
Chinese Journal OF Rice Science ›› 2025, Vol. 39 ›› Issue (5): 624-634.DOI: 10.16819/j.1001-7216.2025.240106
• Research Papers • Previous Articles Next Articles
SHAO Ye1,2,#, HU Yuanyi1,2,#, PENG Yan1,2,#, MAO Bigang1,2, LIU Huimin1,2, TANG Chanjuan1, LEI Bin1,2, TANG Li1,2, YU Lixia3, LI Wenjian3, LUO Wuzhong3, LUO Zhibin1,2, YUAN Yuantao1,2, LI Yaokui1,2, ZHANG Dan1,2, ZHOU Libin3, BAI Lianyang2,4,*( ), TANG Wenbang1,2,*(
), TANG Wenbang1,2,*( ), ZHAO Bingran1,2,*(
), ZHAO Bingran1,2,*( )
)
Received:2024-01-09
Revised:2024-04-17
Online:2025-09-10
Published:2025-09-10
About author:#These authors contributed equally to this work
韶也1,2,#, 胡远艺1,2,#, 彭彦1,2,#, 毛毕刚1,2, 刘慧敏1,2, 唐婵娟1, 雷斌1,2, 唐丽1,2, 余丽霞3, 李文建3, 罗武中3, 罗治斌1,2, 袁远涛1,2, 李曜魁1,2, 张丹1,2, 周利斌3, 柏连阳2,4,*( ), 唐文帮1,2,*(
), 唐文帮1,2,*( ), 赵炳然1,2,*(
), 赵炳然1,2,*( )
)
作者简介:#共同第一作者
基金资助:SHAO Ye, HU Yuanyi, PENG Yan, MAO Bigang, LIU Huimin, TANG Chanjuan, LEI Bin, TANG Li, YU Lixia, LI Wenjian, LUO Wuzhong, LUO Zhibin, YUAN Yuantao, LI Yaokui, ZHANG Dan, ZHOU Libin, BAI Lianyang, TANG Wenbang, ZHAO Bingran. Directed Improvement of Hybrid Rice Zhuoliangyou 1126 by Heavy Ion Beam Mutagenesis Based on M1TDS Targeted Screening Technology[J]. Chinese Journal OF Rice Science, 2025, 39(5): 624-634.
韶也, 胡远艺, 彭彦, 毛毕刚, 刘慧敏, 唐婵娟, 雷斌, 唐丽, 余丽霞, 李文建, 罗武中, 罗治斌, 袁远涛, 李曜魁, 张丹, 周利斌, 柏连阳, 唐文帮, 赵炳然. 基于M1TDS靶向筛选技术的重离子束诱变定向改良杂交水稻卓两优1126性状的研究[J]. 中国水稻科学, 2025, 39(5): 624-634.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2025.240106
| 名称 Name | 序列 Sequence |
|---|---|
| OsNRAMP5-246-8-F | GAAGGTGACCAAGTTCATGCTTGTGGTTGGCTCCGGCTTAA |
| OsNRAMP5-246-8-H | GAAGGTCGGAGTCAACGGATTTGTGGTTGGCTCCGGCTTGC |
| OsNRAMP5-246-8-C | CACAAAATGAAACAGTGGAAACCGATC |
| OsNRAMP5-234-6-F | GAAGGTGACCAAGTTCATGCTCACAAAATGAAACAGTGGAAACCGAT |
| OsNRAMP5-234-6-H | GAAGGTCGGAGTCAACGGATTCACAAAATGAAACAGTGGAAACCGAC |
| OsNRAMP5-234-6-C | CGTACCTCATATCTGTGGTTGGCTC |
| OsNRAMP5-204-6-F | GAAGGTGACCAAGTTCATGCTCACAAAATGAAACAGTGGAAACCGA |
| OsNRAMP5-204-6-H | GAAGGTCGGAGTCAACGGATTCACAAAATGAAACAGTGGAAACCGC |
| OsNRAMP5-204-6-C | CGTACCTCATATCTGTGGTTGGCTC |
| OsNRAMP5-37-7-F | GAAGGTGACCAAGTTCATGCTTGAAGAAGCTAAGAGAGGAAGCAACAA |
| OsNRAMP5-37-7-H | GAAGGTCGGAGTCAACGGATTTGAAGAAGCTAAGAGAGGAAGCAACAG |
| OsNRAMP5-37-7-C | ACTAGCTCTCCAGCTGATGCTC |
| OsBADH2-168-1-F | GAAGGTGACCAAGTTCATGCTGTGTAGTTGGGTTGATCACACCTT |
| OsBADH2-168-1-H | GAAGGTCGGAGTCAACGGATTGTGTAGTTGGGTTGATCACACCTC |
| OsBADH2-168-1-C | TGGGTCATAAATAAATATAAGCGCAGG |
| OsBADH2-224-8-F | GAAGGTGACCAAGTTCATGCTTTCGTCTTTTCTTGACAGCCTGTT |
| OsBADH2-224-8-H | GAAGGTCGGAGTCAACGGATTTTCGTCTTTTCTTGACAGCCTGTC |
| OsBADH2-224-8-C | AGGACTTTTTCCACCAAGTTCCAG |
| OsLOX3-760-7-F | GAAGGTGACCAAGTTCATGCTTATCTTCACCTATGCCACAAGGC |
| OsLOX3-760-7-H | GAAGGTCGGAGTCAACGGATTTATCTTCACCTATGCCACAAGGA |
| OsLOX3-760-7-C | CAGGGTATCATCATCACGTAAGAAC |
| OsLOX3-115-5-F | GAAGGTGACCAAGTTCATGCTGAGAGCAAAGTGTTGAACATGAAC |
| OsLOX3-115-5-H | GAAGGTCGGAGTCAACGGATTGAGAGCAAAGTGTTGAACATGAAG |
| OsLOX3-115-5-C | GGACCGACCCGGTTCTTGAG |
| OsPAO5-549-5-F | GAAGGTGACCAAGTTCATGCTTGCTTGACAGGAATCCACACCTA |
| OsPAO5-549-5-H | GAAGGTCGGAGTCAACGGATTTGCTTGACAGGAATCCACACCTG |
