.05 | ||
| 位点 QTL | 区间 Marker interval | LOD值LOD value | 贡献率PVE/% | 加性效应Additive effect | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | ||
| qPGWC1 | RM3143-RM1117 | 5.02 | 10.45 | 4.40 | 29.34 | 57.67 | 30.84 | 18.40 | 13.34 | 20.36 | |||
| qPGWC2 | RM5529-RM7581 | 4.27 | 16.61 | 4.58 | |||||||||
| qPGWC5 | RM3345-RM17954 | 2.93 | 10.99 | 8.89 | |||||||||
| qPGWC6 | RM7088-RM7311 | 3.54 | 19.27 | 10.65 | |||||||||
| qGC5 | RM3345-RM17954 | 6.54 | 34.67 | 15.72 | |||||||||
| qGC6.1 | RM1369-RM510 | 6.76 | 5.64 | 41.02 | 25.07 | 15.78 | 13.60 | ||||||
| qGC6.2 | RM510-RM1163 | 2.99 | 22.48 | 10.86 | |||||||||
| qGC9 | STS-P0692F07- STS-OJ1001_G09 | 3.48 | 16.05 | 6.97 | |||||||||
| qPC10.1 | RM6737-RM25923 | 2.71 | 19.04 | ﹣0.59 | |||||||||
| qPC10.2 | RM25923-RM271 | 2.76 | 13.15 | 0.28 | |||||||||
表3 昌恢121/越光CSSLs群体垩白粒率、胶稠度和蛋白质含量QTL分析
Table 3 QTL detected for PGWC, GC, PC in the CSSLs population.
| 位点 QTL | 区间 Marker interval | LOD值LOD value | 贡献率PVE/% | 加性效应Additive effect | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | ||
| qPGWC1 | RM3143-RM1117 | 5.02 | 10.45 | 4.40 | 29.34 | 57.67 | 30.84 | 18.40 | 13.34 | 20.36 | |||
| qPGWC2 | RM5529-RM7581 | 4.27 | 16.61 | 4.58 | |||||||||
| qPGWC5 | RM3345-RM17954 | 2.93 | 10.99 | 8.89 | |||||||||
| qPGWC6 | RM7088-RM7311 | 3.54 | 19.27 | 10.65 | |||||||||
| qGC5 | RM3345-RM17954 | 6.54 | 34.67 | 15.72 | |||||||||
| qGC6.1 | RM1369-RM510 | 6.76 | 5.64 | 41.02 | 25.07 | 15.78 | 13.60 | ||||||
| qGC6.2 | RM510-RM1163 | 2.99 | 22.48 | 10.86 | |||||||||
| qGC9 | STS-P0692F07- STS-OJ1001_G09 | 3.48 | 16.05 | 6.97 | |||||||||
| qPC10.1 | RM6737-RM25923 | 2.71 | 19.04 | ﹣0.59 | |||||||||
| qPC10.2 | RM25923-RM271 | 2.76 | 13.15 | 0.28 | |||||||||
| 位点 QTL | 区间 Marker interval | LOD值LOD value | 贡献率PVE/% | ||||||
|---|---|---|---|---|---|---|---|---|---|
| E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | ||
| qPKV1 | RM6292-RM5362 | 3.08 | 10.44 | ||||||
| qPKV3 | RM6297-RM1278 | 6.30 | 40.98 | ||||||
| qPKV6.1 | RM1369-RM510 | 3.42 | 25.32 | ||||||
| qPKV6.2 | RM510-RM1163 | 5.61 | 20.35 | ||||||
| qPKV10 | RM6737-RM25923 | 4.19 | 22.55 | ||||||
