
中国水稻科学 ›› 2026, Vol. 40 ›› Issue (2): 210-222.DOI: 10.16819/j.1001-7216.2026.250310
薛炮1,2, 王友霜1, 何弯弯1,2, 黄晨博3, 张涵3, 丁震乾1, 陈秋丽1, 范运新1, 丁成伟1, 孙廉平3,*( ), 胡婷婷1,2,*(
), 胡婷婷1,2,*( )
)
收稿日期:2025-03-12
修回日期:2025-06-05
出版日期:2026-03-10
发布日期:2026-03-17
通讯作者:
* email: htt713@163.com; sunlianping@caas.cn基金资助:
XUE Pao1,2, WANG Youshuang1, HE Wanwan1,2, HUANG Chenbo3, ZHANG Han3, DING Zhenqian1, CHEN Qiuli1, FAN Yunxin1, DING Chengwei1, SUN Lianping3,*( ), HU Tingting1,2,*(
), HU Tingting1,2,*( )
)
Received:2025-03-12
Revised:2025-06-05
Online:2026-03-10
Published:2026-03-17
Contact:
* email: htt713@163.com; sunlianping@caas.cn摘要:
【目的】水稻颖壳发育直接影响稻米产量与品质,挖掘颖壳发育基因并探究其分子功能,可为水稻优质高产遗传改良提供更多的基因资源。【方法】利用EMS诱变徐稻3号获得一个颖壳开裂后不闭合的突变体sg5 (split glume 5);对其农艺性状和不同开花时间进行浆片表型和细胞学观察;通过图位克隆与BSA-seq相结合的方法鉴定SG5基因;结合野生型各组织表达量检测及GUS转基因株系各组织染色情况,分析SG5组织表达特异性;利用激光共聚焦显微镜观察SG5蛋白在原生质体中的亚细胞定位;通过进化树分析,探讨了SG5在进化过程中的选择情况。【结果】突变体sg5在开花前颖壳发育正常,开花后颖壳保持开裂状态直至成熟。与野生型相比,突变体株高和有效穗数无显著变化,抽穗期显著提前,穗长增加,粒长、粒宽降低,结实率极显著降低。体视显微镜观察结果显示,突变体颖壳开裂是浆片保持膨大状态导致的。SG5基因定位在第5染色体InDel5- 11−InDel5-13区间内,物理间距为3.72 Mb,图位克隆表明sg5突变体中LOC_Os05g50890的第3内含子和第4外显子连接处发生G-A突变导致发生可变剪接,蛋白翻译提前终止。敲除实验证实SG5基因即为目的基因。SG5基因为组成型表达,在根、茎、叶中表达水平较高。SG5蛋白定位在细胞质和细胞核中。SG5在进化过程中发生籼稻、粳稻分化。【结论】本研究鉴定了OsJar1基因一个新的等位突变,发现SG5在自然选择过程中发生了籼、粳分化。本研究为后续深入研究颖壳调控分子机制提供了新的线索,并为水稻设计育种提供新的基因资源。
薛炮, 王友霜, 何弯弯, 黄晨博, 张涵, 丁震乾, 陈秋丽, 范运新, 丁成伟, 孙廉平, 胡婷婷. 水稻颖壳不闭合基因SG5的鉴定与克隆[J]. 中国水稻科学, 2026, 40(2): 210-222.
XUE Pao, WANG Youshuang, HE Wanwan, HUANG Chenbo, ZHANG Han, DING Zhenqian, CHEN Qiuli, FAN Yunxin, DING Chengwei, SUN Lianping, HU Tingting. Identification and Cloning of SG5 in Rice[J]. Chinese Journal OF Rice Science, 2026, 40(2): 210-222.
| 引物名称 Primer name | 引物序列(5'→3') Primer sequence (5'→3') | 功能 Function |
|---|---|---|
| InDel 5-11F | GGAGTTCTGGTGGCTTTCC | 基因定位 Gene mapping |
| InDel 5-11R | CAGTTCCCCATTTCCCTCTA | |
| InDel 5-13F | TGAGTTTCCGGTGTTCCATA | |
| InDel 5-13R | AAGGCAAAGTCGTTCAGCTT | |
| SG5-Cas9-F | ggcaATATTGAGCCCTATATCCAG | 基因编辑 Gene editing |
| SG5- Cas9-R | aaacCTGGATATAGGGCTCAATAT | |
| SG5-1305-GUS-F | catgattacgaattcATCACCTCTGCCCAAAACCA | GUS载体构建 Construction of GUS vector |
| SG5-1305-GUS-R | tcagatctaccatggTGTAATGGTAGAATCCTGGCTTTACCA | |
| SG5-1305-GFP-F | aagtccggagctagctctagaATGACGATCTGCAGCTGTGAGG | 亚细胞定位 Subcellular localization |
| SG5-1305-GFP-R | ggtcctcgagacgtctctagaAAATCCATAGGCAGTACTGAAATAACTTTGGG | |
| Hyg-F | ACGGTGTCGTCCATCACAGTTTGCC | 潮霉素基因检测 Detection of hygromycin resistance gene |
| Hyg-R | TTCCGGAAGTGCTTGACATTGGGGA | |