| OsPAO5-549-5-C | GCACCACCATTCCAGATATAGGTG |
| OsPAO5-1028-4-F | GAAGGTGACCAAGTTCATGCTAACTTTCCTCCAGGGTTACCAAAAT |
| OsPAO5-1028-4-H | GAAGGTCGGAGTCAACGGATTAACTTTCCTCCAGGGTTACCAAATC |
| OsPAO5-1028-4-C | GCTTGTCCCATCTTCAACACATAC |
| OsPAO5-84-2-F | GAAGGTGACCAAGTTCATGCTCTCAGCTCAAGAAAATGCTACCAGG |
| OsPAO5-84-2-H | GAAGGTCGGAGTCAACGGATTCTCAGCTCAAGAAAATGCTACCACT |
| OsPAO5-84-2-C | GACAGTACTCAACTGTACATGGTCTG |
| OsSSIIIa-598-7-F | GAAGGTGACCAAGTTCATGCTCCATTTGGTTCAAGGCCTAGAACTC |
| OsSSIIIa-598-7-H | GAAGGTCGGAGTCAACGGATTCCATTTGGTTCAAGGCCTAGAACTG |
| OsSSIIIa-598-7-C | CACTGTTTTCGACGTAGACCATG |
| OsBEIIb-931-5-F | GAAGGTGACCAAGTTCATGCTGTGTGATTGTTATTAAATCTCACCAGGA |
| OsBEIIb-931-5-H | GAAGGTCGGAGTCAACGGATTGTGTGATTGTTATTAAATCTCACCAGGC |
| OsBEIIb-931-5-C | GGCTATCTTAACTTTATGGGAAATGAGTTC |
Table 1. Primer sequence for KASP genotyping
| 名称 Name | 序列 Sequence |
|---|---|
| OsNRAMP5-246-8-F | GAAGGTGACCAAGTTCATGCTTGTGGTTGGCTCCGGCTTAA |
| OsNRAMP5-246-8-H | GAAGGTCGGAGTCAACGGATTTGTGGTTGGCTCCGGCTTGC |
| OsNRAMP5-246-8-C | CACAAAATGAAACAGTGGAAACCGATC |
| OsNRAMP5-234-6-F | GAAGGTGACCAAGTTCATGCTCACAAAATGAAACAGTGGAAACCGAT |
| OsNRAMP5-234-6-H | GAAGGTCGGAGTCAACGGATTCACAAAATGAAACAGTGGAAACCGAC |
| OsNRAMP5-234-6-C | CGTACCTCATATCTGTGGTTGGCTC |
| OsNRAMP5-204-6-F | GAAGGTGACCAAGTTCATGCTCACAAAATGAAACAGTGGAAACCGA |
| OsNRAMP5-204-6-H | GAAGGTCGGAGTCAACGGATTCACAAAATGAAACAGTGGAAACCGC |
| OsNRAMP5-204-6-C | CGTACCTCATATCTGTGGTTGGCTC |
| OsNRAMP5-37-7-F | GAAGGTGACCAAGTTCATGCTTGAAGAAGCTAAGAGAGGAAGCAACAA |
| OsNRAMP5-37-7-H | GAAGGTCGGAGTCAACGGATTTGAAGAAGCTAAGAGAGGAAGCAACAG |
| OsNRAMP5-37-7-C | ACTAGCTCTCCAGCTGATGCTC |
| OsBADH2-168-1-F | GAAGGTGACCAAGTTCATGCTGTGTAGTTGGGTTGATCACACCTT |
| OsBADH2-168-1-H | GAAGGTCGGAGTCAACGGATTGTGTAGTTGGGTTGATCACACCTC |
| OsBADH2-168-1-C | TGGGTCATAAATAAATATAAGCGCAGG |
| OsBADH2-224-8-F | GAAGGTGACCAAGTTCATGCTTTCGTCTTTTCTTGACAGCCTGTT |
| OsBADH2-224-8-H | GAAGGTCGGAGTCAACGGATTTTCGTCTTTTCTTGACAGCCTGTC |
| OsBADH2-224-8-C | AGGACTTTTTCCACCAAGTTCCAG |
| OsLOX3-760-7-F | GAAGGTGACCAAGTTCATGCTTATCTTCACCTATGCCACAAGGC |
| OsLOX3-760-7-H | GAAGGTCGGAGTCAACGGATTTATCTTCACCTATGCCACAAGGA |
| OsLOX3-760-7-C | CAGGGTATCATCATCACGTAAGAAC |
| OsLOX3-115-5-F | GAAGGTGACCAAGTTCATGCTGAGAGCAAAGTGTTGAACATGAAC |
| OsLOX3-115-5-H | GAAGGTCGGAGTCAACGGATTGAGAGCAAAGTGTTGAACATGAAG |
| OsLOX3-115-5-C | GGACCGACCCGGTTCTTGAG |
| OsPAO5-549-5-F | GAAGGTGACCAAGTTCATGCTTGCTTGACAGGAATCCACACCTA |
| OsPAO5-549-5-H | GAAGGTCGGAGTCAACGGATTTGCTTGACAGGAATCCACACCTG |
| OsPAO5-549-5-C | GCACCACCATTCCAGATATAGGTG |
| OsPAO5-1028-4-F | GAAGGTGACCAAGTTCATGCTAACTTTCCTCCAGGGTTACCAAAAT |
| OsPAO5-1028-4-H | GAAGGTCGGAGTCAACGGATTAACTTTCCTCCAGGGTTACCAAATC |
| OsPAO5-1028-4-C | GCTTGTCCCATCTTCAACACATAC |
| OsPAO5-84-2-F | GAAGGTGACCAAGTTCATGCTCTCAGCTCAAGAAAATGCTACCAGG |
| OsPAO5-84-2-H | GAAGGTCGGAGTCAACGGATTCTCAGCTCAAGAAAATGCTACCACT |
| OsPAO5-84-2-C | GACAGTACTCAACTGTACATGGTCTG |
| OsSSIIIa-598-7-F | GAAGGTGACCAAGTTCATGCTCCATTTGGTTCAAGGCCTAGAACTC |
| OsSSIIIa-598-7-H | GAAGGTCGGAGTCAACGGATTCCATTTGGTTCAAGGCCTAGAACTG |
| OsSSIIIa-598-7-C | CACTGTTTTCGACGTAGACCATG |