| qPKV11 | STS-OSJNBa0095P22-RM5766 | 3.05 | 15.69 | ||||||
| qHPV3 | RM6297-RM1278 | 5.61 | 17.01 | ||||||
| qHPV6 | RM1369-RM510 | 12.22 | 4.98 | 12.64 | 30.11 | 64.75 | 30.21 | 53.41 | 76.71 |
| qHPV8 | RM6838-RM5767 | 2.79 | 3.21 | ||||||
| qBDV3 | RM15887-RM15903 | 3.79 | 3.34 | ||||||
| qBDV5 | RM3345-RM17954 | 3.72 | 14.25 | ||||||
| qBDV6 | RM1369-RM510 | 25.94 | 4.98 | 11.39 | 35.00 | 89.05 | 32.21 | 61.47 | 77.94 |
| qCPV3.1 | RM15887-RM15903 | 2.78 | 4.40 | ||||||
| qCPV3.2 | RM6297-RM1278 | 11.11 | 13.98 | ||||||
| qCPV6 | RM1369-RM510 | 20.80 | 10.46 | 22.26 | 45.45 | 80.40 | 50.81 | 60.34 | 90.23 |
| qCPV8 | RM5556-RM6838 | 11.58 | 14.92 | ||||||
| qSBV2 | RM5529-RM7581 | 2.88 | 5.12 | ||||||
| qSBV3 | RM15887-RM15903 | 5.06 | 1.61 | ||||||
| qSBV5 | RM3345-RM17954 | 3.54 | 6.45 | ||||||
| qSBV6.1 | RM1369-RM510 | 33.68 | 19.23 | 22.89 | 56.05 | 94.34 | 70.67 | 85.28 | 90.49 |
| qSBV6.2 | RM510-RM1163 | 5.81 | 1.89 | ||||||
| qSBV8 | RM7049-RM7556 | 3.37 | 1.03 | ||||||
| qCSV3 | RM15887-RM15903 | 3.23 | 3.72 | ||||||
| qCSV5 | RM3476-RM3790 | 2.74 | 2.97 | ||||||
| qCSV6 | RM1369-RM510 | 24.77 | 12.92 | 11.12 | 30.99 | 85.33 | 63.53 | 60.61 | 76.79 |
| qPaT4 | RM3839-RM303 | 3.53 | 5.04 | ||||||
| qPaT6.1 | RM1369-RM510 | 2.70 | 11.27 | 15.05 | 19.62 | ||||
| qPaT6.2 | RM510-RM1163 | 4.63 | 32.63 | ||||||
| qPaT10 | RM6737-RM25923 | 4.59 | 30.13 | ||||||
| qPaT12 | RM3331-RM5479 | 3.90 | 20.99 | 22.88 | 48.48 | ||||
| qPeT6 | RM1369-RM510 | 11.05 | 3.59 | 48.48 | 16.77 | ||||
| qPeT7.1 | RM1132-RM8261 | 3.51 | 10.78 | ||||||
| qPeT7.2 | STS-OJ1123_C12-RM5752 | 4.47 | 23.25 | ||||||
| qPeT7.3 | RM7273-RM3404 | 2.95 | 14.44 | ||||||
表4 昌恢121/越光CSSLs群体8个RVA谱特征值的QTL分析
Table 4 QTL detected for RVA in CSSLs population.
| 位点 QTL | 区间 Marker interval | LOD值LOD value | 贡献率PVE/% | ||||||
|---|---|---|---|---|---|---|---|---|---|
| E1 | E2 | E3 | E4 | E1 | E2 | E3 | E4 | ||
| qPKV1 | RM6292-RM5362 | 3.08 | 10.44 | ||||||
| qPKV3 | RM6297-RM1278 | 6.30 | 40.98 | ||||||
| qPKV6.1 | RM1369-RM510 | 3.42 | 25.32 | ||||||
| qPKV6.2 | RM510-RM1163 | 5.61 | 20.35 | ||||||
| qPKV10 | RM6737-RM25923 | 4.19 | 22.55 | ||||||
