
Chinese Journal OF Rice Science ›› 2025, Vol. 39 ›› Issue (5): 703-710.DOI: 10.16819/j.1001-7216.2025.240906
• Research Papers • Previous Articles
YANG Xingzhou1,2, CUI Miaomiao1,2, WEI Lihui2, GU Aiguo3, LI Dongxia1, LE Xiuhu1,*( ), FENG Hui2,*(
), FENG Hui2,*( )
)
Received:2024-09-18
Revised:2024-11-27
Online:2025-09-10
Published:2025-09-10
杨行洲1,2, 崔苗苗1,2, 魏利辉2, 顾爱国3, 李东霞1, 乐秀虎1,*( ), 冯辉2,*(
), 冯辉2,*( )
)
基金资助:YANG Xingzhou, CUI Miaomiao, WEI Lihui, GU Aiguo, LI Dongxia, LE Xiuhu, FENG Hui. Effects of Exogenous miR3979 on Chemotaxis, Infection and Development of Root Knot Nematode Meloidogyne graminicola in Rice[J]. Chinese Journal OF Rice Science, 2025, 39(5): 703-710.
杨行洲, 崔苗苗, 魏利辉, 顾爱国, 李东霞, 乐秀虎, 冯辉. 外源miR3979处理水稻对拟禾本科根结线虫趋性、侵染和发育的影响[J]. 中国水稻科学, 2025, 39(5): 703-710.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2025.240906
| 引物名称 Primer name | 序列(5’-3’) Sequence(5’-3’) | 目的 Aim |
|---|---|---|
| miR3979-3p.RT | GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGCTTCTCTCC | RT-PCR |
| miR390-5p.RT | GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGGCGCT | |
| miR3979-3p.F | TCCCGCCTTCGGGGGAGGAG | qPCR |
| miR390-5p.F | TGACCAAAGCTCAGGAGGGAT | |
| Universal R | GTGCAGGGTCCGAGGT |
Table 1. Primers used in this study
| 引物名称 Primer name | 序列(5’-3’) Sequence(5’-3’) | 目的 Aim |
|---|---|---|
| miR3979-3p.RT | GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGCTTCTCTCC | RT-PCR |
| miR390-5p.RT | GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACGGCGCT | |
| miR3979-3p.F | TCCCGCCTTCGGGGGAGGAG | qPCR |
| miR390-5p.F | TGACCAAAGCTCAGGAGGGAT | |
| Universal R | GTGCAGGGTCCGAGGT |
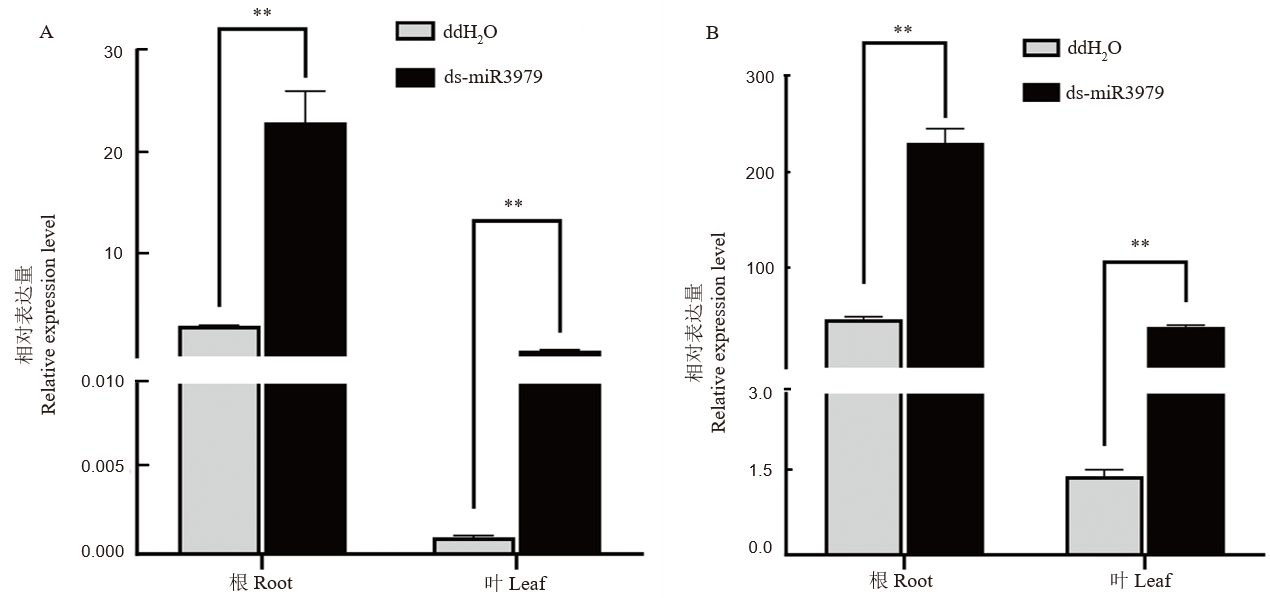
Fig. 2. Relative expression of miR3979-3p in rice plants treated with ds-miR3979 solution A-B, Relative expression of miR3979-3p in roots and shoots of rice treated with 400 nmol/L ds-miR3979 solution for 12 h (A) or 24 h (B). Each treatment repeated three times. **, P < 0.01.