| OsBEIIb-931-5-F | GAAGGTGACCAAGTTCATGCTGTGTGATTGTTATTAAATCTCACCAGGA |
| OsBEIIb-931-5-H | GAAGGTCGGAGTCAACGGATTGTGTGATTGTTATTAAATCTCACCAGGC |
| OsBEIIb-931-5-C | GGCTATCTTAACTTTATGGGAAATGAGTTC |
| 名称 Name | 序列 Sequence |
|---|---|
| OsNRAMP5-3E-540F | GTTGGTCCTGGATTCATGGTGTC |
| OsNRAMP5-3E-540R | TCCAATCAGAATCACCCAGAGCAG |
| OsNRAMP5-1E-265F | TCTTCGTCTACTTCCAGCTAGCC |
| OsNRAMP5-1E-265R | ACAGTGAACAACTGTCCATGGTG |
| OsBADH-4-471F | TTGATGAAGCAGCATGGGACATG |
| OsBADH-4-471R | GCTGCTAGGTACAATTTGTGAGAC |
| OsBADH-8-357F | CTTCAGCTGCTCCTATGGTTAAGG |
| OsBADH-8-357R | TCTAGCATCCAGCTCAGTTAAGTGC |
| OsLOX3-8-535F | GAGAAGAACCTCGAAGGCCTCAG |
| OsLOX3-8-535R | CCATCACAGCATGTGTGTTGAGC |
| OsPAO-6/7-563F | GCTCTCTTTGATAAGGATGGTCGTC |
| OsPAO-6/7-563R | ATGGCCACCAGTAAGAACATGTTC |
| OsBEIIb-17-359F | GGCATGAGAACCTCACATACTG |
| OsBEIIb-17-359R | GGAAGTACTTGTGGAGCTCTTGG |
Table 2. Primers for Sanger sequencing
| 名称 Name | 序列 Sequence |
|---|---|
| OsNRAMP5-3E-540F | GTTGGTCCTGGATTCATGGTGTC |
| OsNRAMP5-3E-540R | TCCAATCAGAATCACCCAGAGCAG |
| OsNRAMP5-1E-265F | TCTTCGTCTACTTCCAGCTAGCC |
| OsNRAMP5-1E-265R | ACAGTGAACAACTGTCCATGGTG |
| OsBADH-4-471F | TTGATGAAGCAGCATGGGACATG |
| OsBADH-4-471R | GCTGCTAGGTACAATTTGTGAGAC |
| OsBADH-8-357F | CTTCAGCTGCTCCTATGGTTAAGG |
| OsBADH-8-357R | TCTAGCATCCAGCTCAGTTAAGTGC |
| OsLOX3-8-535F | GAGAAGAACCTCGAAGGCCTCAG |
| OsLOX3-8-535R | CCATCACAGCATGTGTGTTGAGC |
| OsPAO-6/7-563F | GCTCTCTTTGATAAGGATGGTCGTC |
| OsPAO-6/7-563R | ATGGCCACCAGTAAGAACATGTTC |
| OsBEIIb-17-359F | GGCATGAGAACCTCACATACTG |
| OsBEIIb-17-359R | GGAAGTACTTGTGGAGCTCTTGG |
| 目标基因 Target gene | 基因编号 Gene ID | 目标表型 Target phenotype | 外显子数 No. of exons | 目标区域覆盖度 Coverage of target area(%) | 平均测序深度 Average sequencing depth |
|---|---|---|---|---|---|
| OsNRAMP5 | Os07g0257200 | 镉低积累 Low cadmium accumulation | 13 | 100 | 53398× |
| OsBADH2 | Os08g0424500 | 香味Aroma | 15 | 100 | 53086× |
| OsLOX3 | Os03g0699700 | 耐储藏Storage-tolerance | 9 | 100 | 50202× |
| OsPAO5 | Os04g0671300 | 耐淹 Submergence-tolerance | 10 | 100 | 54263× |
| OsSSIIIa | Os08g0191433 | 低升糖指数 Low glycemic index (GI) | 14 | 100 | 54510× |
| OsBEIIb | Os02g0528200 | 低升糖指数 Low glycemic index (GI) | 22 | 100 | 52192× |
Table 3. Target gene and targeted sequencing analysis
| 目标基因 Target gene | 基因编号 Gene ID | 目标表型 Target phenotype | 外显子数 No. of exons | 目标区域覆盖度 Coverage of target area(%) | 平均测序深度 Average sequencing depth |
|---|---|---|---|---|---|
| OsNRAMP5 | Os07g0257200 | 镉低积累 Low cadmium accumulation | 13 | 100 | 53398× |
| OsBADH2 | Os08g0424500 | 香味Aroma | 15 | 100 | 53086× |
| OsLOX3 | Os03g0699700 | 耐储藏Storage-tolerance | 9 | 100 | 50202× |
| OsPAO5 | Os04g0671300 | 耐淹 Submergence-tolerance | 10 | 100 | 54263× |
| OsSSIIIa | Os08g0191433 | 低升糖指数 Low glycemic index (GI) | 14 | 100 | 54510× |
| OsBEIIb | Os02g0528200 | 低升糖指数 Low glycemic index (GI) | 22 | 100 | 52192× |
| 目标基因 Target gene | 亲本 Parent | 混池编号 No. of the pool | 突变位置 Mutation site | 野生/突变基因型 Wild/mutant genotype |
|---|---|---|---|---|
| OsNRAMP5 | 卓234S Zhuo 234S | 217-288E | 第3外显子Exon 3 | C/CTTAA |
