
中国水稻科学 ›› 2023, Vol. 37 ›› Issue (5): 478-485.DOI: 10.16819/j.1001-7216.2023.220907
李景芳1, 温舒越2, 赵利君2, 陈庭木1, 周振玲1, 孙志广1, 刘艳1, 陈海元3, 张云辉3, 迟铭1, 邢运高1, 徐波1, 徐大勇1, 王宝祥1,*( )
)
收稿日期:2022-09-28
修回日期:2022-12-19
出版日期:2023-09-10
发布日期:2023-09-13
通讯作者:
*email:基金资助:
LI Jingfang1, WEN Shuyue2, ZHAO Lijun2, CHEN Tingmu1, ZHOU Zhenling1, SUN Zhiguang1, LIU Yan1, CHEN Haiyuan3, ZHANG Yunhui3, CHI Ming1, XING Yungao1, XU Bo1, XU Dayong1, WANG Baoxiang1,*( )
)
Received:2022-09-28
Revised:2022-12-19
Online:2023-09-10
Published:2023-09-13
Contact:
*email:摘要:
【目的】 为了促进耐盐香稻育种,利用CRISPR/Cas9系统对粳稻品种连粳11的Badh2和OsRR22基因进行编辑,以期快速获得一批不含有转基因成分且具有耐盐性和香味的纯合水稻材料。【方法】 根据Badh2和OsRR22基因序列中编辑位点的敲除效率设计靶位点,构建pH-Ubi-Cas9-Badh2-OsRR22敲除载体,利用农杆菌介导法转入受体品种连粳11中。对转基因后代进行潮霉素和Cas9标记PCR检测以及靶基因测序,获得无外源基因插入的badh2-osrr22纯合株系,并对后代种子性状和苗期耐盐性进行分析。【结果】 T2代成熟种子中2-AP含量较背景材料连粳11显著增加,千粒重、粒长、粒宽无明显变化;128 mmol/L氯化钠处理14 d后株系21-30较连粳11苗高增加15.2%,苗鲜质量增加45.2%,苗干质量增加13.2%。【结论】 利用CRISPR/Cas9技术对Badh2和OsRR22基因编辑获得能稳定遗传且无外源基因插入的耐盐香稻材料,加快了水稻多性状聚合的选育进程。
李景芳, 温舒越, 赵利君, 陈庭木, 周振玲, 孙志广, 刘艳, 陈海元, 张云辉, 迟铭, 邢运高, 徐波, 徐大勇, 王宝祥. 基于CRISPR/Cas9技术创制耐盐香稻[J]. 中国水稻科学, 2023, 37(5): 478-485.
LI Jingfang, WEN Shuyue, ZHAO Lijun, CHEN Tingmu, ZHOU Zhenling, SUN Zhiguang, LIU Yan, CHEN Haiyuan, ZHANG Yunhui, CHI Ming, XING Yungao, XU Bo, XU Dayong, WANG Baoxiang. Development of Aromatic Salt-tolerant Rice Based on CRISPR/Cas9 Technology[J]. Chinese Journal OF Rice Science, 2023, 37(5): 478-485.