| qPKV11 | STS-OSJNBa0095P22-RM5766 | 3.05 | 15.69 | ||||||
| qHPV3 | RM6297-RM1278 | 5.61 | 17.01 | ||||||
| qHPV6 | RM1369-RM510 | 12.22 | 4.98 | 12.64 | 30.11 | 64.75 | 30.21 | 53.41 | 76.71 |
| qHPV8 | RM6838-RM5767 | 2.79 | 3.21 | ||||||
| qBDV3 | RM15887-RM15903 | 3.79 | 3.34 | ||||||
| qBDV5 | RM3345-RM17954 | 3.72 | 14.25 | ||||||
| qBDV6 | RM1369-RM510 | 25.94 | 4.98 | 11.39 | 35.00 | 89.05 | 32.21 | 61.47 | 77.94 |
| qCPV3.1 | RM15887-RM15903 | 2.78 | 4.40 | ||||||
| qCPV3.2 | RM6297-RM1278 | 11.11 | 13.98 | ||||||
| qCPV6 | RM1369-RM510 | 20.80 | 10.46 | 22.26 | 45.45 | 80.40 | 50.81 | 60.34 | 90.23 |
| qCPV8 | RM5556-RM6838 | 11.58 | 14.92 | ||||||
| qSBV2 | RM5529-RM7581 | 2.88 | 5.12 | ||||||
| qSBV3 | RM15887-RM15903 | 5.06 | 1.61 | ||||||
| qSBV5 | RM3345-RM17954 | 3.54 | 6.45 | ||||||
| qSBV6.1 | RM1369-RM510 | 33.68 | 19.23 | 22.89 | 56.05 | 94.34 | 70.67 | 85.28 | 90.49 |
| qSBV6.2 | RM510-RM1163 | 5.81 | 1.89 | ||||||
| qSBV8 | RM7049-RM7556 | 3.37 | 1.03 | ||||||
| qCSV3 | RM15887-RM15903 | 3.23 | 3.72 | ||||||
| qCSV5 | RM3476-RM3790 | 2.74 | 2.97 | ||||||
| qCSV6 | RM1369-RM510 | 24.77 | 12.92 | 11.12 | 30.99 | 85.33 | 63.53 | 60.61 | 76.79 |
| qPaT4 | RM3839-RM303 | 3.53 | 5.04 | ||||||
| qPaT6.1 | RM1369-RM510 | 2.70 | 11.27 | 15.05 | 19.62 | ||||
| qPaT6.2 | RM510-RM1163 | 4.63 | 32.63 | ||||||
| qPaT10 | RM6737-RM25923 | 4.59 | 30.13 | ||||||
| qPaT12 | RM3331-RM5479 | 3.90 | 20.99 | 22.88 | 48.48 | ||||
| qPeT6 | RM1369-RM510 | 11.05 | 3.59 | 48.48 | 16.77 | ||||
| qPeT7.1 | RM1132-RM8261 | 3.51 | 10.78 | ||||||
| qPeT7.2 | STS-OJ1123_C12-RM5752 | 4.47 | 23.25 | ||||||
| qPeT7.3 | RM7273-RM3404 | 2.95 | 14.44 | ||||||
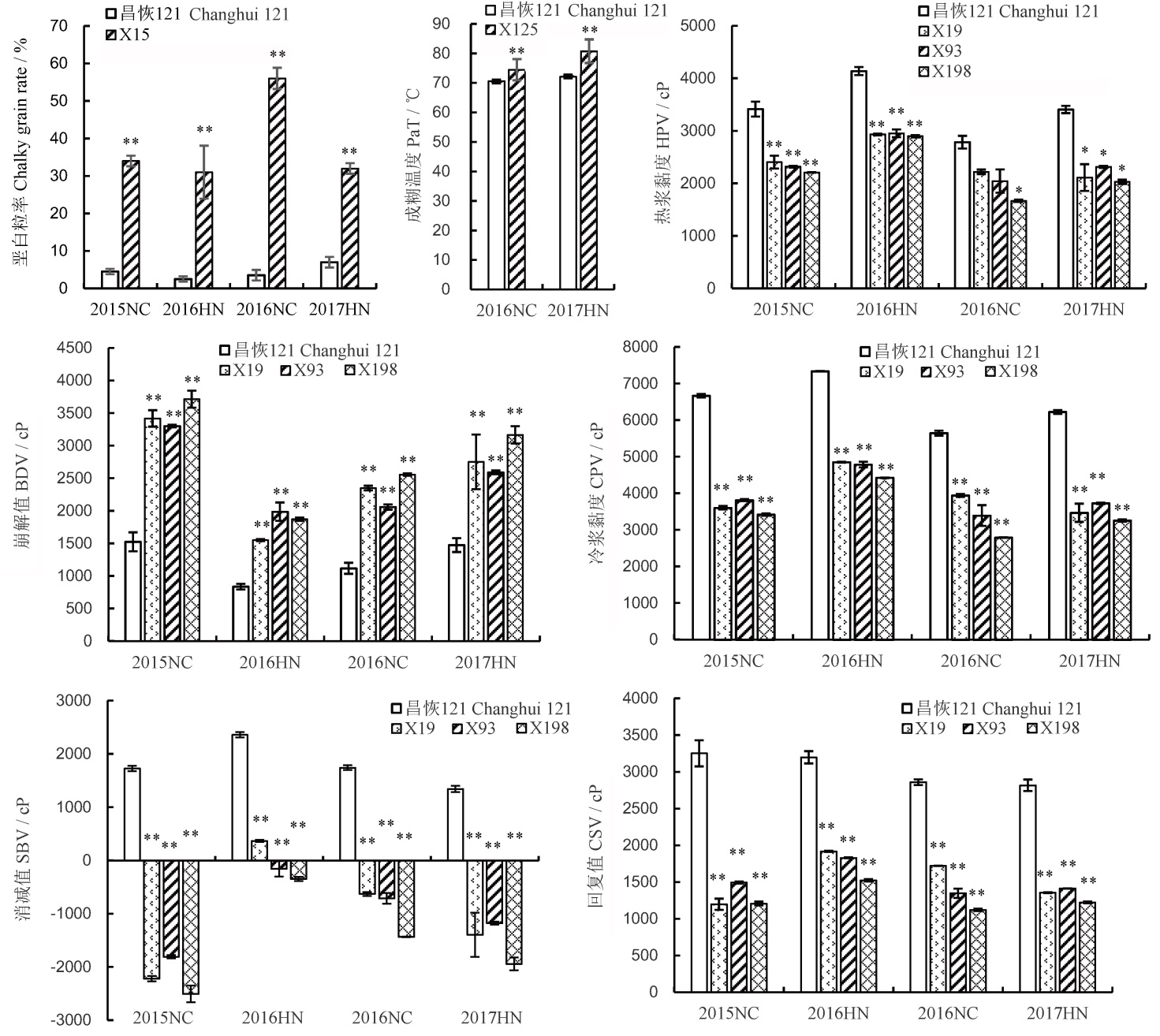
图2 稻米品质性状QTL对应片段代换系与背景亲本昌恢121的表型差异比较*和**分别表示昌恢121的品质性状与目标片段代换系之间的差异达5%和1%显著水平。NC和HN分别表示江西南昌和海南。
Fig. 2. Differences of phenotypic values of rice quality traits between genetic background parent Changhui 121 and the CSSLs harboring the QTL alleles. *and** mean significant difference at 5% and 1% levels between rice quality traits of Changhui 121 and the CSSLs harboring the QTL alleles. NC and HN mean Nanchang City, Jiangxi Province and Hainan Province, respectively.