| Cas9-F | CGTGGAAGATCGGTTCAACGC | 阳性株检测 Sequencing of positive plants |
| Cas9-R | CTGCCGGCCAGATTGGCA | |
| CR-SG5-F | TGATGCCAACTCCTACTGCAC | 突变位点检测 Mutation site detection |
| CR-SG5-R | ACTTGGGCTTTCCGTGTGTT | |
| UBQ10-qPCR-F | TGGTCAGTAATCAGCCAGTTTGG | qRT-PCR |
| UBQ10-qPCR-R | GCACCACAAATACTTGACGAACAG | |
| SG5-qPCR-F | GATACCTCACCAGTGGTCACTG | |
| SG5-qPCR-R | TCCTAAACGCGTAAGATGTACGG |
表1 本研究所用的引物序列
Table 1. Primer sequences used in this study
| 引物名称 Primer name | 引物序列(5'→3') Primer sequence (5'→3') | 功能 Function |
|---|---|---|
| InDel 5-11F | GGAGTTCTGGTGGCTTTCC | 基因定位 Gene mapping |
| InDel 5-11R | CAGTTCCCCATTTCCCTCTA | |
| InDel 5-13F | TGAGTTTCCGGTGTTCCATA | |
| InDel 5-13R | AAGGCAAAGTCGTTCAGCTT | |
| SG5-Cas9-F | ggcaATATTGAGCCCTATATCCAG | 基因编辑 Gene editing |
| SG5- Cas9-R | aaacCTGGATATAGGGCTCAATAT | |
| SG5-1305-GUS-F | catgattacgaattcATCACCTCTGCCCAAAACCA | GUS载体构建 Construction of GUS vector |
| SG5-1305-GUS-R | tcagatctaccatggTGTAATGGTAGAATCCTGGCTTTACCA | |
| SG5-1305-GFP-F | aagtccggagctagctctagaATGACGATCTGCAGCTGTGAGG | 亚细胞定位 Subcellular localization |
| SG5-1305-GFP-R | ggtcctcgagacgtctctagaAAATCCATAGGCAGTACTGAAATAACTTTGGG | |
| Hyg-F | ACGGTGTCGTCCATCACAGTTTGCC | 潮霉素基因检测 Detection of hygromycin resistance gene |
| Hyg-R | TTCCGGAAGTGCTTGACATTGGGGA | |
| Cas9-F | CGTGGAAGATCGGTTCAACGC | 阳性株检测 Sequencing of positive plants |
| Cas9-R | CTGCCGGCCAGATTGGCA | |
| CR-SG5-F | TGATGCCAACTCCTACTGCAC | 突变位点检测 Mutation site detection |
| CR-SG5-R | ACTTGGGCTTTCCGTGTGTT | |
| UBQ10-qPCR-F | TGGTCAGTAATCAGCCAGTTTGG | qRT-PCR |
| UBQ10-qPCR-R | GCACCACAAATACTTGACGAACAG | |
| SG5-qPCR-F | GATACCTCACCAGTGGTCACTG | |
| SG5-qPCR-R | TCCTAAACGCGTAAGATGTACGG |
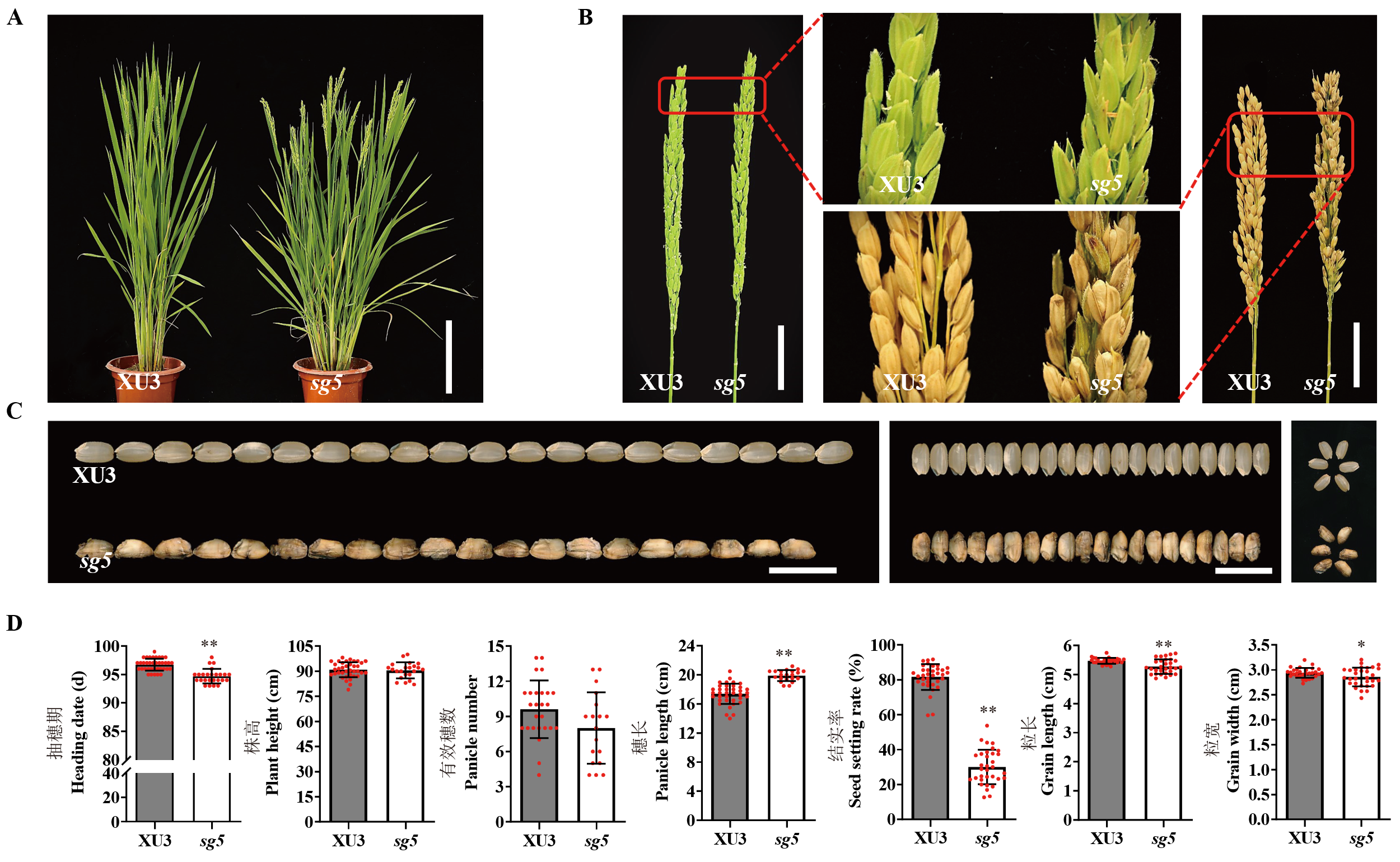
图1 野生型徐稻3号与sg5表型特征 A: 抽穗时野生型及突变体单株表型,比例尺=20 cm;B: 野生型及突变体颖壳及穗部形态,比例尺=5 cm;C: 野生型及突变体籽粒性状,比例尺=1 cm;D: 抽穗期、株高、有效穗数、穗长、结实率、粒长、粒宽表型差异。数据表示为平均值±标准差(n≥20),学生氏t-检验计算差异显著性。*突变体与野生型间的表型差异在P < 0.05水平上显著;**突变体与野生型间的表型差异在P < 0.01水平上极显著。
Fig. 1. Phenotypic characterization of wildtype (Xudao 3, XU3) and sg5 A, Characterization of plant type in wild type and sg5. Scale bar=20 cm. B, Glume morphology and panicle morphology in wild type and sg5. Scale bar=5 cm. C, Comparison of grain size in wild type and sg5. Scale bar=10 mm. D, Comparison of heading date, plant height, panicle number, panicle length, seed setting rate, grain length and grain width in wild type and sg5. Data are given as Mean ± SD (n≥20). Student’s t-test was used to generate P value. *, P < 0.05; **, P < 0.01.