| 卓234S Zhuo 234S | 217-288W | 第3外显子Exon 3 | C/CTTAA | |
| 卓234S Zhuo 234S | 217-288E | 第3外显子Exon 3 | GA/G | |
| 湘农恢1126 Xiangnonghui 1126 | 145-216S | 第3外显子Exon 3 | GCAGAT/G | |
| 湘农恢1126 Xiangnonghui 1126 | 1-72N | 第1外显子Exon 1 | CTCAATCTCCAT/C | |
| OsBADH2 | 卓234S Zhuo 234S | 145-216E | 第4外显子和第4内含子Exon 4/Intron 4 | CCTTGGTATTTCACATTTTTCT/C |
| 湘农恢1126 Xiangnonghui 1126 | 217-238S | 第8外显子Exon 8 | GTT/G | |
| OsLOX3 | 卓234S Zhuo 234S | 718-789W | 第7外显子Exon 7 | G/GC |
| 湘农恢1126 Xiangnonghui 1126 | 73-144S | 第9外显子Exon 9 | GAAC/G | |
| 湘农恢1126 Xiangnonghui 1126 | 73-144E | 第9外显子Exon 9 | GAAC/G | |
| OsPAO5 | 卓234S Zhuo 234S | 504-575W | 第6外显子和第6内含子Exon 6/Intron 6 | CACTTACTTT/C |
| 卓234S Zhuo 234S | 1004-1040E | 第9外显子Exon 9 | ATGAT/A | |
| 卓234S Zhuo 234S | 991-1040W | 第9外显子Exon 9 | ATGAT/A | |
| 湘农恢1126 Xiangnonghui 1126 | 73-144E | 第9外显子Exon 9 | GTAGCTCC/G | |
| OsSSIIIa | 卓234S Zhuo 234S | 576-647E | 第13外显子Exon 13 | CTCG/C |
| OsBEIIb | 卓234S Zhuo 234S | 863-932W | 第17外显子Exon 17 | GATGT/G |
Table 4. Mutations were captured by mixed-pool targeted gene sequencing
| 目标基因 Target gene | 亲本 Parent | 混池编号 No. of the pool | 突变位置 Mutation site | 野生/突变基因型 Wild/mutant genotype |
|---|---|---|---|---|
| OsNRAMP5 | 卓234S Zhuo 234S | 217-288E | 第3外显子Exon 3 | C/CTTAA |
| 卓234S Zhuo 234S | 217-288W | 第3外显子Exon 3 | C/CTTAA | |
| 卓234S Zhuo 234S | 217-288E | 第3外显子Exon 3 | GA/G | |
| 湘农恢1126 Xiangnonghui 1126 | 145-216S | 第3外显子Exon 3 | GCAGAT/G | |
| 湘农恢1126 Xiangnonghui 1126 | 1-72N | 第1外显子Exon 1 | CTCAATCTCCAT/C | |
| OsBADH2 | 卓234S Zhuo 234S | 145-216E | 第4外显子和第4内含子Exon 4/Intron 4 | CCTTGGTATTTCACATTTTTCT/C |
| 湘农恢1126 Xiangnonghui 1126 | 217-238S | 第8外显子Exon 8 | GTT/G | |
| OsLOX3 | 卓234S Zhuo 234S | 718-789W | 第7外显子Exon 7 | G/GC |
| 湘农恢1126 Xiangnonghui 1126 | 73-144S | 第9外显子Exon 9 | GAAC/G | |
| 湘农恢1126 Xiangnonghui 1126 | 73-144E | 第9外显子Exon 9 | GAAC/G | |
| OsPAO5 | 卓234S Zhuo 234S | 504-575W | 第6外显子和第6内含子Exon 6/Intron 6 | CACTTACTTT/C |
| 卓234S Zhuo 234S | 1004-1040E | 第9外显子Exon 9 | ATGAT/A | |
| 卓234S Zhuo 234S | 991-1040W | 第9外显子Exon 9 | ATGAT/A | |
| 湘农恢1126 Xiangnonghui 1126 | 73-144E | 第9外显子Exon 9 | GTAGCTCC/G | |
| OsSSIIIa | 卓234S Zhuo 234S | 576-647E | 第13外显子Exon 13 | CTCG/C |
| OsBEIIb | 卓234S Zhuo 234S | 863-932W | 第17外显子Exon 17 | GATGT/G |
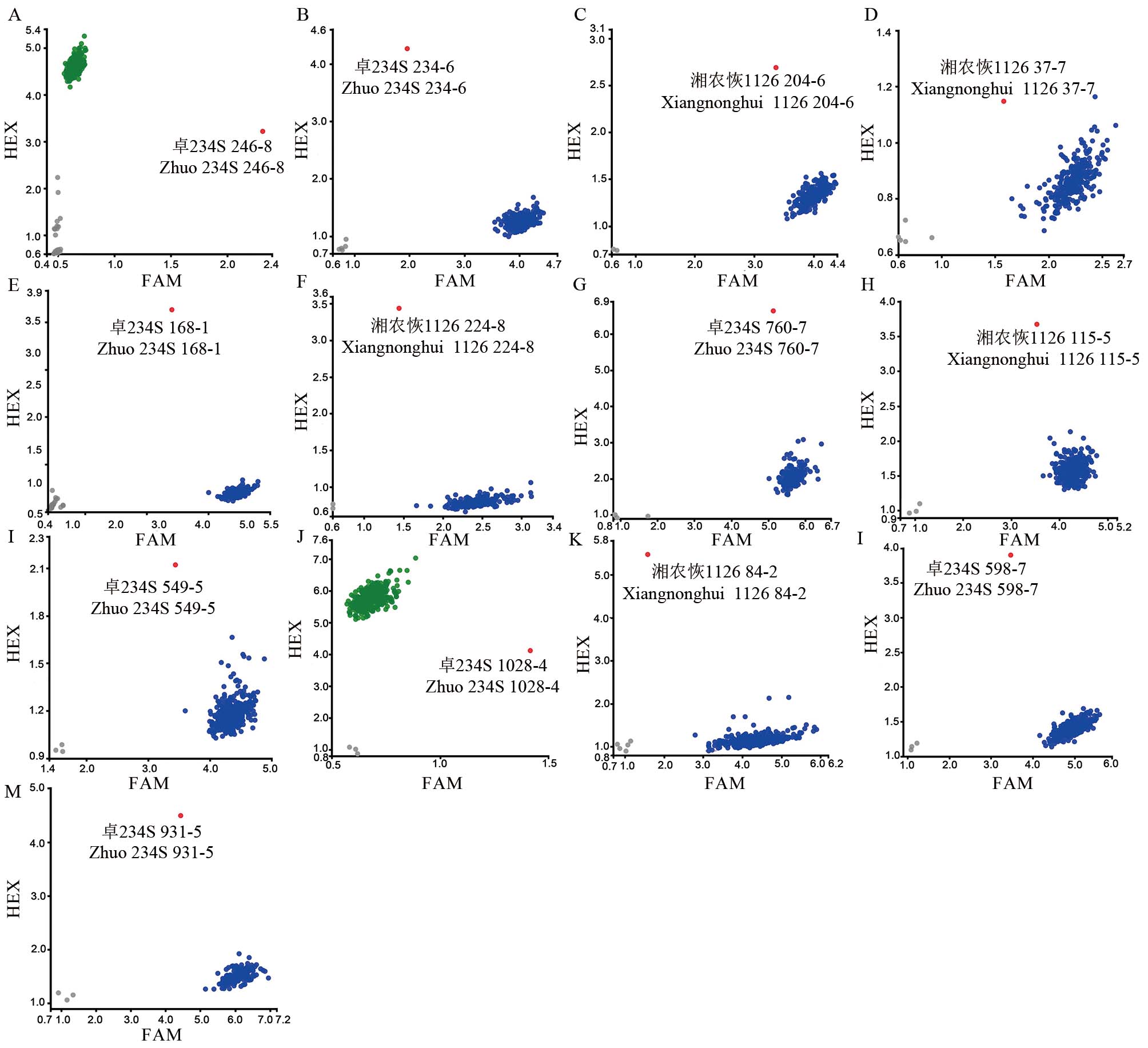
Fig. 1. KASP genotyping in the mixed pool of target gene mutations A-D represent the genotyping of different mutation sites of OsNRAMP5 gene; E and F represent the genotyping of different mutation sites of OsBADH2 gene, respectively; G and H represent genotyping of different mutation sites of OsLOX3 gene, respectively; I-K represent genotyping of different mutation sites of OsPAO5 gene; L, M represent the genotyping of mutation sites in OsSSIIIa gene and OsBEIIb gene, respectively; In panel A, 246-8 represents the 8th plant in 246 rows of M1 population (the same below); Blue/green dots represent wild genotypes, red dots represent mutant genotypes, and gray dots represent negative controls; FAM and HEX represent fluorescently modified labels of two different colors, respectively.