| 引物名称 Primer name | 引物序列 Primer sequence(5’-3’) | 用途 Usage | |
|---|---|---|---|
| Badh2-S1-F | GGCATGGTGGAAAAAGTCCTATAG | 构建Badh2敲除载体 Construction of Badh2 knockout vector | |
| Badh2-S1-R | AAACCTATAGGACTTTTTCCACCA | ||
| Badh2-F | CCTTTGTCATCACACCCTGG | Badh2敲除靶点测序鉴定 | |
| Badh2-R | TAACTGCCTTCCTTGCCACG | Sequencing for Badh2 knockout target | |
| OsRR22-S1-F | GGCACTCAACTGACCATACAACTC | 构建OsRR22敲除载体 | |
| OsRR22-S1-R | AAACGAGTTGTATGGTCAGTTGAG | Construction of OsRR22 knockout vector | |
| OsRR22-F | TAGGAGGAAGTTCGGTAATCGT | OsRR22敲除靶点测序鉴定 | |
| OsRR22-R | ACATTTTCCCTGGTGAGTTTCT | Sequencing for OsRR22 knockout target | |
| Cas9-F | CCTTCTGTGTGGTCTGGTTC | 载体检测 | |
| Cas9-R | AGACAATCACCCCCTGGAAC | Vector assay | |
| Hyg-F | CTATTTCTTTGCCCTCGGAC | 转基因检测 | |
| Hyg-R | ATGCCTGAACTCACCGCGAC | Transgenic detection | |
| qPCR SOS1-F | ATTGCAGCTGAGCATGTACG | 实时荧光定量PCR | |
| qPCR SOS1-R | AGAGCTTGCTTTCGTGTGAC | Quantitative real-time PCR | |
| qPCR OsMYB2-F | GGGCTGAAACGCACAGGCAAGA | 实时荧光定量PCR | |
| qPCR OsMYB2-R | CTGCTTGGCGTGCTTCTGC | Quantitative real-time PCR | |
| qPCR OsNHX1-F | GTGACAGACCTGGCAAATCC | 实时荧光定量PCR | |
| qPCR OsNHX1-R | TCGACACAGCTCCTCTCATC | Quantitative real-time PCR | |
| qPCR OsHKT1;1-F | TGCCAGAAGTTGTTGAAGCC | 实时荧光定量PCR | |
| qPCR OsHKT1;1-R | CCCAGGAACATCACCAGGAT | Quantitative real-time PCR | |
| qPCR OsP5CS1-F | GTCAGAGTGGACTGATGGCT | 实时荧光定量PCR | |
| qPCR OsP5CS1-R | GCCTTTCTAGTGCTGATGGC | Quantitative real-time PCR | |
| UBIQUITIN-F | GCTCCGTGGCGGTATCAT | 实时荧光定量PCR | |
| UBIQUITIN-R | CGGCAGTTGACAGCCCTAG | Quantitative real-time PCR | |
表1 本研究中使用的引物
Table 1. Primers used in this study.
| 引物名称 Primer name | 引物序列 Primer sequence(5’-3’) | 用途 Usage | |
|---|---|---|---|
| Badh2-S1-F | GGCATGGTGGAAAAAGTCCTATAG | 构建Badh2敲除载体 Construction of Badh2 knockout vector | |