图2 野生型(XU3)和sg5在开花前后不同时间颖壳及浆片的发育形态变化 野生型及突变体未开花前穗部状态(A)、颖壳形态(B)、开花后穗部状态(C)和开花后颖壳形态(D),比例尺=5 cm;E: 野生型及突变体前期长势一致的颖花,在开花前后不同时间的颖壳及浆片形态变化,比例尺=20μm。白框中是相对应的颖花。
Fig. 2. Developmental morphological changes of glumes and lodicules in wild type (XU3) and sg5 at various time points around flowering A, Characterization of panicle before flowering(A), glume before flowering(B), panicle after flowering(C), glume after flowering(D) in the wild type and sg5(Scale bar=5 cm). E, Florets of consistent early growth from wild type and sg5, showing morphological changes of glumes and lodicules at various time points around flowering. Scale bar = 20 μm. The white boxes indicate the corresponding florets.
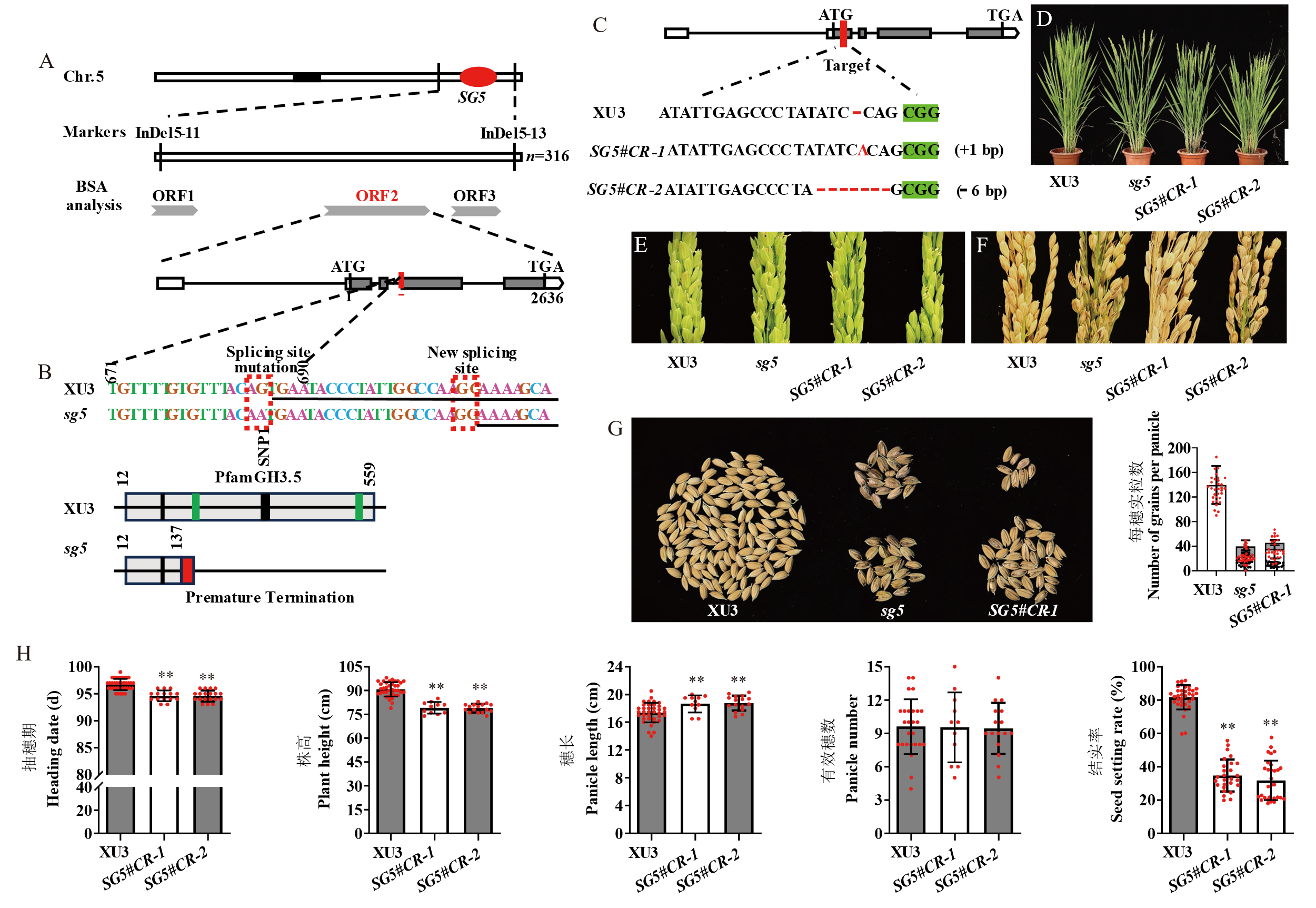
图3 SG5图位克隆及功能验证 A:SG5图位克隆;B: 野生型、突变体DNA序列及蛋白序列分析;下划线表示野生型SG5第4外显子前端序列,其他序列代表第3内含子后半部分;虚线红框表示剪接突变位点及新的剪接位点。黑色竖线表示茉莉酮酸酯位点;绿色竖线表示L-α-氨基酸位点。C:SG5基因编辑及纯合突变株序列;D: 野生型、突变体、SG5#CR-1、SG5#CR-2抽穗时单株表型,比例尺=20 cm, E: 野生型、突变体、SG5#CR-1、SG5#CR-2受精后颖壳形态;F: 野生型、突变体、SG5#CR-1、SG5#CR-2成熟期穗部形态,比例尺=5 cm;G:野生型、突变体、SG5#CR-1每穗实粒数;红点:正常籽粒;黑点:发霉籽粒。H:抽穗期、株高、有效穗数、穗长、结实率、粒长、粒宽表型差异。红点:单株数目。数据表示为平均值±标准差(n≥10),学生氏t检验计算差异显著性。*表示表型在P < 0.05水平显著;**表示表型在P < 0.01水平极显著。
Fig. 3. Map-based cloning and functional verification of SG5 A, Map-based cloning of SG5. B, Characterization of DNA and protein sequences in Xudao 3(XU3) and sg5. Underlined nucleotides are the sequence of the first half of exons 4 in XU3, others are at the terminal of the intron 3. Red boxes are the splicing site mutation and new splicing sites. Black vertical line, the site of jasmonate. Green vertical line, the site of L-alpha-amino acid. C, Gene editing and homozygous mutant sequences. D, Characterization of plant type in XU3, sg5 mutant, SG5#CR-1, and SG5#CR-2. Scale bar=20 cm. E, Glume morphology of XU3, sg5 mutant, SG5#CR-1, and SG5#CR-2; F, Panicle morphology of wild type, sg5 mutant, SG5#CR-1, and SG5#CR-2. Scale bar=5 cm. G, Number of grains per panicle in XU3, sg5 mutant, and SG5#CR-1; Red dot, Normal seeds. Black dot, Moldy seeds. H, Comparison of heading date, plant height, panicle length, panicle number, and seed setting rate in XU3, SG5#CR-1, and SG5#CR-2. Red dot, Number of plants investigated. Data are given as Mean ± SD (n≥10). Student’s t-test was used to generate P value. * P < 0.05; ** P < 0.01.