| 基因 Gene | 品种 Variety | 突变株定位(行-株) Location of mutants (row-plant) | 突变分蘖占比 Proportion of tillers containing mutations |
|---|---|---|---|
| OsNRAMP5 | 卓234S Zhuo 234S | 246-8 | 18/18 |
| OsNRAMP5 | 卓234S Zhuo 234S | 234-6 | 4/18 |
| OsNRAMP5 | 湘农恢1126 Xiangnonghui 1126 | 204-6 | 3/6 |
| OsNRAMP5 | 湘农恢1126 Xiangnonghui 1126 | 37-7 | 1/8 |
| OsBADH2 | 卓234S Zhuo 234S | 168-1 | 3/18 |
| OsBADH2 | 湘农恢1126 Xiangnonghui 1126 | 224-8 | 1/6 |
| OsLOX3 | 卓234S Zhuo 234S | 760-7 | 1/18 |
| OsLOX3 | 湘农恢1126 Xiangnonghui 1126 | 115-5 | 2/6 |
| OsPAO5 | 卓234S Zhuo 234S | 549-5 | 1/18 |
| OsPAO5 | 卓234S Zhuo 234S | 1028-4 | 5/18 |
| OsPAO5 | 湘农恢1126 Xiangnonghui 1126 | 84-2 | 1/6 |
| OsSSIIIa | 卓234S Zhuo 234S | 598-7 | 1/18 |
| OsBEIIb | 卓234S Zhuo 234S | 931-5 | 4/18 |
Table 5. Tiller ratio of mutagenic M1 mutants
| 基因 Gene | 品种 Variety | 突变株定位(行-株) Location of mutants (row-plant) | 突变分蘖占比 Proportion of tillers containing mutations |
|---|---|---|---|
| OsNRAMP5 | 卓234S Zhuo 234S | 246-8 | 18/18 |
| OsNRAMP5 | 卓234S Zhuo 234S | 234-6 | 4/18 |
| OsNRAMP5 | 湘农恢1126 Xiangnonghui 1126 | 204-6 | 3/6 |
| OsNRAMP5 | 湘农恢1126 Xiangnonghui 1126 | 37-7 | 1/8 |
| OsBADH2 | 卓234S Zhuo 234S | 168-1 | 3/18 |
| OsBADH2 | 湘农恢1126 Xiangnonghui 1126 | 224-8 | 1/6 |
| OsLOX3 | 卓234S Zhuo 234S | 760-7 | 1/18 |
| OsLOX3 | 湘农恢1126 Xiangnonghui 1126 | 115-5 | 2/6 |
| OsPAO5 | 卓234S Zhuo 234S | 549-5 | 1/18 |
| OsPAO5 | 卓234S Zhuo 234S | 1028-4 | 5/18 |
| OsPAO5 | 湘农恢1126 Xiangnonghui 1126 | 84-2 | 1/6 |
| OsSSIIIa | 卓234S Zhuo 234S | 598-7 | 1/18 |
| OsBEIIb | 卓234S Zhuo 234S | 931-5 | 4/18 |
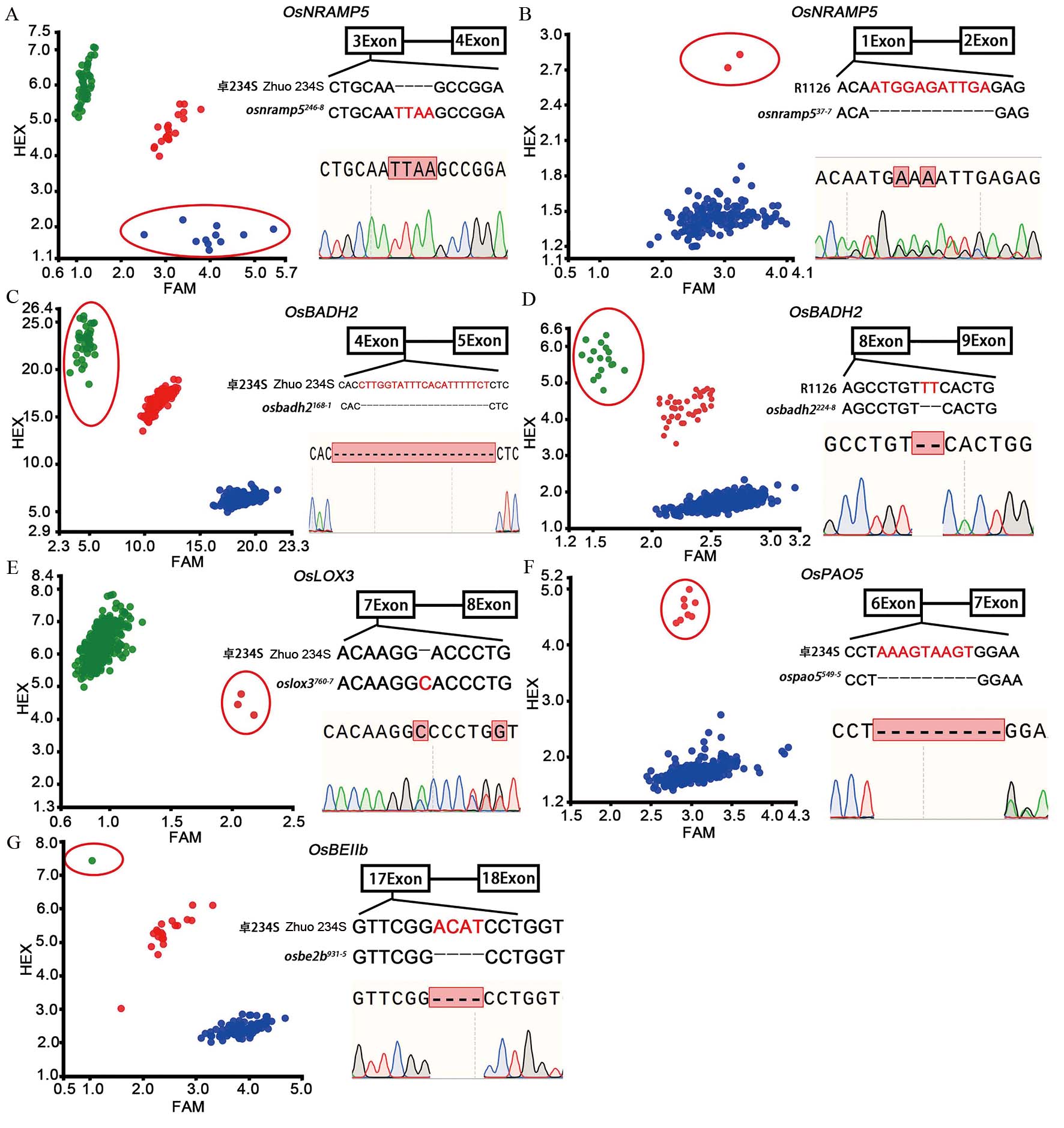
Fig. 2. KASP identification of M2 generation and Sanger sequencing of mutant plants A and B, KASP identification of OsNRAMP5 gene in Zhuo 234S and Xiangnonghui 1126 M2 population and Sanger sequencing of mutants. osnramp5246-8 and osnramp537-7 represent mutant plant of OsNRAMP5 gene in Zhuo 234S and Xiangnonghui 1126, respectively; C and D, KASP identification of OsBADH2 gene in Zhuo 234S and Xiangnonghui 1126 M2 population and Sanger sequencing of mutants; osbadh2168-1and osbadh2224-8 represent mutant plant of OsBADH2 gene in Zhuo234S and Xiang Nonghui 1126, respectively. E-G, KASP identification of OsLOX3, OsPAO5 and OsBEⅡb gene in Zhuo 234S M2 population and Sanger sequencing of mutants. oslox3760-7, ospao5549-5 and osbe2b931-5 represented mutant plant of OsLOX3, OsPAO5 and OsBEⅡb gene of Zhuo 234S, respectively. In panel A, red dots represent heterozygous genotype, blue dots represent homozygous mutant genotype, green dots represent wild genotype. In panels B-G, red dots represent heterozygous genotype, green dots represent homozygous mutant genotype, blue dots represent wild genotype. The dots in the red oval box represent the homozygous/heterozygous genotype of target genes.