| 染色体 Chr | 主效QTL簇 Major QTL cluster | 区间 Marker interval | 性状 Trait | 多效性QTL位点 Pleiotropic QTL cluster |
|---|---|---|---|---|
| 2 | qPGWC2 | RM5529–RM7581 | PGWC, SBV | qPGWC2, qSBV2 |
| 3 | qCPV3.2 | RM6297–RM1278 | CPV, PKV, HPV | qPKV3, qHPV3, qCPV3.2 |
| 3 | qSBV3 | RM15887–RM15903 | BDV, CPV, SBV, CSV | qBDV3, qCPV3.1, qSBV3, qCSV3 |
| 5 | qBDV5 | RM3345–RM17954 | BDV, SBV, PGWC, GC | qPGWC5, qGC5, qBDV5, qSBV5 |
| 6 | qSBV6.1 | RM1369–RM510 | GC, PKV, HPV, BDV, CPV, SBV, CSV, PaT, PeT | qGC6.1, qPKV6.1, qHPV6, qBDV6, qCPV6, qSBV6.1, qCSV6, qPaT6.1, qPeT6 |
| 6 | qSBV6.2 | RM510–RM1163 | GC, PKV, SBV, PaT | qGC6.2, qPKV6.2, qSBV6.2, qPaT6.2 |
| 10 | qPaT10 | RM6737–RM25923 | PKV, PaT, PC | qPKV10, qPaT10, qPC10.1 |
表5 水稻品质多效性区域分析
Table 5 Grain quality traits QTL hotspot and pleiotropic region in rice.
| 染色体 Chr | 主效QTL簇 Major QTL cluster | 区间 Marker interval | 性状 Trait | 多效性QTL位点 Pleiotropic QTL cluster |
|---|---|---|---|---|
| 2 | qPGWC2 | RM5529–RM7581 | PGWC, SBV | qPGWC2, qSBV2 |
| 3 | qCPV3.2 | RM6297–RM1278 | CPV, PKV, HPV | qPKV3, qHPV3, qCPV3.2 |
| 3 | qSBV3 | RM15887–RM15903 | BDV, CPV, SBV, CSV | qBDV3, qCPV3.1, qSBV3, qCSV3 |
| 5 | qBDV5 | RM3345–RM17954 | BDV, SBV, PGWC, GC | qPGWC5, qGC5, qBDV5, qSBV5 |
| 6 | qSBV6.1 | RM1369–RM510 | GC, PKV, HPV, BDV, CPV, SBV, CSV, PaT, PeT | qGC6.1, qPKV6.1, qHPV6, qBDV6, qCPV6, qSBV6.1, qCSV6, qPaT6.1, qPeT6 |
| 6 | qSBV6.2 | RM510–RM1163 | GC, PKV, SBV, PaT | qGC6.2, qPKV6.2, qSBV6.2, qPaT6.2 |
| 10 | qPaT10 | RM6737–RM25923 | PKV, PaT, PC | qPKV10, qPaT10, qPC10.1 |
| [1] | Chen L K, Gao W W, Chen S P, Wang L P, Zou J Y, Liu Y Z, Wang H, Chen Z Q, Guo T.High-resolution QTL mapping for grain appearance traits and co-localization of chalkiness-associated differentially expressed candidate genes in rice[J]. Rice, 2016, 9(1): 48-65. |
| [2] | Gao Y, Liu C L, Li Y Y, Zhang A P, Dong G J, Xie L H, Zhang B, Ruan B P, Hong K, Xue, D W, Zeng D L, Guo L B, Qian Q, Gao Z Y. QTL analysis for chalkiness of rice and fine mapping of a candidate gene for qACE9[J]. Rice, 2016, 9(1): 41. |
| [3] | Xu X M, Xu Z J, Matsue Y J, Xu Q.Effects of genetic background and environmental conditions on texture properties in a recombinant inbred population of an inter-subspecies cross[J]. Rice, 2019, 12(1): 32. |
| [4] | Tan Y F, Sun M, Xing Y Z, Hua J P, Sun X L, Zhang Q F, Corke H.Mapping quantitative trait loci for milling quality, protein content and color characteristics of rice using a recombinant inbred line population derived from an elite rice hybrid[J]. Theoretical and Applied Genetics, 2001, 103(6/7): 1037-1045. |