| 差异种类 Category | 差异基因数目 Number of differential genes |
|---|---|
| 基因上游 Upstream | 638 |
| 功能获得型 Stop gain | 6 |
| 功能缺失型 Stop loss | 2 |
| 5' UTR | 10 |
| 3' UTR | 14 |
| 保守结构缺失/插入 Conservative inframe deletion / insertion | 4 |
| 散乱结构缺失/插入 Disruptive inframe deletion / insertion | 3 |
| 移码突变 Frameshift variant | 23 |
| 错义突变 Missense variant | 119 |
| 同义突变 Synonymous variant | 98 |
| 剪接受体变异 Splice acceptor variant & intron variant | 4 |
| 内含子变异 Intronic | 60 |
| 基因间变异 Intergenic region | 72 |
| 基因下游 Downstream | 212 |
| 合计 Total | 1265 |
表2 BSA分析检测的差异位点汇总
Table 2. Summary of differentially expressed genes detected using BSA analysis
| 差异种类 Category | 差异基因数目 Number of differential genes |
|---|---|
| 基因上游 Upstream | 638 |
| 功能获得型 Stop gain | 6 |
| 功能缺失型 Stop loss | 2 |
| 5' UTR | 10 |
| 3' UTR | 14 |
| 保守结构缺失/插入 Conservative inframe deletion / insertion | 4 |
| 散乱结构缺失/插入 Disruptive inframe deletion / insertion | 3 |
| 移码突变 Frameshift variant | 23 |
| 错义突变 Missense variant | 119 |
| 同义突变 Synonymous variant | 98 |
| 剪接受体变异 Splice acceptor variant & intron variant | 4 |
| 内含子变异 Intronic | 60 |
| 基因间变异 Intergenic region | 72 |
| 基因下游 Downstream | 212 |
| 合计 Total | 1265 |
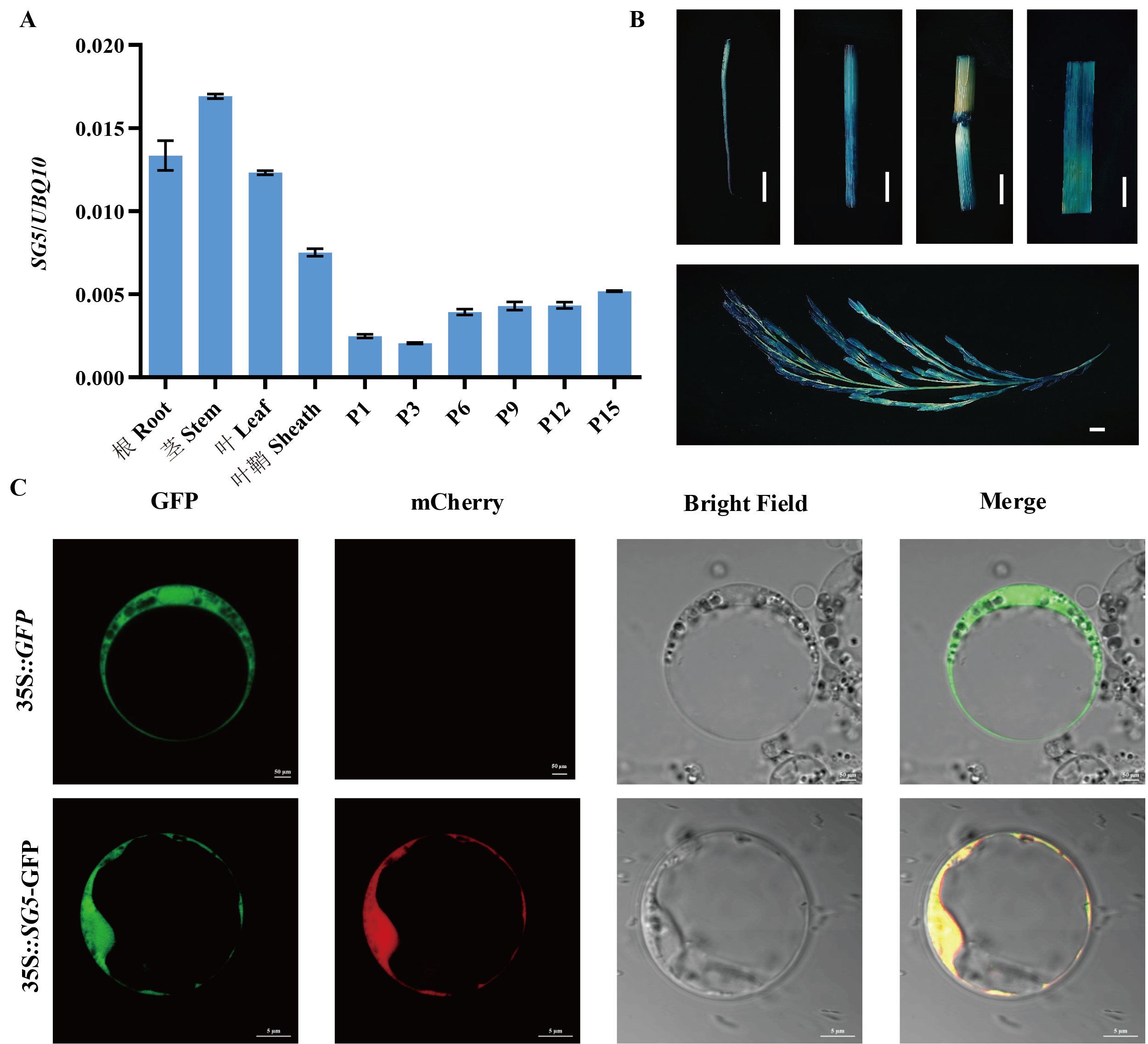
图4 SG5时空表达模式及亚细胞定位 A:SG5基因的组织表达模式分析,数据表示为平均值±标准差(n=3);B:SG5启动子启动GUS蛋白在各个组织部位中的染色情况(比例尺=10 mm)。C:SG5在原生质体中的亚细胞定位,35S::GFP空载 (比例尺=50 μm);35S::SG5-GFP载体(比例尺=5 μm)。
Fig. 4. Expression pattern analysis of SG5 and subcellular localization A, qRT-PCR analysis of SG5 gene expression in various tissues. Data are given as Mean ± SD (n=3). B, Histochemical analysis of GUS activity in various tissues.( Scale bar=10 mm). C, Subcellular localization of SG5 in rice protoplast. 35S::GFP(scale bar=50 μm). 35S::SG5-GFP(scale bar=5 μm).