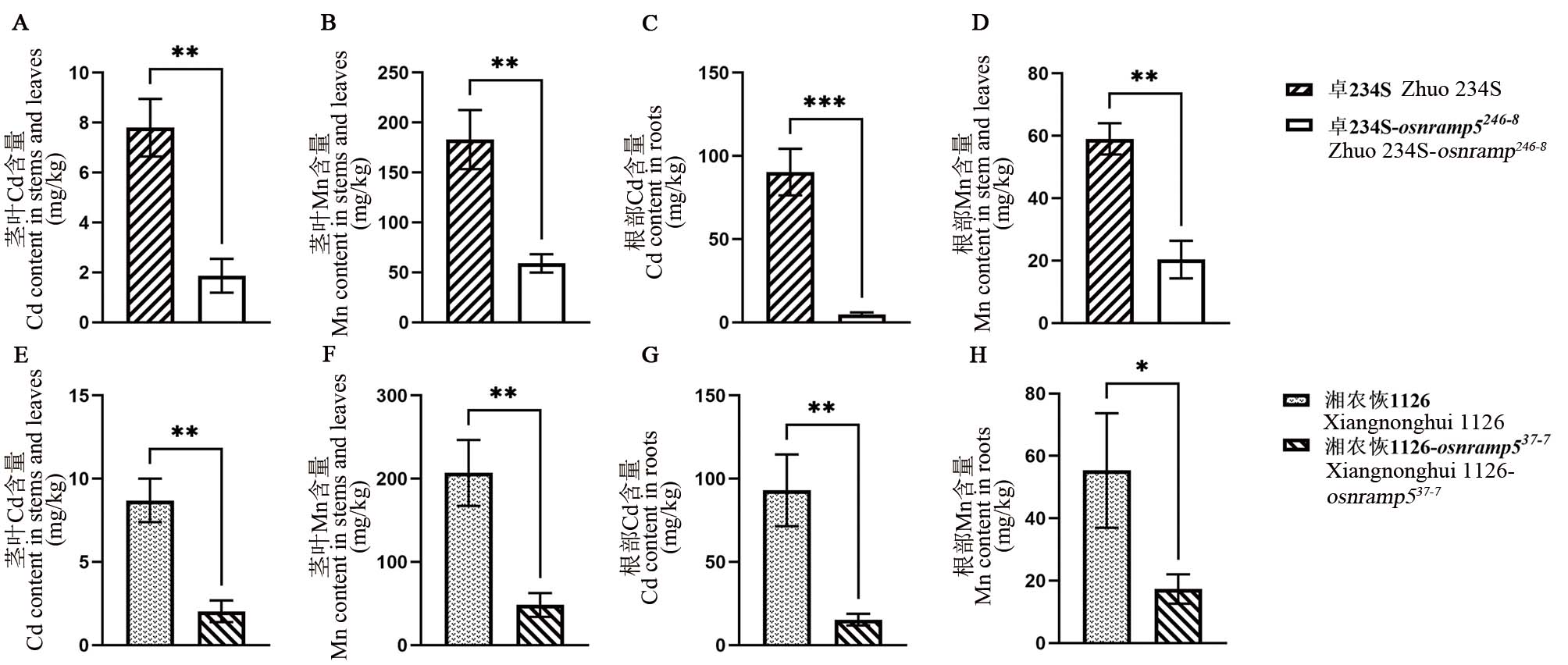
Fig. 3. Contents of Cd and Mn in homozygous mutant of OsNRAMP5 and control A-D, Contents of Cd and Mn in homozygous mutant Zhuo 234S-osnramp5246-8 and its original control material; E-H, Contents of Cd and Mn in the homozygous mutant Xiangnonghui 1126-osnramp537-7and its original control material; *, **, and *** represent difference was significant at 0.05, 0.01, and 0.001 levels, respectively (t-test).
| 材料 Material | 株高 Plant height (cm) | 有效分蘖数 No. of effective tillers | 主穗长 Panicle length(cm) | 一次枝梗数 No. of primary branches | 二次枝梗数 No. of secondary branches |
|---|---|---|---|---|---|
| 卓两优1126(对照) Zhuoliangyou 1126(CK) | 124.6±3.3 a | 13.2±1.3 a | 28.9±1.4 a | 18.2±1.3 a | 99.4±6.4 a |
| 镉低积累卓两优1126 Zhuoliangyou 1126 with low cadmium accumulation | 124.8±2.7 a | 13.6±1.1 a | 27.9±0.9 a | 19.0±0.7 a | 104.2±5.7 a |
| 材料 Material | 总粒数 Spikelets per panicle | 实粒数 Grains per panicle | 结实率 Seed setting rate(%) | 千粒重 1000-grain weight(g) | |
| 卓两优1126(对照) Zhuoliangyou 1126(CK) | 347.4±39.1 a | 301.6±28.9 a | 87.0±2.0 a | 22.9±0.4 a | |
| 镉低积累卓两优1126 Zhuoliangyou 1126 with low cadmium accumulation | 360.8±24.1 a | 316.8±19.6 a | 88.0±2.0 a | 22.9±0.5 a |
Table 6. Comparison of agronomic traits of Zhuoliangyou 1126 with low cadmium accumulation and its original control varieties
| 材料 Material | 株高 Plant height (cm) | 有效分蘖数 No. of effective tillers | 主穗长 Panicle length(cm) | 一次枝梗数 No. of primary branches | 二次枝梗数 No. of secondary branches |
|---|---|---|---|---|---|
| 卓两优1126(对照) Zhuoliangyou 1126(CK) | 124.6±3.3 a | 13.2±1.3 a | 28.9±1.4 a | 18.2±1.3 a | 99.4±6.4 a |
| 镉低积累卓两优1126 Zhuoliangyou 1126 with low cadmium accumulation | 124.8±2.7 a | 13.6±1.1 a | 27.9±0.9 a | 19.0±0.7 a | 104.2±5.7 a |
| 材料 Material | 总粒数 Spikelets per panicle | 实粒数 Grains per panicle | 结实率 Seed setting rate(%) | 千粒重 1000-grain weight(g) | |
| 卓两优1126(对照) Zhuoliangyou 1126(CK) | 347.4±39.1 a | 301.6±28.9 a | 87.0±2.0 a | 22.9±0.4 a | |
| 镉低积累卓两优1126 Zhuoliangyou 1126 with low cadmium accumulation | 360.8±24.1 a | 316.8±19.6 a | 88.0±2.0 a | 22.9±0.5 a |
| [1] | Oladosu Y, Rafii M, Abdullah N, Hussin G, Ramli A, Rahim H, Miah G, Usman M. Principle and application of plant mutagenesis in crop improvement: A review[J]. Biotechnology & Biotechnological Equipment, 2016, 30(1): 1-16. |
| [2] | Sahu P, Sao R, Mondal S, Vishwakarma G, Gupta S, Kumar V, Singh S, Sharma D, Das B. Next generation sequencing based forward genetic approaches for identification and mapping of causal mutations in crop plants: A comprehensive review[J]. Plants (Basel), 2020, 9(10): 1355. |
| [3] | Bagher A, Nahid A, Mohsen M, Vahid M. Nuclear techniques in agriculture and genetics[J]. American Journal of Bioscience, 2014, 2(3): 102-105. |
| [4] | Jankowicz-Cieslak J, Tai T, Kumlehn J, Till B. Biotechnologies for plant mutation breeding: Protocols[M]. Springer Nature, 2017. |
| [5] | Spencer-Lopes M, Forster B, Jankuloski L. Manual on mutation breeding[M]. Food and Agriculture Organization of the United Nations (FAO), 2018. |
| [6] | Tokuyama Y, Furusawa Y, Ide H, Yasui A, Terato H. Role of isolated and clustered DNA damage and the post-irradiating repair process in the effects of heavy ion beam irradiation[J]. Journal of Radiation Research, 2015, 56(3): 446-455. |
| [7] | Hamada N, Imaoka T, Masunaga S, Ogata T, Okayasu R, Takahashi A, Kato T A, Kobayashi Y, Ohnishi T, Ono K, Shimada Y, Teshima T. Recent advances in the biology of heavy-ion cancer therapy[J]. Journal of Radiation Research, 2010, 51(4): 365-383. |
| [8] | Tanaka A, Shikazono N, Hase Y. Studies on biological effects of ion beams on lethality, molecular nature of mutation, mutation rate, and spectrum of mutation phenotype for mutation breeding in higher plants[J]. Journal of Radiation Research, 2010, 51(3): 223-233. |
| [9] | Hirano T, Kazama Y, Ishii K, Ohbu S, Shirakawa Y, Abe T. Comprehensive identification of mutations induced by heavy-ion beam irradiation in Arabidopsis thaliana[J]. Plant Journal, 2015, 82(1): 93-104. |
| [10] | Dong X, Yan X, Li W. Plant mutation breeding with heavy ion irradiation at IMP[J]. Journal of Agricultural Science, 2016, 8(5): 34-41. |
| [11] | Abe T, Kazama Y, Hirano T. Ion beam breeding and gene discovery for function analyses using mutants[J]. Nuclear Physics News, 2015, 25(4): 30-34. |
| [12] | 韶也, 彭彦, 毛毕刚, 余丽霞, 唐丽, 李曜魁, 胡远艺, 张丹, 袁智成, 罗武中, 彭选明, 李文建, 周利斌, 柏连阳, 赵炳然. M1TDS技术及镉低积累杂交水稻亲本创制与组合选育[J]. 杂交水稻, 2022, 37(1): 1-11. |