| [5] | 翁建峰, 万向元, 吴秀菊, 王海莲, 翟虎渠, 万建民. 利用CSSL群体研究稻米AC和PC相关QTL表达稳定性[J]. 作物学报, 2006, 32(1): 14-19. |
| Weng J F, Wan X Y, Wu X J, Wang H L, Zhai H Q, Wan J M.Stable expression of QTL for AC and PC of milled rice (Oryza sativa L.) using a CSSL population[J]. Acta Agronomica Sinica, 2006, 32(1): 14-19. (in Chinese with English abstract) | |
| [6] | 张涛, 郑家奎, 吴先军, 蒋开锋, 杨乾华, 陈温福, 杨莉. 水稻糙米蛋白质含量的QTL定位[J]. 分子植物育种, 2009, 7(1): 67-72. |
| Zhang T, Zheng J K, Wu X J, Jang K F, Yang Q H, Chen W F, Yang L.QTL mapping of brown rice protein content in a RIL population of rice[J]. Molecular Plant Breeding, 2009, 7(1): 67-72. (in Chinese with English abstract) | |
| [7] | 李晨, 潘大建, 孙传清, 周汉钦, 范芝兰, 陈建酉, 王象坤. 水稻糙米高蛋白基因的QTL定位[J]. 植物遗传资源学报, 2006, 7(2): 170-174. |
| Li C, Pan D J, Sun C Q, Zhou H Q, Fan Z L, Chen J Y, Wang X K.QTL location of high protein gene in rice grain[J]. Journal of Plant Genet, 2006, 7(2): 170-174. (in Chinese with English abstract) | |
| [8] | 杨亚春, 倪大虎, 宋丰顺, 李莉, 冯光, 李泽福, 杨剑波. 两个环境下糙米和精米蛋白质含量的QTL分析[J]. 中国水稻科学, 2012, 26(3): 351-355. |
| Yang Y C, Ni D H, Song F S, Li L, Feng G, Li Z F, Yang J B.Identification of QTL for protein content in brown and milled rice in two environments[J]. Chinese Journal of Rice Science, 2012, 26(3): 351-355. (in Chinese with English abstract) | |
| [9] | Bao J S, Harold C, He P, Zhu L H.Analysis of quantitative trait loci for starch properties of rice based on an RIL population[J]. Acta Botany Sinica, 2003, 45(8): 986-994. |
| [10] | Fan C C, Yu X Q, Xing Y Z, Xu C G, Luo L J, Zhang Q F.The main effects, epistatic effects and environmental interactions of QTLs on the cooking and eating quality of rice in a double-haploid line population[J]. Theoretical and Applied Genetics, 2005, 110(8): 1445-1452. |
| [11] | 吴长明, 孙传清, 王象坤, 李自超, 付秀林, 张强. 稻米食味品质性状的QTL分析[J]. 吉林农业科学, 2003, 28(2): 6-14. |
| Wu C M, Sun C Q, Wang X K, Li Z C, Fu X L, Zhang Q.Study on QTLs of grain eating quality characters in rice[J]. Journal of Jilin Agricultural Science, 2003, 28(2): 6-14. (in Chinese with English abstract) | |
| [12] | 邵高能, 唐绍清, 焦桂爱, 罗炬, 唐傲, 胡培松. 稻米蒸煮品质性状的QTL定位[J]. 中国水稻科学, 2009, 23(1): 94-98. |
| Shao G N, Tang S Q, Jiao G A, Luo J, Tang A, Hu P S.Mapping of QTL for cooking quality traits of rice[J]. Chinese Journal of Rice Science, 2009, 23(1): 94-98. (in Chinese with English abstract) | |
| [13] | Lanceras J C, Huang Z L, Naivikul O, Vanavichit A, Ruanjaichon V, Tragoonrung S.Mapping of genes for cooking and eating qualities in Thai Jasmine rice (KDML105)[J]. DNA Research, 2000, 7(2): 93-101. |
| [14] | 张巧凤, 张亚东, 朱镇, 赵凌, 赵庆勇, 许凌, 王才林. 稻米淀粉黏滞性(RVA谱)特征值的遗传及QTL定位分析[J]. 中国水稻科学, 2007, 21(6): 591-598. |