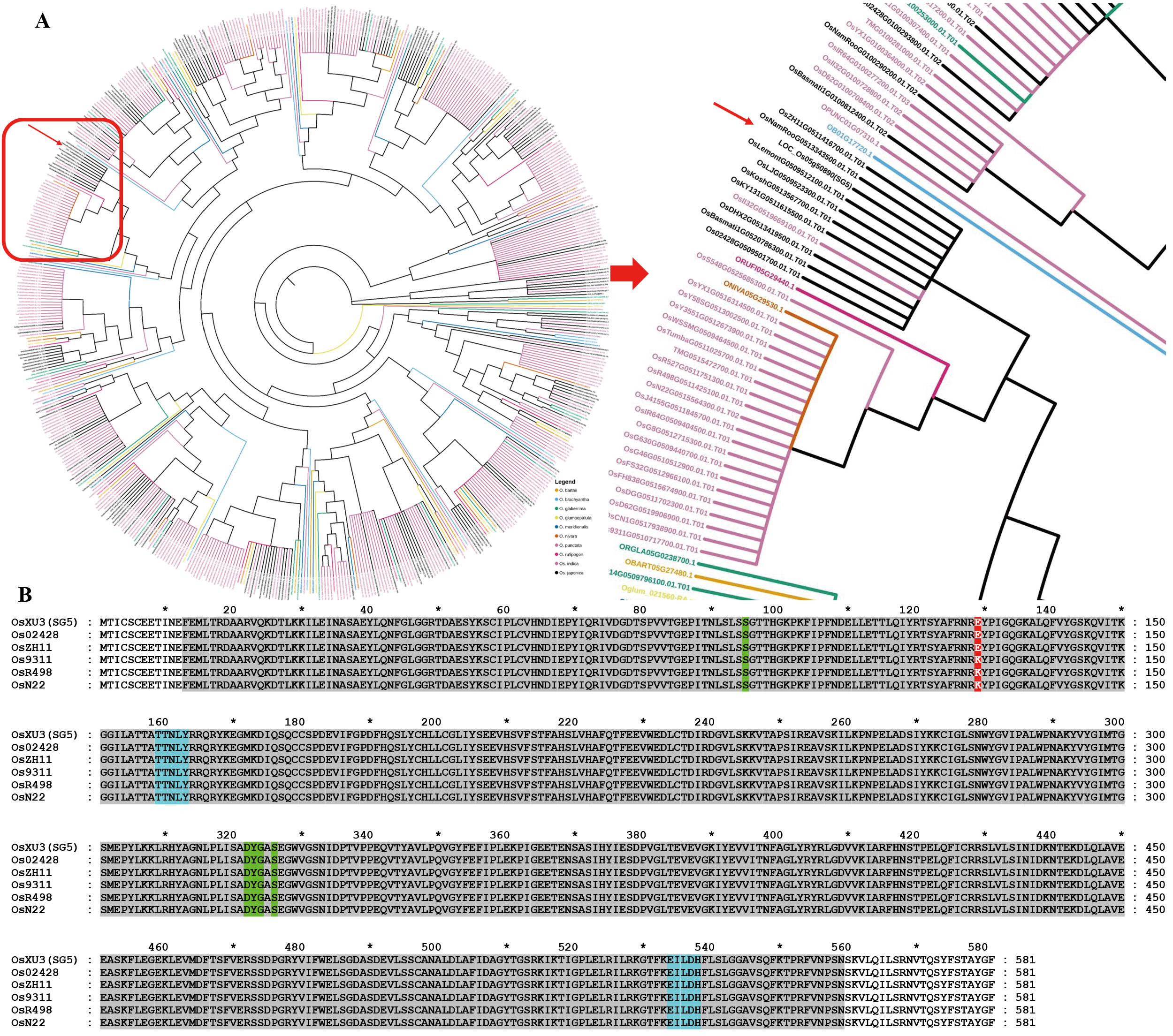
图5 SG5进化树分析及蛋白多序列比对 A:SG5蛋白进化树分析。红框:具有GH3-5结构域的种质资源;红色箭头指向:SG5目的蛋白。B:SG5蛋白多序列对分析。灰色框:GH3-5结构域;绿色框:茉莉酮酸酯;红色框:变异位点氨基酸;蓝色框:L-α-氨基酸位点。
Fig. 5. Phylogenetic tree and multiple sequence alignment analysis of SG5 in rice A, Phylogenetic tree analysis of SG5 in rice. Red frame, Rice resources with the GH3-5 domain. Red arrow, SG5 protein. B, Multiple sequence alignment of SG5 in rice. Gray frame, The domain of GH3-5. Green frame, The site of jasmonate. Blue frame, The site of L-alpha-amino acid.