| Badh2-S1-R | AAACCTATAGGACTTTTTCCACCA | ||
| Badh2-F | CCTTTGTCATCACACCCTGG | Badh2敲除靶点测序鉴定 | |
| Badh2-R | TAACTGCCTTCCTTGCCACG | Sequencing for Badh2 knockout target | |
| OsRR22-S1-F | GGCACTCAACTGACCATACAACTC | 构建OsRR22敲除载体 | |
| OsRR22-S1-R | AAACGAGTTGTATGGTCAGTTGAG | Construction of OsRR22 knockout vector | |
| OsRR22-F | TAGGAGGAAGTTCGGTAATCGT | OsRR22敲除靶点测序鉴定 | |
| OsRR22-R | ACATTTTCCCTGGTGAGTTTCT | Sequencing for OsRR22 knockout target | |
| Cas9-F | CCTTCTGTGTGGTCTGGTTC | 载体检测 | |
| Cas9-R | AGACAATCACCCCCTGGAAC | Vector assay | |
| Hyg-F | CTATTTCTTTGCCCTCGGAC | 转基因检测 | |
| Hyg-R | ATGCCTGAACTCACCGCGAC | Transgenic detection | |
| qPCR SOS1-F | ATTGCAGCTGAGCATGTACG | 实时荧光定量PCR | |
| qPCR SOS1-R | AGAGCTTGCTTTCGTGTGAC | Quantitative real-time PCR | |
| qPCR OsMYB2-F | GGGCTGAAACGCACAGGCAAGA | 实时荧光定量PCR | |
| qPCR OsMYB2-R | CTGCTTGGCGTGCTTCTGC | Quantitative real-time PCR | |
| qPCR OsNHX1-F | GTGACAGACCTGGCAAATCC | 实时荧光定量PCR | |
| qPCR OsNHX1-R | TCGACACAGCTCCTCTCATC | Quantitative real-time PCR | |
| qPCR OsHKT1;1-F | TGCCAGAAGTTGTTGAAGCC | 实时荧光定量PCR | |
| qPCR OsHKT1;1-R | CCCAGGAACATCACCAGGAT | Quantitative real-time PCR | |
| qPCR OsP5CS1-F | GTCAGAGTGGACTGATGGCT | 实时荧光定量PCR | |
| qPCR OsP5CS1-R | GCCTTTCTAGTGCTGATGGC | Quantitative real-time PCR | |
| UBIQUITIN-F | GCTCCGTGGCGGTATCAT | 实时荧光定量PCR | |
| UBIQUITIN-R | CGGCAGTTGACAGCCCTAG | Quantitative real-time PCR | |
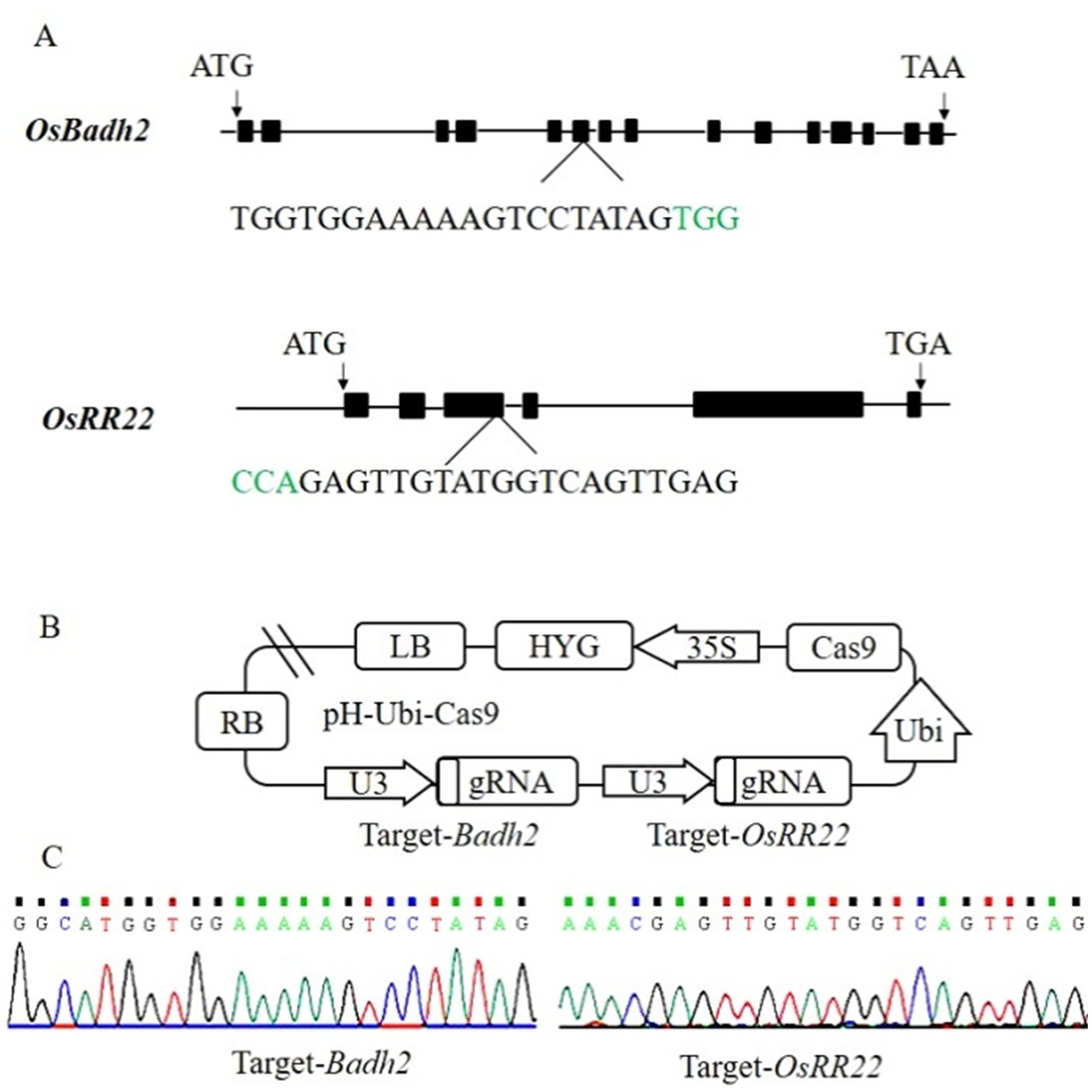
图1 Badh2和OsRR22基因靶点和载体构建 A―Badh2和OsRR22基因结构及靶位点位置信息,绿色碱基代表PAM序列;B―Badh2和OsRR22基因靶点CRISPR/Cas9载体构建;C―表达载体中靶位点测序结果。
Fig. 1. Badh2 and OsRR22 target sites and CRISPR/Cas9 vector construction. A, Badh2 and OsRR22 structure and target site. The PAM motif is shown in green. B, Construction of CRISPR/Cas9 vector for Badh2 and OsRR22 gene target; C, Target site sequencing results in the expression vector.