| Zhang Q F, Zhang Y D, Zhu Z, Zhao L, Zhao Q Y, Xu L, Wang C L.Analysis of inheritance and QTLs of rice starch viscosity (RVA profile) characteristics[J]. Chinese Journal of Rice Science, 2007, 21(6): 591-598. (in Chinese with English abstract) | |
| [15] | 张永生, 江玲, 刘喜, 刘世家, 陈亮明, 翟虎渠, 万建民. 优质水稻品种越光(Koshihikari)中控制稻米淀粉RVA谱特征值的QTL分析[J]. 中国水稻科学, 2010, 24(2): 137-144. |
| Zhang Y S, Jiang L, Liu X, Liu S J, Chen L M, Zhai H Q, Wan J M.Analysis of QTLs for starch RVA profile properties in the superior rice cultivar Koshihikari[J]. Chinese Journal of Rice Science, 2010, 24(2): 137-144. (in Chinese with English abstract) | |
| [16] | 杨亚春, 倪大虎, 宋丰顺, 李莉, 陆徐忠, 李泽福, 杨剑波. 不同生态环境下稻米淀粉RVA谱特征值的QTL定位分析[J]. 作物学报, 2012, 38(2): 264-274. |
| Yang Y C, Ni D H, Song F S, Li L, Lu X Z, Li Z F, Yang J B.Identification of QTL for rice starch RVA profile properties under different ecological sites[J]. Acta Agronomy Sinica, 2012, 38(2): 264-274. (in Chinese with English abstract) | |
| [17] | 张昌泉, 胡冰, 朱孔志, 张华, 冷亚麟, 汤述翥, 顾铭洪, 刘巧泉. 利用重测序的水稻染色体片段代换系定位控制稻米淀粉黏滞性谱QTL[J]. 中国水稻科学, 2013, 27(1): 56-64. |
| Zhang C Q, Hu B, Zhu K Z, Zhang H, Leng Y L, Tang S Z, Gu M H, Liu Q Q.Mapping of QTLs for rice RVA properties using high-throughput re-sequenced chromosome segment substitution lines[J]. Chinese Journal of Rice Science, 2013, 27(1): 56-64. (in Chinese with English abstract) | |
| [18] | 张杰, 郑蕾娜, 蔡跃, 尤小满, 孔飞, 汪国湘, 燕海刚, 金洁, 王亮. 稻米淀粉RVA谱特征值与直链淀粉、蛋白含量的相关性及QTL定位分析[J]. 中国水稻科学, 2017, 31(1): 31-39. |
| Zhang J, Zheng L N, Cai Y, You X M, Kong F, Wang G X, Yan H G, Jin J, Wang L.Correlation analysis and QTL mapping for starch RVA profile properties and amylose and protein contents in rice[J]. Chinese Journal of Rice Science, 2017, 31(1): 31-39. (in Chinese with English abstract) | |
| [19] | Matsushima R, Maekawa M, Kusano M, Kondo H, Naoko F, Kawagoe Y, Sakamoto W.Amyloplast- localized SUBSTANDARD STARCH GRAIN4 protein influences the size of starch grains in rice endosperm[J]. Plant Physiology, 2014, 164(2): 623-636. |
| [20] | She K, Kusano H, Koizumi K, Yamakawa H, Hakata M, Imamura T, Fukuda M, Naito N, Tsurumaki Y, Yaeshima M, Tsuge T, Kudoh M, Itoh E, Kikuchi S, Kishimoto N, Yazaki J, Ando T, Yano M, Aoyama T, Sasaki T, Satoh H, Shimada H.A novel factor FLOURYENDOSPERM2 is involved in regulation of rice grain size and starch quality[J]. The Plant Cell, 2010, 22(10): 3280-3294. |
| [21] | Wang E, Wang J J, Zhu X D, Hao W, Wang L Y, Li Q, Zhang L X, He W, Lu B, Lin H X, Ma H, Zhang G Q, He Z H.Control of rice grain-filling and yield by a gene with a potential signature of domestication[J]. Nature Genetics, 2008, 40(11): 1370-1374. |