| Shao Y, Peng Y, Mao B G, Yu L X, Tang L, Li Y K, Hu Y Y, Zhang D, Yuan Z C, Luo W Z, Peng X M, Li W J, Zhou L B, Bai L Y, Zhao B R. M1TDS technology and breeding of Cd low-accumulating hybrid rice parents and combinations[J]. Hybrid Rice, 2022, 37(1): 1-11. (in Chinese with English abstract) | |
| [13] | 赵连芝, 王浩瀚, 王勇, 李雁民, 甄东升, 颉红梅. 重离子辐照选育春小麦新品种初探[J]. 西北农业学报, 2006(3): 17-19. |
| Zhao L Z, Wang H H, Wang Y, Li Y M, Zhen D S, Xie H M. Preliminary study on breeding new spring wheat varieties by heavy-ion irradiation[J]. Acta Agriculturae Boreali-occidentalis Sinica, 2006(3): 17-19. (in Chinese with English abstract) | |
| [14] | Melsen K, van de Wouw M, Contreras R. Mutation breeding in ornamentals[J]. HortScience, 2021, 56(10): 1154-1165. |
| [15] | Hu W, Li W, Chen J. Recent advances of microbial breeding via heavy-ion mutagenesis at IMP[J]. Letters in Applied Microbiology, 2017, 65(4): 274-280. |
| [16] | 陆栋, 王颖, 刘青芳, 吴鑫, 王菊芳, 马爽, 李文建. 12C离子束辐照对酵母发酵能力的影响[J]. 核技术, 2010, 33(5): 350-353. |
| Lu D, Wang Y, Liu Q F, Wu X, Wang J F, Ma S, Li W J. Effects of 12C ion beam irradiation on yeast fermentation capacity[J]. Nuclear Techniques, 2010, 33(5): 350-353. (in Chinese with English abstract) | |
| [17] | McCallum C M, Comai L, Greene E A, Henikoff S. Targeted screening for induced mutations[J]. Nature Biotechnology, 2000, 18(4): 455-7. |
| [18] | Colbert T, Till B, Tompa R, Reynolds S, Steine M, Yeung A, McCallum C, Comai L, Henikoff S. High-throughput screening for induced point mutations[J]. Plant Physiology, 2001, 126(2): 480-4. |
| [19] | Taheri S, Abdullah T, Jain S, Sahebi M, Azizi P. TILLING, high-resolution melting (HRM), and next-generation sequencing (NGS) techniques in plant mutation breeding[J]. Molecular Breeding, 2017, 37: 1-23. |
| [20] | Till B J, Datta S, Jankowicz-Cieslak J. TILLING: The next generation[J]. Advances in Biochemical Engineering-biotechnology, 2018, 164: 139-160. |
| [21] | Kumar A, McKeown P, Boualem A, Ryder P, Brychkova G, Bendahmane A, Sarkar A, Chatterjee M, Spillane C. TILLING by Sequencing (TbyS) for targeted genome mutagenesis in crops[J]. Molecular Breeding, 2017, 37: 1-12. |
| [22] | Kashtwari M, Wani A, Rather R. TILLING: An alternative path for crop improvement[J]. Journal of Crop Improvement, 2019, 33(1): 83-109. |
| [23] | Dai P, Wu L R, Chen S X, Wang M X, Cheng L Y, Zhang J X, Hao P, Yao W, Zarka J, Issa G C, Kwong L, Zhang D Y. Calibration-free NGS quantitation of mutations below 0.01% VAF[J]. Nature Communications, 2021, 12(1): 6123. |
| [24] | Song P, Chen S X, Yan Y H, Pinto A, Cheng L Y, Dai P, Patel A A, Zhang D Y. Selective multiplexed enrichment for the detection and quantitation of low-fraction DNA variants via low-depth sequencing[J]. Nature Biomedical Engineering, 2021, 5(7): 690-701. |
| [25] | Tang L, Mao B, Li Y, Lü Q, Zhang L, Chen C, He H, Wang W, Zeng X, Shao Y, Pan Y, Hu Y, Peng Y, Fu X, Li H, Xia S, Zhao B. Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield[J]. Scientific Reports, 2017, 7(1): 14438. |
| [26] | Hui S, Li H, Mawia AM, Zhou L, Cai J, Ahmad S, Lai C, Wang J, Jiao G, Xie L, Shao G, Sheng Z, Tang S, Wang J, Wei X, Hu S, Hu P. Production of aromatic three-line hybrid rice using novel alleles of BADH2[J]. Plant Biotechnology Journal, 2022, 20(1): 59-74. |
| [27] | Xu H, Wei Y, Zhu Y, Lian L, Xie H, Cai Q, Chen Q, Lin Z, Wang Z, Xie H, Zhang J. Antisense suppression of LOX3 gene expression in rice endosperm enhances seed longevity[J]. Plant Biotechnology Journal, 2015, 13(4): 526-39. |
| [28] | Lü Y, Shao G, Jiao G, Sheng Z, Xie L, Hu S, Tang S, Wei X, Hu P. Targeted mutagenesis of POLYAMINE OXIDASE 5 that negatively regulates mesocotyl elongation enables the generation of direct-seeding rice with improved grain yield[J]. Molecular Plant, 2021, 14(2): 344-351. |
| [29] | Sun Y, Jiao G, Liu Z, Zhang X, Li J, Guo X, Du W, Du J, Francis F, Zhao Y, Xia L. Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes[J]. Frontiers in Plant Science, 2017, 8: 298. |
| [30] | Wang A, Jing Y, Cheng Q, Zhou H, Wang L, Gong W, Kou L, Liu G, Meng X, Chen M, Ma H, Shu X, Yu H, Wu D, Li J. Loss of function of SSIIIa and SSIIIb coordinately confers high RS content in cooked rice[J]. Proceedings of the National Academy of Sciences of the United States America, 2023, 120(19): e2220622120. |
| [31] | Kitamura S, Satoh K, Oono Y. Detection and characterization of genome-wide mutations in M1 vegetative cells of gamma-irradiated Arabidopsis[J]. PLoS Genetics, 2022, 18(1): e1009979. |