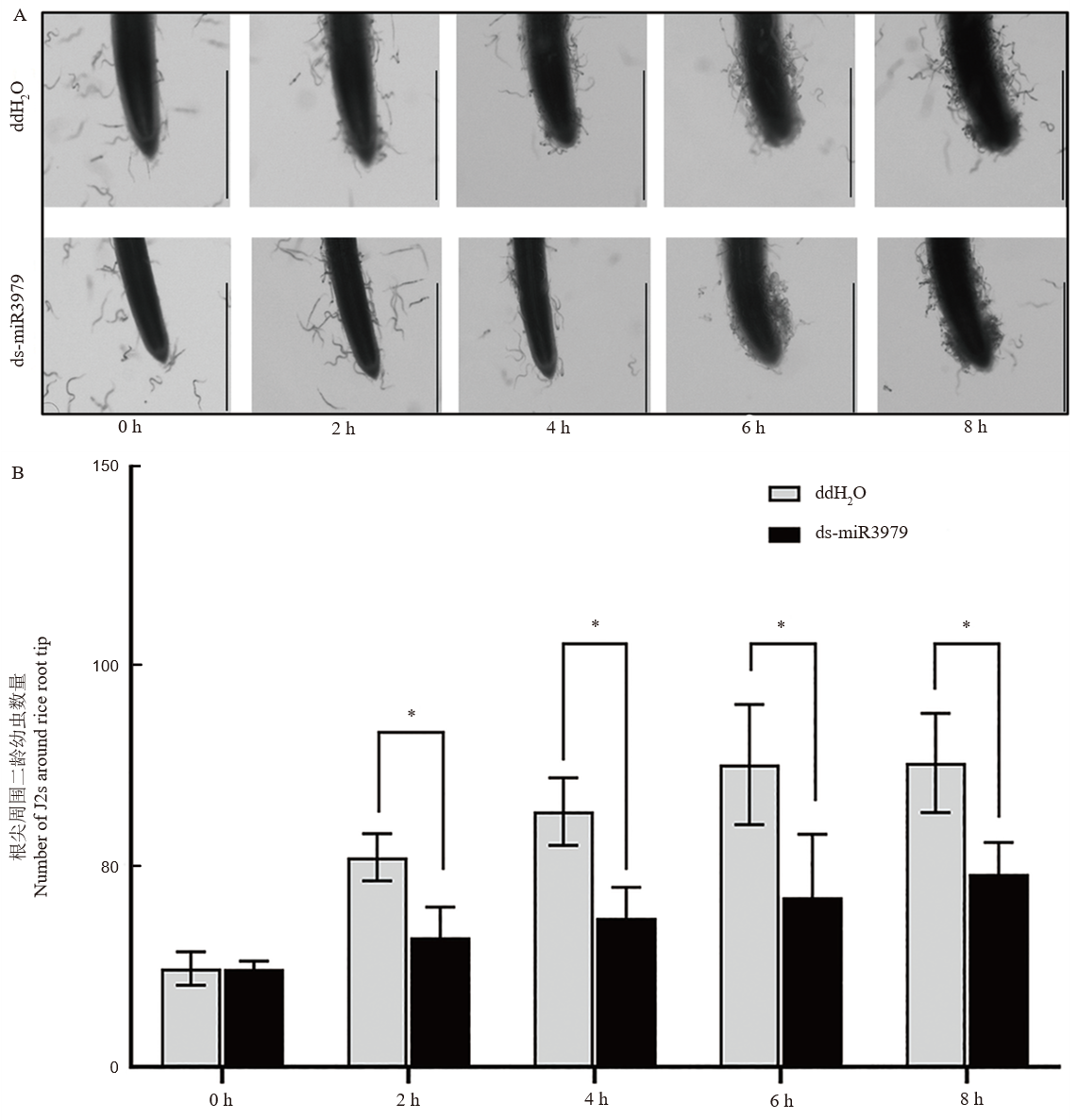
Fig. 3. Chems tabis of J2 towards rice roots treated with ds-miR3979 solution A. Chemotaxis of J2s towards rice root tips. B. Number of J2s around rice root tips. Rice roots were soaked into 400 nmol/L ds-miR3979 solution or ddH2O for 12 h and then cut into 2 cm length for the assay. Each treatment repeated three times. Scale bar = 1 mm; *, P < 0.05.