| [1] | Xing Y Z, Zhang Q F. Genetic and molecular bases of rice yield[J]. Annual Review of Plant Biology, 2010, 61: 421-442. |
| [2] | Ren D Y, Ding C Q, Qian Q. Molecular bases of rice grain size and quality for optimized productivity[J]. Science Bulletin, 2023, 68(3): 314-350. |
| [3] | Theissen G, Melzer R. Molecular mechanisms underlying origin and diversification of the Angiosperm flower[J]. Annals of Botany, 2007, 100(3): 603-619. |
| [4] | Soltis D E, Chanderbali A S, Kim S, Buzgo M, Soltis P S. The ABC model and its applicability to basal angiosperms[J]. Annals of Botany, 2007, 100(2): 155-163. |
| [5] | Maoileidigh D S O, Graciet E, Wellmer F. Gene networks controlling Arabidopsis thaliana flower development[J]. New Phytologist, 2014, 201: 16-30. |
| [6] | Krizek B A, Fletcher J C. Molecular mechanisms of flower development: An armchair guide[J]. Nature Reviews Genetics, 2005, 6(9): 688-698. |
| [7] | Causier B, Schwarz-Sommer Z, Davies B. Floral organ identity: 20 years of ABCs[J]. Seminars in Cell & Developmental Biology, 2010, 21(1): 73-79. |
| [8] | 郭爽, 李云峰, 任德勇, 王增, 杜青, 何光华. 水稻内稃扭曲突变体palea distortion 1 (pd1) 的鉴定与基因定位[J]. 分子植物育种, 2011, 9(3): 256-260. |
| Guo S, Li Y F, Ren D Y, Wang Z, Du Q, He G H. Identification and gene mapping of a palea distortion 1 (pd1) mutant in rice (Oryza sativa L.)[J]. Molecular Plant Breeding, 2011, 9(3): 256-260. (in Chinese with English abstract) | |
| [9] | Jin Y, Luo Q, Tong H N, Wang A J, Cheng Z J, Tang J F, Li D Y, Zhao X F, Li X B, Wan J M, Jiao Y L, Chu C C, Zhu L H. An AT-hook gene is required for palea formation and floral organ number control in rice[J]. Developmental Biology, 2011, 359(2): 277-288. |
| [10] | 陈东, 毛毕刚, 潘银林, 彭彦, 韶也, 吴天昊, 赵炳然. 水稻颖花畸形突变体(gm(t))的遗传分析与基因定位[J]. 分子植物育种, 2019, 17(15): 5026-5031. |
| Chen D, Mao B G, Pan Y L, Peng Y, Shao Y, Wu T H, Zhao B R. Genetic analysis and gene mapping of glume malformation mutant(gm(t)) in rice[J]. Molecular Plant Breeding, 2019, 17(15): 5026-5031. (in Chinese with English abstract) | |
| [11] | 薛大伟. 两个水稻花器官突变体的遗传分析[D]. 北京: 中国农业科学院, 2006. |
| Xue D W. The genetic analysis of two floral organ mutants from rice[D]. Chinese Academy of Agricultural Sciences, 2006. Beijing: Chinese Academy of Agricultural Sciences, 2006. (in Chinese with English abstract) | |
| [12] | Jeon J S, Jang S, Lee S, Nam J, Kim C, Lee S H, Chung Y Y, Kim S R, Lee Y H, Cho Y G, An G. leafy hull sterile1 is a homeotic mutation in a rice MADS box gene affecting rice flower development[J]. The Plant Cell, 2000, 12(6): 871-884. |
| [13] | Agrawal G K, Abe K, Yamazaki M, Miyao A, Hirochika H. Conservation of the E-function for floral organ identity in rice revealed by the analysis of tissue culture-induced loss-of-function mutants of the OsMADS1 gene[J]. Plant Molecular Biology, 2005, 59(1): 125-135. |
| [14] | Prasad K, Parameswaran S, Vijayraghavan U. OsMADS1, a rice MADS‐box factor, controls differentiation of specific cell types in the lemma and palea and is an early‐acting regulator of inner floral organs[J]. The Plant Journal, 2005, 43(6): 915-928. |
| [15] | Hu Y, Liang W Q, Yin C S, Yang X L, Ping B Z, Li A X, Jia R, Chen M J, Luo Z J, Cai Q, Zhao X X, Zhang D B, Yuan Z. Interactions of OsMADS1 with floral homeotic genes in rice flower development[J]. Molecular plant, 2015, 8(9): 1366-1384. |
| [16] | Pelucchi N, Fornara F, Favalli C, Masiero S, Lago C, P E M, Colombo L, Kater M M. Comparative analysis of rice MADS-box genes expressed during flower development[J]. Sexual Plant Reproduction, 2002, 15(3): 113-122. |
| [17] | Wang K J, Tang D, Hong L L, Xu W Y, Huang J, Li M, Gu M H, Xue Y B, Cheng Z K. DEP and AFO regulate reproductive habit in rice[J]. PLoS Genetics, 2010, 6(1): e1000818. |
| [18] | Kobayashi K, Yasuno N, Sato Y, Yoda M, Yamazaki R, Kimizu M, Yoshida H, Nagamura Y, Kyozuka J. Inflorescence meristem identity in rice is specified by overlapping functions of three AP1/FUL-like MADS box genes and PAP2 a SEPALLATA MADS box gene[J]. The Plant Cell, 2012, 24(5): 1848-1859. |
| [19] | Yadav S R, Prasad K, Vijayraghavan U. Divergent regulatory OsMADS2 functions control size, shape and differentiation of the highly derived rice floret second-whorl organ[J]. Genetics, 2007, 176(1): 283-294. |
| [20] | Yao S G, Ohmori S, Kimizu M, Yoshida H. Unequal genetic redundancy of rice PISTILLATA orthologs, OsMADS2 and OsMADS4, in lodicule and stamen development[J]. Plant & Cell Physiology, 2008, 49(5): 853-857. |
| [21] | Yu X Q, Xia S S, Xu Q K, Cui Y J, Gong M, Zeng D L, Zhang Q, Shen L, Jiao G A, Gao Z Y, Hu J, Zhang G H, Zhu L, Guo L B, Ren D Y, Qian Q. ABNORMAL FLOWER AND GRAIN 1 encodes OsMADS6 and determines palea identity and affects rice grain yield and quality[J]. Science China Life Sciences, 2020, 63(2): 228-238. |