| [22] | Li Y B, Fan C C, Xing Y Z, Yun P, Luo L J, Yan B, Peng B, Xie W B, Wang G W, Li X H, Xiao J H, Xu C G, He Y Q.Chalk5 encodes a vacuolar H+ translocating pyrophosphatase influencing grain chalkiness in rice[J]. Nature Genetics, 2014, 46(4): 398-404. |
| [23] | Luo Q, Tang D, Wang M, Luo W X, Zhang L, Qin B X, Shen Y, Wang K J, Li Y F, Cheng Z K.The role ofOsMSH5 in crossover formation during rice meiosis. Molecular Plant, 2013, 3(6): 729-742. |
| [24] | Han X H, Wang Y H, Liu X, Jiang L, Ren Y L, Liu F, Peng C, Li J J, Jin X M, Wu F Q, Wang J L, Guo X P, Zhang X, Cheng Z J, Wan J M.The failure to express a protein disulphide isomerase-like protein results in a floury endosperm and an endoplasmic reticulum stress response in rice[J]. Journal of Experiment Botany, 2012, 63(1): 121-130. |
| [25] | Wan Y H, Ren Y L, Liu X, Jiang L, Chen L, Han X H, Jin M N, Liu S J, Liu F, Lv J, Zhou K N, Su N, Wan J M.OsRab5a regulates endomembrane organization and storage protein trafficking in rice endosperm cells[J]. Plant Journal, 2010, 64(5): 812-824. |
| [26] | Yang R F, Sun C L, Bai J J, Luo Z X, Shi B, Zhang J M, Yan W G, Piao Z Z.A putative gene sbe3-rs for resistant starch mutated from SBE3 for starch branching enzyme in rice (Oryza sativa L.)[J]. PLoS ONE, 2012, 7(8): e43026. |
| [27] | Cai Y C, Li S F, Jiao G A, Sheng Z H, Wu Y W, Shao G N, Xie L H, Peng C, Xu J F, Tang S Q, Wei X J, Hu P S.OsPK2 encodes a plastidic pyruvate kinase involved in rice endosperm starch synthesis, compound granule formation and grain filling[J]. Plant Biotechnology Journal, 2018, 16(11): 1878-1891. |
| [28] | Tuncel A, Kawaguchi J, Ihara Y, Matsusaka H, Nishi A, Nakamura T, Kuhara S, Hirakawa H, Nakamura Y, Cakir B, Nagamine A, Okita T W, Hwang S K, Satoh H.The rice endosperm ADP-glucose pyrophosphorylase large subunit is essential for optimal catalysis and allosteric regulation of the heterotetrameric enzyme[J]. Plant Cell Physiology, 2014, 55(6):1169-83. |
| [29] | McCouch S R. Gene nomenclature system for rice[J]. Rice, 2008, 1(1): 72-84. |
| [30] | 吴殿星, 舒庆尧, 夏英武. 利用RVA谱快速鉴别不同表观直链淀粉含量早籼稻的淀粉黏滞特性[J]. 中国水稻科学, 2001, 15(1): 58-60. |
| Wu D X, Shu Q Y, Xia Y W.Rapid identification of starch viscosity property of early indica rice varieties with different apparent amylose content by RVA profile[J]. Chinese Journal of Rice Science, 2001, 15(1): 58-60. (in Chinese with English abstract) | |
| [31] | 李欣, 张蓉, 隋炯明, 梁国华, 沈新平, 严长杰, 顾世梁, 顾铭洪. 稻米淀粉黏滞性谱特征的表现及其遗传[J]. 中国水稻科学, 2004, 18(5): 10-16. |