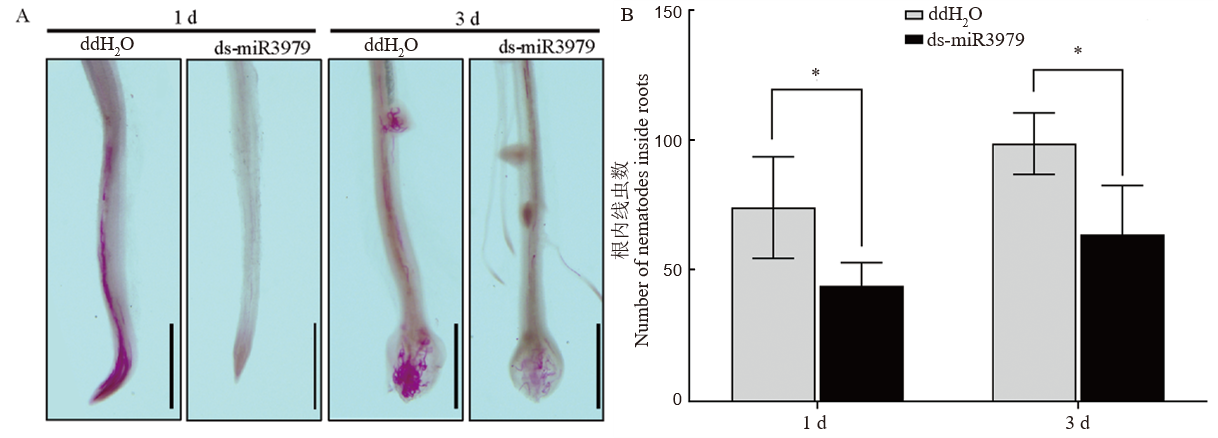
Fig. 4. Early infection of J2 in rice roots treated with ds-miR3979 solution A, J2s inside the roots stained with acid fuchsin at 1 and 3 days after inoculation (dpi); B, Number of J2s in roots at 1 and 3 days after inoculation. Rice root were soaked into 400 nmol/L ds-miR3979 solution or ddH2O for 12 h. Each treatment repeated six times. Scale = 1 mm. *, P < 0.05.
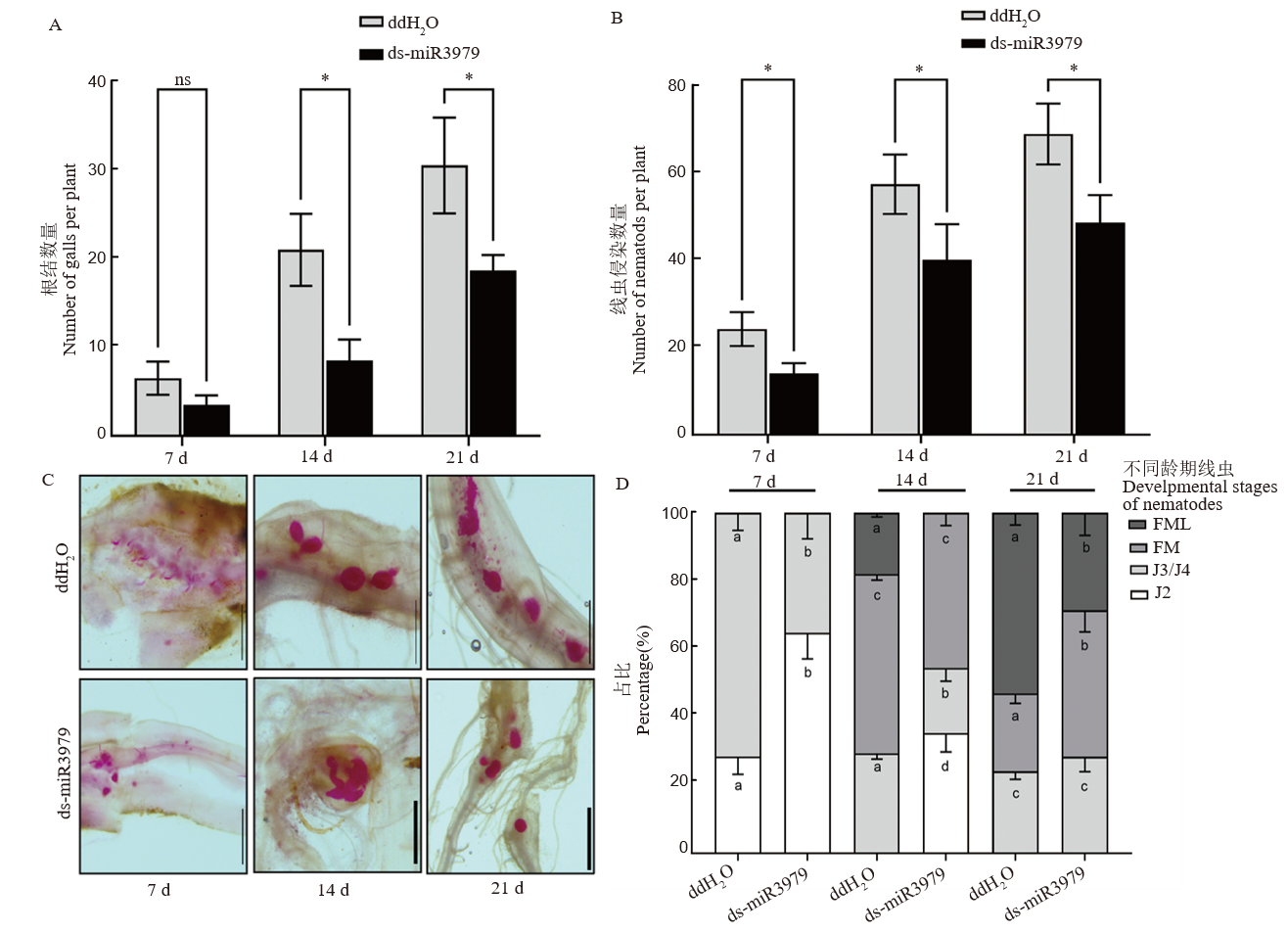
Fig. 5. Development of M. graminicola in rice treated with ds-miR3979 solution A and B, Number of galls (A) and nematodes (B) in roots at 7, 14, and 21 day after inoculation (dpi); C, Nematodes in the roots stained with acid fuchsin; D, percentage of different stages of nematodes in the roots. J2, Second stage juvenile; J3/4, Third/fourth stage juvenile; FM, Developing female, FML, Egg-caying female. Rice roots were soaked into 400 nmol/L ds-miR3979 solution or ddH2O for 24 h. Each treatment repeated six times. Scale = 1 mm. Different letters in a column (D) indicate significant difference for identical developmental stages between control and treatment at the same time point at P<0.05; ns, No significant difference; *, P<0.05.