| [22] | Ohmori S, Kimizu M, Sugita M, Miyao A, Hirochika H, Uchida E, Nagato Y, Yoshida H. MOSAIC FLORAL ORGANS1, an AGL6 -like MADS box gene, regulates floral organ identity and meristem fate in rice[J]. The Plant Cell, 2009, 21(10): 3008-3025. |
| [23] | Zhang J, Nallamilli B R, Mujahid H, Peng Z H. OsMADS6 plays an essential role in endosperm nutrient accumulation and is subject to epigenetic regulation in rice (Oryza sativa)[J]. The Plant Journal, 2010, 64(4): 604-617. |
| [24] | Duan Y L, Xing Z, Diao Z J, Xu W Y, Li S P, Du X Q, Wu G H, Wang C L, Lan T, Meng Z, Liu H Q, Wang F, Wu W R, Xue Y B. Characterization of Osmads6-5, a null allele, reveals that OsMADS6 is a critical regulator for early flower development in rice (Oryza sativa L.)[J]. Plant Molecular Biology, 2012, 80(4-5): 429-442. |
| [25] | Wang H H, Zhang L, Cai Q, Hu Y, Jin Z M, Zhao X X, Fan W, Huang Q M, Luo Z J, Chen M J, Zhang D B, Yuan Z. OsMADS32 interacts with PI-like proteins and regulates rice flower development[J]. Journal of Integrative Plant Biology, 2015, 57(5): 504-513. |
| [26] | Luo Q, Zhou K D, Zhao X F, Zeng Q C, Xia H A, Zhai W X, Xu J C, Wu X J, Yang H S, Zhu L H. Identification and fine mapping of a mutant gene for palealess spikelet in rice[J]. Planta, 2005, 221(2): 222-230. |
| [27] | Xiao H, Tang J F, Li Y F, Wang W M, Li X B, Jin L, Xie R, Luo H F, Zhao X F, Meng Z, He G H, Zhu L H. STAMENLESS 1, encoding a single C2H2 zinc finger protein, regulates floral organ identity in rice[J]. The Plant Journal, 2009, 59(5): 789-801. |
| [28] | Yan D W, Zhang X M, Zhang L, Ye S H, Zeng L J, Liu J Y, Li Q, He Z H. CURVED CHIMERIC PALEA 1 encoding an EMF1-like protein maintains epigenetic repression of OsMADS58in rice palea development[J]. The Plant Journal, 2015, 82(1): 12-24. |
| [29] | Zheng M, Wang Y H, Wang Y L, Wang C M, Ren Y L, Lv J, Peng C, Wu T, Liu K, Zhao S L, Liu X, Guo X P, Jiang L, Terzaghi W, Wan J M. DEFORMED FLORAL ORGAN1 (DFO1) regulates floral organ identity by epigenetically repressing the expression of OsMADS58 in rice (Oryza sativa)[J]. The New Phytologist, 2015, 206(4): 1476-1490. |
| [30] | Yuan Z, Gao S, Xue D W, Luo D, Li L T, Ding S Y, Yao X, Wilson Z A, Qian Q, Zhang D B. RETARDED PALEA1 controls palea development and floral zygomorphy in rice[J]. Plant Physiology, 2009, 149(1): 235-244. |
| [31] | Zeng D D, Qin R, Alamin M, Liang R, Yang C C, Jin X L, Shi C H. DBOP specifies palea development by suppressing the expansion of the margin of palea in rice[J]. Genes and Genomics, 2016, 38(11): 1095-1103. |
| [32] | Lee D Y, Lee J, Moon S, Park S Y, An G. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem[J]. The Plant Journal, 2007, 49(1): 64-78. |
| [33] | Ren D Y, Li Y F, Zhao F M, Sang X C, Shi J Q, Wang N, Guo S, Ling Y H, Zhang C W, Yang Z L, He G H. MULTI-FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice[J]. Plant Physiology, 2013, 162(2): 872-884. |
| [34] | Ren D Y, Rao Y C, Wu L W, Xu Q K, Li Z Z, Yu H P, Zhang Y, Leng Y J, Hu J, Zhu L, Gao Z Y, Dong G J, Zhang G H, Guo L B, Zeng D L, Qian Q. The pleiotropic ABNORMAL FLOWER AND DWARF1 affects plant height, floral development and grain yield in rice[J]. Journal of Integrative Plant Biology, 2016, 58(6): 529-539. |
| [35] | Yan D W, Zhou Y, Ye S H, Zeng L J, Zhang X M, He Z H. BEAK-SHAPED GRAIN 1/TRIANGULAR HULL 1, a DUF640 gene, is associated with grain shape, size and weight in rice[J]. Science China Life Sciences, 2013, 56(3): 275-283. |
| [36] | Livak K J, Schmittgen T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCTmethod[J]. Methods, 2001, 25(4): 402-408. |
| [37] | 王忠, 顾蕴洁, 高煜珠. 水稻开颖机理的探讨: Ⅲ.浆片的结构及其在开颖过程中内含物的变化[J]. 作物学报, 1991, 17(2): 96-101, 161-162. |
| Wang Z, Gu Y J, Gao Y Z. Studies on the mechanism of the anthesis of rice Ⅲ.Structure of the lodicule and changes of its contents during flowering[J]. Acta Agronomica Sinica, 1991, 17(2): 96-101, 161-162. (in Chinese with English abstract) | |
| [38] | 秦鹏. 水稻颖壳发育基因hzp和ug1的研究[D]. 长沙: 湖南农业大学, 2021. |
| Qin P. The study on genes hzp and ug1 controlling rice glume development[D]. Changsha: Hunan Agricultural University, 2021. (in Chinese with English abstract) | |
| [39] | Terol J, Domingo C, Talón M. The GH3 family in plants: Genome wide analysis in rice and evolutionary history based on EST analysis[J]. Gene, 2006, 371(2): 279-290. |
| [40] | Riemann M, Riemann M, Takano M. Rice JASMONATE RESISTANT 1 is involved in phytochrome and jasmonate signalling[J]. Plant, Cell & Environment, 2008, 31(6): 783-792. |
| [41] | Xiao Y G, Chen Y, Charnikhova T, Mulder P P J, Heijmans J, Hoogenboom A, Agalou A, Michel C, Morel J B, Dreni L, Kater M M, Bouwmeester H, Wang M, Zhu Z, Ouwerkerk P B F. OsJAR1 is required for JA-regulated floret opening and anther dehiscence in rice[J]. Plant Molecular Biology, 2014, 86(1/2): 19-33. |
| [42] | Jain M, Kaur N, Tyagi A K, Khurana J P. The auxin-responsive GH3 gene family in rice (Oryza sativa)[J]. Functional & Integrative Genomics, 2006, 6(1): 36-46. |