| Li X, Zhang R, Sui J M, Liang G H, Shen X P, Yan C J, Gu S L, Gu M H.Performance and inheritance of rice starch viscosity (RVA Profile) characteristics[J]. Chinese Journal of Rice Science, 2004, 18(5): 10-16. (in Chinese with English abstract) | |
| [32] | 张巧凤, 张亚东, 朱镇, 赵凌, 赵庆勇, 许凌, 王才林. 稻米淀粉黏滞性(RVA谱)特征值的遗传及QTL定位分析[J]. 中国水稻科学, 2007, 21(6): 591-598. |
| Zhang Q F, Zhang Y D, Zhu Z, Zhao L, Zhao Q Y, Xu L, Wang C L.Analysis of inheritance and QTLs of rice starch viscosity (RVA profile) characteristics[J]. Chinese Journal of Rice Science, 2007, 21(6): 591-598. (in Chinese with English abstract) | |
| [33] | 刘鑫燕, 朱孔志, 张昌泉, 洪燃, 孙鹏, 汤述翥, 顾铭洪, 刘巧泉. 利用9311来源的粳型染色体片段代换系定位控制稻米糊化温度的微效QTL[J]. 作物学报, 2014, 40(10): 1740-1747. |
| Liu X Y, Zhu K Z, Zhang C Q, Hong R, Sun P, Tang S Z, Gu M H, Liu Q Q.Mapping of minor QTLs for rice gelatinization temperature using chromo-some segment substitution lines from indica 9311 in the japonica background[J]. Acta Agronomy Sinica, 2014, 40(10): 1740-1747. (in Chinese with English abstract) |
| [1] | 汪邑晨, 朱本顺, 周磊, 朱骏, 杨仲南. 光/温敏核不育系的不育机理及两系杂交稻的发展与展望[J]. 中国水稻科学, 2024, 38(5): 463-474. |
| [2] | 许用强, 徐军, 奉保华, 肖晶晶, 王丹英, 曾宇翔, 符冠富. 水稻花粉管生长及其对非生物逆境胁迫的响应机理研究进展[J]. 中国水稻科学, 2024, 38(5): 495-506. |
| [3] | 何勇, 刘耀威, 熊翔, 祝丹晨, 王爱群, 马拉娜, 王廷宝, 张健, 李建雄, 田志宏. 利用CRISPR/Cas9技术编辑OsOFP30基因创制水稻粒型突变体[J]. 中国水稻科学, 2024, 38(5): 507-515. |
| [4] | 吕阳, 刘聪聪, 杨龙波, 曹兴岚, 王月影, 童毅, Mohamed Hazman, 钱前, 商连光, 郭龙彪. 全基因组关联分析(GWAS)鉴定水稻氮素利用效率候选基因[J]. 中国水稻科学, 2024, 38(5): 516-524. |
| [5] | 杨好, 黄衍焱, 王剑, 易春霖, 石军, 谭楮湉, 任文芮, 王文明. 水稻中八个稻瘟病抗性基因特异分子标记的开发及应用[J]. 中国水稻科学, 2024, 38(5): 525-534. |
| [6] | 杨铭榆, 陈志诚, 潘美清, 张汴泓, 潘睿欣, 尤林东, 陈晓艳, 唐莉娜, 黄锦文. 烟-稻轮作下减氮配施生物炭对水稻茎鞘同化物转运和产量 形成的影响[J]. 中国水稻科学, 2024, 38(5): 555-566. |
| [7] | 熊家欢, 张义凯, 向镜, 陈惠哲, 徐一成, 王亚梁, 王志刚, 姚坚, 张玉屏. 覆膜稻田施用炭基肥对水稻产量及氮素利用的影响[J]. 中国水稻科学, 2024, 38(5): 567-576. |
| [8] | 郭展, 张运波. 水稻对干旱胁迫的生理生化响应及分子调控研究进展[J]. 中国水稻科学, 2024, 38(4): 335-349. |
| [9] | 韦还和, 马唯一, 左博源, 汪璐璐, 朱旺, 耿孝宇, 张翔, 孟天瑶, 陈英龙, 高平磊, 许轲, 霍中洋, 戴其根. 盐、干旱及其复合胁迫对水稻产量和品质形成影响的研究进展[J]. 中国水稻科学, 2024, 38(4): 350-363. |
| [10] | 许丹洁, 林巧霞, 李正康, 庄小倩, 凌宇, 赖美玲, 陈晓婷, 鲁国东. OsOPR10正调控水稻对稻瘟病和白叶枯病的抗性[J]. 中国水稻科学, 2024, 38(4): 364-374. |
| [11] | 候小琴, 王莹, 余贝, 符卫蒙, 奉保华, 沈煜潮, 谢杭军, 王焕然, 许用强, 武志海, 王建军, 陶龙兴, 符冠富. 黄腐酸钾提高水稻秧苗耐盐性的作用途径分析[J]. 中国水稻科学, 2024, 38(4): 409-421. |
| [12] | 胡继杰, 胡志华, 张均华, 曹小闯, 金千瑜, 章志远, 朱练峰. 根际饱和溶解氧对水稻分蘖期光合及生长特性的影响[J]. 中国水稻科学, 2024, 38(4): 437-446. |
| [13] | 刘福祥, 甄浩洋, 彭焕, 郑刘春, 彭德良, 文艳华. 广东省水稻孢囊线虫病调查与鉴定[J]. 中国水稻科学, 2024, 38(4): 456-461. |
| [14] | 陈浩田, 秦缘, 钟笑涵, 林晨语, 秦竞航, 杨建昌, 张伟杨. 水稻根系和土壤性状与稻田甲烷排放关系的研究进展[J]. 中国水稻科学, 2024, 38(3): 233-245. |
| [15] | 缪军, 冉金晖, 徐梦彬, 卜柳冰, 王平, 梁国华, 周勇. 过量表达异三聚体G蛋白γ亚基基因RGG2提高水稻抗旱性[J]. 中国水稻科学, 2024, 38(3): 246-255. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||