图2 T1代部分植株Cas9标记鉴定结果 M―DNA标记;“+”―阳性对照;“−”―阴性对照;1~9为T1代部分植株。Cas9标记目标片段大小为906 bp。
Fig. 2. Identification of Cas9 markers in some T1 plants. M, DNA Marker; “+”, Positive control; “−”, Negative control; Lanes 1 to 9, Some plants of T1. The size of the Cas9 marker target fragment is 906 bp.

图3 T1代纯合株系Badh2和OsRR22突变类型 红色箭头表示突变位置。“+”代表插入;“-”代表缺失。黑色箭头表示氨基酸位置。
Fig. 3. Mutation types at the Badh2 and OsRR22 loci of the homozygous lines in T1. Red arrows indicate mutation positions. “+” means insertion; “-” means deletion. Black arrows indicate amino acid positions.

图4 T2代两个纯合株系21-30、21-31苗期耐盐性鉴定 数据用平均数±标准差表示。*和**分别表示在5%和1%水平上差异显著(t检验)。
Fig. 4. Salt tolerance identification of T2 homozygous lines 21-30, 21-31 at the seedling stage. Values are shown as mean ± SD. *, ** Significantly different at 0.05 and 0.01 levels by t-test, respectively.
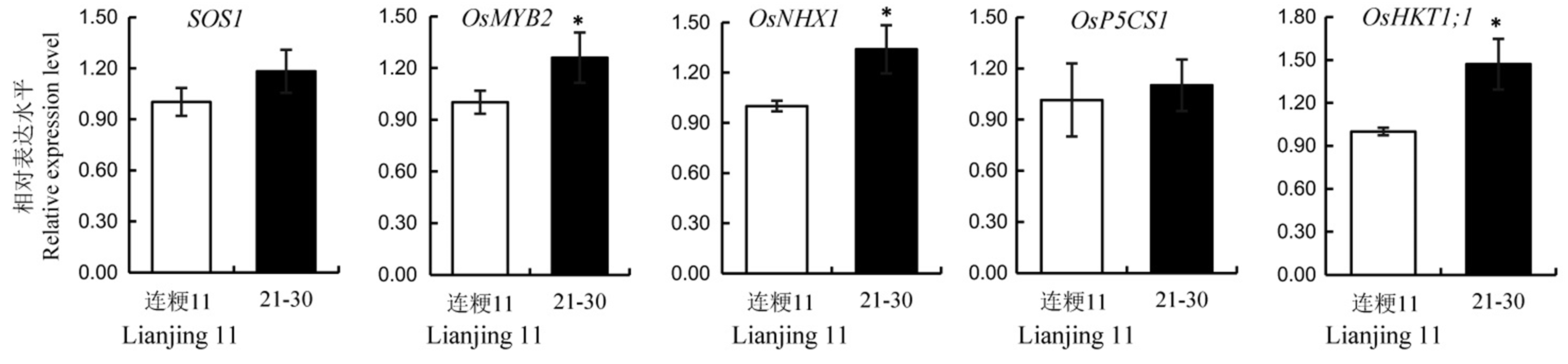
图5 野生型连粳11与T2代21-30株系中耐盐相关基因的相对表达量 数值用平均数±标准差表示。*表示在5%水平上差异显著(t检验)。
Fig. 5. Relative expression levels of salt resistance related genes in the wild-type and its T2 line 21-30. Values are shown as mean ± SD.*Significantly different at 0.05 level by t-test.