| [32] | Sasikala R, Kalaiyarasi R. Sensitivity of rice varieties to gamma irradiation[J]. Electronic Journal of Plant Breeding, 2010, 1(4): 845-889. |
| [33] | Gowthami R, Vanniarajan C, Souframanien J, Arumugam M. Comparison of radiosensitivity of two rice (Oryza sativa L.) varieties to gamma rays and electron beam in M1 generation[J]. Electronic Journal of Plant Breeding, 2017, 8(3): 732-741. |
| [34] | Yamaguchi H. Characteristics of ion beams as mutagens for mutation breeding in rice and chrysanthemums[J]. Japan Agricultural Research Quarterly, 2013, 47(4): 339-346. |
| [1] | YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang. Advances in Molecular Mechanism and Breeding Application of Heading Date Regulation in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 145-154. |
| [2] | NI Chen, ZHANG Jiahao, ZHU Changjin, XU Jiwei, HU Qiuqian, HUO Zhongyang, DAI Qigen, XU Ke, LI Guohui. Research Progress on the Formation and Regulation of Rice Source, Flow and Sink and Their Influencing Factors [J]. Chinese Journal OF Rice Science, 2026, 40(2): 155-170. |
| [3] | WANG Mengning, XIE Keran, GAO Ti, WANG Zhenmei, XIONG Dongliang, CUI Kehui. Research Progress on Effects of High Temperature on Rice Grain Weight Formation and Cultivation Strategies [J]. Chinese Journal OF Rice Science, 2026, 40(2): 171-180. |
| [4] | LUO Xiaoyun, ZHENG Xingfei, PENG Xuanguo, YU Qizhi, DONG Hualin, YIN Desuo, WANG Hongbo, HU Jianlin, XUE Lian, HU Peng, XU Deze. Research on Rice Lodging Resistance: Current Status, Challenges, and Future Directions [J]. Chinese Journal OF Rice Science, 2026, 40(2): 181-195. |
| [5] | XUE Pao, WANG Youshuang, HE Wanwan, HUANG Chenbo, ZHANG Han, DING Zhenqian, CHEN Qiuli, FAN Yunxin, DING Chengwei, SUN Lianping, HU Tingting. Identification and Cloning of SG5 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 210-222. |
| [6] | DUAN Min, XIE Liujie, YUE Yani, HUANG Shanjun. Development of High-quality Aromatic japonica Rice Lines by Using CRISPR/Cas9 Technology [J]. Chinese Journal OF Rice Science, 2026, 40(2): 235-243. |
| [7] | ZHANG Mengke, LU Jiayu, HE Jin, XU Xue, WU Shuang, WANG Peiran, CHEN Ruofan, JIN Qing, WANG Xiufeng. Development and Application of Functional Markers for Purity Identification of Wild-abortive Three-line Hybrid Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 244-252. |
| [8] | WANG Weiying, ZHAO Hong, CHEN Yu, YAO Xiaoming, LU Jianfei, GUO Qianshuang, DU Yongjun. Disruption Efficacy of Intelligent Active Aerosol Sex Pheromone on Mating Behavior of Major Lepidopteran Pests in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 264-272. |
| [9] | ZHENG Xusong, XU Hongxing, LÜ Haitao, LI Jianzhong, LÜ Zhongxian, LU Yanhui. Feeding and Damage Characteristics of Pomacea canaliculata on Direct-seeded Rice Seedlings of Different Varieties [J]. Chinese Journal OF Rice Science, 2026, 40(2): 273-280. |
| [10] | LI Xingyi, CHEN Ling, SHAO Jiantao, XIAO Suqin, LI Jinlu, FU Huixian, YIN Fuyou, ZHANG Jianhong, CHENG Zaiquan, LIU Li. Progress in Molecular Genetic Research on Co-regulation of Grain Yield and Starch Quality in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 1-17. |
| [11] | MIAO Zhening, CHEN Jijin, LI Ziming, LIU Yi, LUO Lijun. Advances and Prospects in Breeding Bacterial Blight-resistant Rice in South China [J]. Chinese Journal OF Rice Science, 2026, 40(1): 18-26. |
| [12] | YUE Xuanyu, XIE Wenya, FENG Zhiming, CHEN Zongxiang, HU Keming, ZUO Shimin. Function of OsERF93 in Regulating Resistance to Sheath Blight in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 37-50. |
| [13] | WANG Yixin, LIN Shen, MA Liuyang, CHEN Long, FENG Baohua, NI Shen, WEI Xiangjin, HE Jiwai, CHEN Tianxiao. Regulation of Nitrogen Uptake and Yield in Rice by the Alanine Aminotransferase Gene OsAlaAT4 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 51-60. |
| [14] | HUANG Qina, JIANG Hongrui, YANG Jie, YU Kunyu, YANG Changdeng, LIANG Yan. Bioinformatics Analysis, Development and Application of Molecular Markers for Seed Dormancy Gene Sdr4 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 61-71. |
| [15] | CHENG Zhaoping, HE Niqing, BAI Kangcheng, LIN Shaojun, HUANG Fenghuang, LIU Junhua, CHENG Zuxin, HUANG Chengzhi, YANG Dewei. Breeding and Utilization of the Three-Line Male Sterile Line Fumeng A with Pyramided Rice Blast Resistance Genes Pigm-1 and Pid2 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 72-84. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||