| [43] | Zhao S Q, Xiang J J, Xue H W. Studies on the rice LEAF INCLINATION1 (LC1), an IAA-amido synthetase, reveal the effects of auxin in leaf inclination control[J]. Molecular Plant, 2013, 6(1): 174-187. |
| [44] | Du H, Wu N, Fu J, Wang S P, Li X H, Xiao J H, Xiong L Z. A GH3 family member, OsGH3-2, modulates auxin and abscisic acid levels and differentially affects drought and cold tolerance in rice[J]. Journal of Experimental Botany, 2012, 63(18): 6467-6480. |
| [45] | Fu J, Yu H H, Li X H, Xiao J H, Wang S P. Rice GH3 gene family: Regulators of growth and development[J]. Plant Signaling Behavior, 2011, 6(4): 570-574. |
| [46] | Kong W L, Zhong H, Deng X X, Gautam M, Gong Z Y, Zhang Y, Zhao G Q, Liu C, Li Y S. Evolutionary analysis of GH3 genes in six Oryza Species/Subspecies and their expression under salinity stress in Oryza sativa ssp. japonica[J]. Plants, 2019, 8(2): 30. |
| [47] | Mao C J, He J M, Liu L, Deng Q M, Yao X F, Liu C M, Qiao Y L, Li P, Ming F. OsNAC2 integrates auxin and cytokinin pathways to modulate rice root development[J]. Plant Biotechnology Journal, 2020, 18(2): 429-442. |
| [48] | Ding X H, Cao Y L, Huang L L, Zhao J, Xu C G, Li X H, Wang S P. Activation of the Indole-3-Acetic Acid-Amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice[J]. The Plant Cell, 2008, 20(1): 228-240. |
| [49] | Yadav S R, Khanday I, Majhi B B, Veluthambi K, Vijayraghavan U. Auxin-responsive OsMGH3, a common downstream target of OsMADS1 and OsMADS6, controls rice floret fertility[J]. Plant and Cell Physiology, 2011, 52(12): 2123-2135. |
| [50] | Fukumoto K, Alamgir K M, Yamashita Y, Mori I C, Matsuura H, Galis I. Response of rice to insect elicitors and the role of OsJAR1 in wound and herbivory-induced JA-Ile accumulation[J]. Journal of Integrative Plant Biology, 2013, 55(8): 775-784. |
| [51] | Song S S, Qi T C, Huang H, Ren Q C, Wu D W, Chang C Q, Peng W, Liu Y, Peng J R, Xie D X. The jasmonate-ZIM domain proteins interact with the R2R3-MYB transcription factors MYB21 and MYB24 to affect jasmonate-regulated stamen development in Arabidopsis[J]. The Plant Cell, 2011, 23(3): 1000-1013. |
| [52] | Wakuta S, Suzuki E, Saburi W, Matsuura H, Nabeta K, Imai R, Matsui H. OsJAR1 and OsJAR2 are jasmonyl-L-isoleucine synthases involved in wound- and pathogen-induced Jasmonic acid signalling[J]. Biochemical and Biophysical Research Communications, 2011, 409: 634-639. |
| [53] | Zhang S N, Wang S K, Xu Y X, Yu C L, Shen C J, Qian Q, Geisler M, Jiang D A, Qi Y H. The auxin response factor, OsARF19, controls rice leaf angles through positively regulating OsGH3-5 and OsBRI1[J]. Plant, Cell and Environment, 2015, 38: 638-654. |
| [54] | Ding W Y, Gou Y J, Li Y J, Li J J, Fang Y D, Liu X P, Zhu X Y, Ye R J, Heng Y Q, Wang H Y, Shen R X. A jasmonate-mediated regulatory network modulates diurnal floret opening time in rice[J]. New phytologist, 2024, 244(1): 176-191. |
| [55] | 王忠, 顾蕴洁, 高煜珠. 水稻开颖机理的探讨: V.不育系与可育系浆片和花丝结构的比较[J]. 作物学报, 1994, 20(1): 13-17, 129-131. |
| Wang Z, Gu Y J, Gao Y Z. Studies on the mechanism of the anthesis of rice V. comparison of lodicule and filament structure between sterile line and fertile line[J]. Acta Agronomica Sinica, 1994, 20(1): 13-17, 129-131. (in Chinese with English abstract) |
| [1] | 杨大兵, 杜雪树, 李进波, 夏明元, 胡亮, 石桓, 万丙良. 水稻抽穗期调控的分子机理及育种应用进展[J]. 中国水稻科学, 2026, 40(2): 145-154. |
| [2] | 倪晨, 张家豪, 朱昌进, 徐继伟, 胡秋倩, 霍中洋, 戴其根, 许轲, 李国辉. 水稻源流库形成与调控及其影响因素研究进展[J]. 中国水稻科学, 2026, 40(2): 155-170. |
| [3] | 王梦宁, 谢可冉, 高逖, 王真梅, 熊栋梁, 崔克辉. 高温对水稻粒重形成的影响及其栽培调控研究进展[J]. 中国水稻科学, 2026, 40(2): 171-180. |
| [4] | 罗肖郧, 郑兴飞, 彭宣国, 余启芝, 董华林, 殷得所, 王红波, 胡建林, 薛莲, 胡鹏, 徐得泽. 水稻抗倒伏研究:现状、挑战与未来方向[J]. 中国水稻科学, 2026, 40(2): 181-195. |
| [5] | 段敏, 谢留杰, 岳雅妮, 黄善军. 基于CRISPR/Cas9技术创制优质香味粳稻品系[J]. 中国水稻科学, 2026, 40(2): 235-243. |
| [6] | 张梦柯, 陆佳雨, 何金, 许学, 吴爽, 王沛然, 陈若凡, 金青, 汪秀峰. 水稻野败型三系杂交种纯度检测功能标记的开发与应用[J]. 中国水稻科学, 2026, 40(2): 244-252. |
| [7] | 李兴沂, 陈玲, 邵建韬, 肖素勤, 李金璐, 付惠仙, 殷富有, 张建红, 程在全, 刘丽. 水稻产量与淀粉品质协同调控的分子遗传研究进展[J]. 中国水稻科学, 2026, 40(1): 1-17. |
| [8] | 岳轩宇, 谢文亚, 冯志明, 陈宗祥, 胡珂鸣, 左示敏. OsERF93参与调控水稻纹枯病抗性的研究[J]. 中国水稻科学, 2026, 40(1): 37-50. |
| [9] | 王轶欣, 林参, 马刘洋, 陈龙, 奉保华, 倪深, 魏祥进, 贺记外, 陈天晓. 谷丙转氨酶基因OsAlaAT4调控水稻氮素吸收和产量[J]. 中国水稻科学, 2026, 40(1): 51-60. |
| [10] | 黄奇娜, 姜鸿瑞, 杨婕, 于坤宇, 杨长登, 梁燕. 种子休眠基因Sdr4的生物信息学分析与分子标记开发和应用[J]. 中国水稻科学, 2026, 40(1): 61-71. |
| [11] | 程朝平, 何旎清, 白康呈, 林少俊, 黄凤凰, 刘军化, 程祖锌, 黄成志, 杨德卫. 聚合稻瘟病抗性基因Pigm-1和Pid2的水稻三系不育系福梦A的选育与利用[J]. 中国水稻科学, 2026, 40(1): 72-84. |
| [12] | 刘亚萍, 董译词, 郑君妍, 邱绚, 刘鹏程, 叶亚峰, 刘斌美, 陈析丰, 马伯军. 水稻类病变早衰突变体lmes7的鉴定与基因精细定位[J]. 中国水稻科学, 2026, 40(1): 85-94. |
| [13] | 谢世民, 周誉株, 薛晓迪, 朱广飞, 孙良, 陈建能. 水稻钵苗取栽协同作业式移栽机构设计与试验[J]. 中国水稻科学, 2026, 40(1): 131-144. |
| [14] | 王娟, 吴丽娟, 洪海波, 姚志文, 王磊, 鄂志国. 水稻泛素结合酶E2的生物学功能研究进展[J]. 中国水稻科学, 2025, 39(6): 744-750. |
| [15] | 陶士博, 许娜, 徐正进, 刘畅, 徐铨. 水稻发芽期耐冷基因Cold6的克隆[J]. 中国水稻科学, 2025, 39(6): 751-759. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||