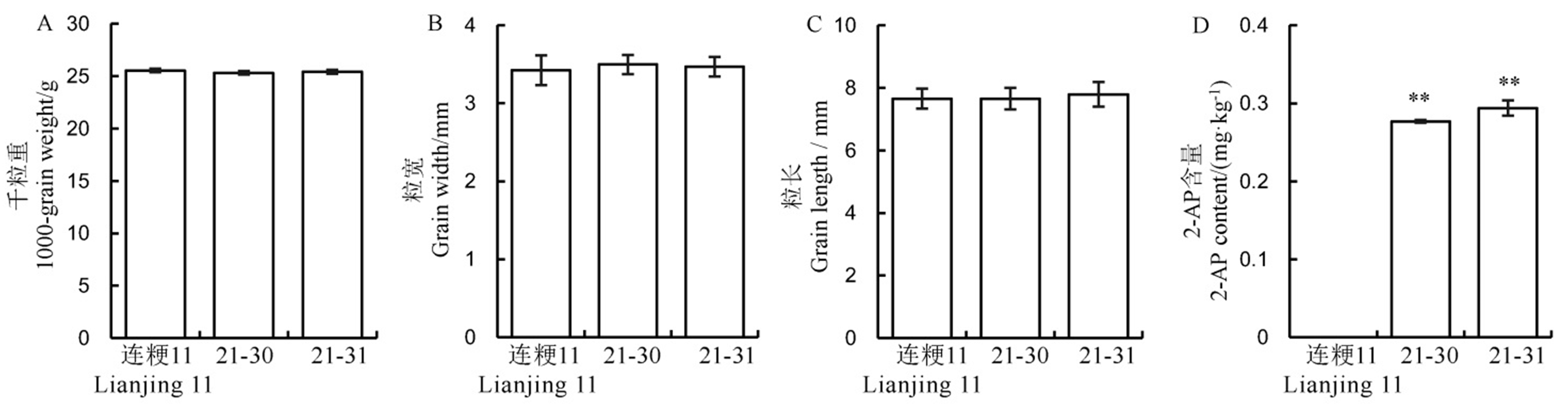
图6 野生型和转基因阳性株系中水稻籽粒和香味相关性状表现 数值用平均数±标准差表示。**表示在1%水平上差异显著 (t 检验)。
Fig. 6. Performance of rice yield and fragrance related traits in the wild-type and its positive transgenic lines. Values are shown as mean ± SD.** Significantly different at 0.01 level by t-test.
| [1] | El Mahi H, Perez-Hormaeche J, de Luca A, Villalta I, Espartero J, Gamez-Arjona F, Fernandez J L, Bundo M, Mendoza I, Mieulet D. A critical role of sodium flux via the plasma membrane Na+/H+ exchanger SOS1 in the salt tolerance of rice[J]. Plant Physiology, 2019, 180(2): 1046-1065. |
| [2] | Yang A, Dai X Y, Zhang W H. A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice[J]. Journal of Experimental Botany, 2012, 63(7): 2541-2556. |
| [3] | Wang R, Jing W, Xiao L G, Jin Y K, Shen L K, Zhang W H. The rice high-affinity potassium pransporter1;1 is involved in salt tolerance and regulated by an MYB-type transcription factor[J]. Plant Physiology, 2015, 168(3): 1076-1090. |
| [4] | Liu S P, Zheng L Q, Xue Y H, Zhang Q, Wang L, Shuo H X. Overexpression of OsVP1 and OsNHX1 increases tolerance to drought and salinity in rice[J]. Journal of Plant Biology, 2010, 53: 444-452. |
| [5] | 张霞, 唐维, 刘嘉, 刘永胜. 过量表达水稻 OsP5CS1 和OsP5CS2 基因提高烟草脯氨酸的生物合成及其非生物胁迫抗性[J]. 应用与环境生物学报, 2014, 20(4): 717-722. |
| Zhang X, Tang W, Liu J, Liu Y S. Co-expression of rice OsP5CS1 and OsP5CS2genes in transgenic tobacco resulted in elevated proline biosynthesis and enhanced abiotic stress tolerance[J]. Chinese Journal of Applied & Environmental Biology, 2014, 20(4): 717-722. (in Chinese with English abstract) | |
| [6] | Li H W, Wang X P. Overexpression of the trehalose- 6-phosphate synthase gene OsTPS1 enhances abiotic stress tolerance in rice[J]. Planta, 2011, 234(5): 1007-1018. |
| [7] | Lan T, Zheng Y L, Su Z L, Yu S B, Song H B, Zheng X Y, Lin G G, Wu W R. OsSPL10, a SBP-box gene, plays a dual role in salt tolerance and trichome formation in rice (Oryza sativa L.)[J]. G3: Genes, Genomes, Genetics, 2019, 9(12): 4107-4114. |
| [8] | Yang C, Ma B, He S J, Xiong Q, Duan K X, Yin C C, Chen H, Lu X, Chen S Y, Zhang J S. MAOHUZI6/ETHYLENE INSENSITIVE3-LIKE1 and ETHYLENE INSENSITIVE3-LIKE2 regulate ethylene response of roots and coleoptiles and negatively affect salt tolerance in rice[J]. Plant Physiology, 2015, 169(1): 148-165. |
| [9] | Koh S, Lee S C, Kim M K, Koh J H, Lee S, An G, Choe S, Kim S R. T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses[J]. Plant Molecular Biology, 2007, 65(4): 453-466. |
| [10] | Takagi H, Tamiru M, Abe A, Yoshida K, Uemura A, Yaegashi H, Obara T, Oikawa K, Utsushi H, Kanzaki E, Mitsuoka C, Natsume S, Kosugi S, Kanzaki H, Matsumura H, Urasaki N, Kamoun S, Terauchi R. MutMap accelerates breeding of a salt-tolerant rice cultivar[J]. Nature Biotechnology, 2015, 33(5): 445-449. |
| [11] | Zhang A N, Liu Y, Wang F M, Li T F, Chen Z H, Kong D Y, Bi J G, Zhang F Y, Luo X X, Wang J H, Tang J J, Yu X Q, Liu G L, Luo L J. Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene[J]. Molecular Breeding, 2019, 39: 47. |
| [12] | 李晶岚, 陈鑫欣, 石翠翠, 刘方惠, 孙静, 葛荣朝. OsRPK1基因过表达和RNA干涉对水稻苗期耐盐性的影响[J]. 作物学报, 2020, 46(8): 1217-1224. |
| Li J L, Chen X X, Shi C C, Liu F H, Sun J, Ge R C. Effects of OsRPK1 gene overexpression and RNAi on the salt-tolerance at seedling stage in rice[J]. Acta Agronomica Sinica, 2020, 46(8): 1217-1224. (in Chinese with English abstract) | |
| [13] | 魏晓东, 张亚东, 赵凌, 路凯, 宋雪梅, 王才林. 稻米香味物质2-乙酰-1-吡咯啉的形成及其影响因素[J]. 中国水稻科学, 2022, 36 (2): 131-138. |
| Wei X D, Zhang Y D, Zhao L, Lu K, Song X M, Wang C L. Research progress in biosynthesis and influencing factors of 2-acetyl-1-pyrroline in fragrant rice[J]. Chinese Journal of Rice Science, 2022, 36(2): 131-138. (in Chinese with English abstract) | |
| [14] | Jiang M, Liu Y H, Li R Q, Li S, Tan Y Y, Huang J Z, Shu Q Y. An inositol 1, 3, 4, 5, 6-pentakisphosphate 2-kinase 1 mutant with a 33-nt deletion showed enhanced tolerance to salt and drought stress in rice[J]. Plants, 2021, 10(1): 23. |
| [15] | Bradbury L M, Fitzgerald T L, Henry R J, Jin Q S, Waters D L. The gene for fragrance in rice[J]. Plant Biotechnology Journal, 2005, 3(3): 363-370. |
| [16] | Chen S H, Yang Y, Shi W W, Ji Q, He F, Zhang Z D, Heng Z K, Liu X N, Xu M L. Badh2, encoding betaine ldehyde dehydrogenase, inhibits the biosynthesis of -acetyl-1-pyrroline, a major component in rice fragrance[J]. Plant Cell, 2008, 20(7): 1850-1861. |
| [17] | Bradbury L M, Gillies S A, Brushett D J, Waters D L, Henry R J. Inactivation of an amino aldehyde dehydrogenase is responsible for fragrance in rice[J]. Plant Molecular Biology, 2008, 68(4-5): 439-449. |
| [18] | Shan Q W, Zhang Y, Chen K L, Zhang K, Gao C X. Creation of fragrant rice by targeted knockout of the OsBadh2 gene using TALEN technology[J]. Plant Biotechnology Journal, 2015, 13(6): 791-800. |
| [19] | Hui S Z, Li H J, Mawia A M, Zhou L, Cai J Y, Ahmad S, Lai C K, Wang J X, Jiao G A, Xie L H, Shao G N, Sheng Z H, Tang S Q, Wang J L, Wei X J, Hu S K, Hu P S. Production of aromatic three-line hybrid rice using novel alleles of BADH2[J]. Plant Biotechnology Journal, 2022, 20(1): 59-74. |
| [20] | Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu L. Targeted mutagenesis in rice using CRISPR-Cas system[J]. Cell Research, 2013, 23(10): 1233-1236. |
| [21] | Yoshida S, Forno D A, Cook J H, Gomez K A. Laboratory Manual for Physiological Studies of Rice[M]. International Rice Research Institute, Philippines, 1976: 61-66. |
| [22] | Liu Y, Wang B X, Li J, Song Z Q, Lu B G, Chi M, Yang B, Liu J B, Lam Y W, Li J X, Xu D Y. Salt-response analysis in two rice cultivars at seedling stage[J]. Acta Physiologiae Plantarum, 2017, 39: 215. |
| [23] | Wang B X, Xu B, Liu Y, Li J F, Sun Z G, Chi M, Xing Y G, Yang B, Li J, Liu J B, Lu B G, Xu D Y, Bello B K. A novel mechanism of the signaling cascade associated with the SAPK10-bZIP20-NHX1 synergistic interaction to enhance tolerance of plant to abiotic stress in rice[J]. Plant Science, 2022, 323: 111393. |
| [24] | Xu Y, Lin Q P, Li X F, Wang F J, Chen Z H, Wang J, Li W Q, Fan F J, Tao Y J, Jiang Y J, Wei X D, Zhang R, Yang J, Gao C X. Fine-tuning the amylose content of rice by precise base editing of the Wx gene[J]. Plant Biotechnology Journal, 2021, 19(1): 11-13. |
| [25] | Liu X X, Liu H L, Zhang Y Y, He M L Li R T, Meng W, Wang Z Y, Li X F, Bu Q Y. Fine-tuning flowering time via genome editing of upstream open reading frames of heading date 2 in rice[J]. Rice, 2021, 14(1): 59. |
| [26] | 徐善斌, 郑洪亮, 刘利锋, 卜庆云, 李秀峰, 邹德堂. 利用CRISPR/Cas9 技术高效创制长粒香型水稻[J]. 中国水稻科学, 2020, 34(5): 406-412. |
| Xu S B, Zheng H L, Liu L F, Bu Q Y, Li X F, Zou D T. Improvement of grain shape and fragrance by using CRISPR/Cas9 system[J]. Chinese Journal of Rice Science, 2020, 34(5): 406-412. (in Chinese with English abstract) |
| [1] | 郭展, 张运波. 水稻对干旱胁迫的生理生化响应及分子调控研究进展[J]. 中国水稻科学, 2024, 38(4): 335-349. |
| [2] | 韦还和, 马唯一, 左博源, 汪璐璐, 朱旺, 耿孝宇, 张翔, 孟天瑶, 陈英龙, 高平磊, 许轲, 霍中洋, 戴其根. 盐、干旱及其复合胁迫对水稻产量和品质形成影响的研究进展[J]. 中国水稻科学, 2024, 38(4): 350-363. |
| [3] | 许丹洁, 林巧霞, 李正康, 庄小倩, 凌宇, 赖美玲, 陈晓婷, 鲁国东. OsOPR10正调控水稻对稻瘟病和白叶枯病的抗性[J]. 中国水稻科学, 2024, 38(4): 364-374. |
| [4] | 候小琴, 王莹, 余贝, 符卫蒙, 奉保华, 沈煜潮, 谢杭军, 王焕然, 许用强, 武志海, 王建军, 陶龙兴, 符冠富. 黄腐酸钾提高水稻秧苗耐盐性的作用途径分析[J]. 中国水稻科学, 2024, 38(4): 409-421. |
| [5] | 胡继杰, 胡志华, 张均华, 曹小闯, 金千瑜, 章志远, 朱练峰. 根际饱和溶解氧对水稻分蘖期光合及生长特性的影响[J]. 中国水稻科学, 2024, 38(4): 437-446. |
| [6] | 刘福祥, 甄浩洋, 彭焕, 郑刘春, 彭德良, 文艳华. 广东省水稻孢囊线虫病调查与鉴定[J]. 中国水稻科学, 2024, 38(4): 456-461. |
| [7] | 陈浩田, 秦缘, 钟笑涵, 林晨语, 秦竞航, 杨建昌, 张伟杨. 水稻根系和土壤性状与稻田甲烷排放关系的研究进展[J]. 中国水稻科学, 2024, 38(3): 233-245. |
| [8] | 缪军, 冉金晖, 徐梦彬, 卜柳冰, 王平, 梁国华, 周勇. 过量表达异三聚体G蛋白γ亚基基因RGG2提高水稻抗旱性[J]. 中国水稻科学, 2024, 38(3): 246-255. |
| [9] | 尹潇潇, 张芷菡, 颜绣莲, 廖蓉, 杨思葭, 郭岱铭, 樊晶, 赵志学, 王文明. 多个稻曲病菌效应因子的信号肽验证和表达分析[J]. 中国水稻科学, 2024, 38(3): 256-265. |
| [10] | 朱裕敬, 桂金鑫, 龚成云, 罗新阳, 石居斌, 张海清, 贺记外. 全基因组关联分析定位水稻分蘖角度QTL[J]. 中国水稻科学, 2024, 38(3): 266-276. |
| [11] | 魏倩倩, 汪玉磊, 孔海民, 徐青山, 颜玉莲, 潘林, 迟春欣, 孔亚丽, 田文昊, 朱练峰, 曹小闯, 张均华, 朱春权. 信号分子硫化氢参与硫肥缓解铝对水稻生长抑制作用的机制[J]. 中国水稻科学, 2024, 38(3): 290-302. |
| [12] | 周甜, 吴少华, 康建宏, 吴宏亮, 杨生龙, 王星强, 李昱, 黄玉峰. 不同种植模式对水稻籽粒淀粉含量及淀粉关键酶活性的影响[J]. 中国水稻科学, 2024, 38(3): 303-315. |
| [13] | 关雅琪, 鄂志国, 王磊, 申红芳. 影响中国水稻生产环节外包发展因素的实证研究:基于群体效应视角[J]. 中国水稻科学, 2024, 38(3): 324-334. |
| [14] | 许用强, 姜宁, 奉保华, 肖晶晶, 陶龙兴, 符冠富. 水稻开花期高温热害响应机理及其调控技术研究进展[J]. 中国水稻科学, 2024, 38(2): 111-126. |
| [15] | 吕海涛, 李建忠, 鲁艳辉, 徐红星, 郑许松, 吕仲贤. 稻田福寿螺的发生、危害及其防控技术研究进展[J]. 中国水稻科学, 2024, 38(2): 127-139. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||