| [1] | Bologna N G, Voinnet O. The diversity, biogenesis, and activities of endogenous silencing small RNAs in Arabidopsis[J]. Annual Review of Plant Biology, 2014, 65: 473-503. |
| [2] | Borges F, Martienssen R A. The expanding world of small RNAs in plants[J]. Nature Reviews Molecular Cell Biology, 2015, 16(12): 727-741. |
| [3] | Molnar A, Melnyk C, Baulcombe D C. Silencing signals in plants: A long journey for small RNAs[J]. Genome Biology, 2011, 12(1): 215. |
| [4] | Li S, Wang X, Xu W, Liu T, Cai C, Chen L, Clark C B, Ma J. Unidirectional movement of small RNAs from shoots to roots in interspecific heterografts[J]. Nature Plants, 2021, 7(1): 50-59. |
| [5] | Betti F, Ladera-Carmona M J, Weits D A, Ferri G, Iacopino S, Novi G, Svezia B, Kunkowska A B, Santaniello A, Piaggesi A, Loreti E, Perata P. Exogenous miRNAs induce post-transcriptional gene silencing in plants[J]. Nature Plants, 2021, 7(10): 1379-1388. |
| [6] | Li T, Li H, Zhang Y X, Liu J Y. Identification and analysis of seven H₂O₂-responsive miRNAs and 32 new miRNAs in the seedlings of rice (Oryza sativa L. ssp. indica)[J]. Nucleic Acids Research, 2011, 39(7): 2821-2833. |
| [7] | Nguyen D Q, Nguyen N L, Nguyen V T, Tran T H G, Nguyen T H, Nguyen T K L, Nguyen H H. Comparative analysis of microRNA expression profiles in shoot and root tissues of contrasting rice cultivars (Oryza sativa L.) with different salt stress tolerance[J]. PLoS One, 2023, 18(5): e0286140. |
| [8] | Bakhshi B, Mohseni Fard E, Nikpay N, Ali Ebrahimi M, Bihamta M R, Mardi M, Salekdeh G H. microRNA signatures of drought signaling in rice root[J]. PLoS One, 2016, 11(6): e0156814. |
| [9] | Liu Q, Zhang H. Molecular identification and analysis of arsenite stress-responsive miRNAs in rice[J]. Journal of Agricultural and Food Chemistry, 2012, 60(26): 6524-6536. |
| [10] | Sato Y, Nosaka-Takahashi M, Suzuki T, Shimizu-Sato S. Small RNAs in Rice: Molecular Species and Their Functions[M]. Rice Genomics, Genetics and Breeding, Singapore: Springer Nature, 2018: 21-36. |
| [11] | Xu D, Mou G, Wang K, Zhou G. MicroRNAs responding to southern rice black-streaked dwarf virus infection and their target genes associated with symptom development in rice[J]. Virus Research, 2014, 190: 60-68. |
| [12] | Wu Y, Lü W, Hu L, Rao W, Zeng Y, Zhu L, He Y, He G. Identification and analysis of brown planthopper-responsive microRNAs in resistant and susceptible rice plants[J]. Scientific Reports, 2017, 7(1): 8712. |
| [13] | Javed M, Reddy B, Sheoran N, Ganesan P, Kumar A. Unraveling the transcriptional network regulated by miRNAs in blast-resistant and blast-susceptible rice genotypes during Magnaporthe oryzae interaction[J]. Gene, 2023, 886: 147718. |
| [14] | Jeong D H, Park S, Zhai J, Gurazada S G R, De Paoli E, Meyers B C, Green P J. Massive analysis of rice small RNAs: Mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage[J]. The Plant Cell, 2011, 23(12): 4185-4207. |
| [15] | Mantelin S, Bellafiore S, Kyndt T. Meloidogyne graminicola: A major threat to rice agriculture[J]. Molecular Plant Pathology, 2016, 18: 3-15. |
| [16] | Kyndt T, Fernandez D, Gheysen G. Plant-parasitic nematode infections in rice: Molecular and cellular insights[J]. Annual Review of Phytopathology, 2014, 52: 135-153. |
| [17] | 刘国坤, 肖顺, 张绍升, 张敦富, 王玉. 拟禾本科根结线虫对水稻根系的侵染特性及其生活史[J]. 热带作物学报, 2011, 32(4): 743-748. |
| Liu G K, Xiao S, Zhang S S, Zhang D F, Wang Y. Infection characteristics and life cycle of Meloidogyne graminicola on rice roots[J]. Chinese Journal of Tropical Crops, 2011, 32(4): 743-748. (in Chinese with English abstract) | |
| [18] | Niu Y, Xiao L, de Almeida-Engler J, Gheysen G, Peng D, Xiao X, Huang W, Wang G, Xiao Y. Morphological characterization reveals new insights into giant cell development of Meloidogyne graminicola on rice[J]. Planta, 2022, 255(3): 70. |
| [19] | Verstraeten B, Atighi M R, Ruiz-Ferrer V, Escobar C, de Meyer T, Kyndt T. Non-coding RNAs in the interaction between rice and Meloidogyne graminicola[J]. BMC Genomics, 2021, 22(1): 560. |
| [20] | Hui F, Zhou C R, Zhu F, Le X H, Jing D D, Daly P, Zhou D M, Wei L H. Resistance analysis of the rice variety Huaidao 5 against root-knot nematode Meloidogyne graminicola[J]. Journal of Integrative Agriculture, 2023, 22(10): 3081-3089. |
| [21] | 冯辉, 聂国嫒, 陈曦, 张金凤, 周冬梅, 魏利辉. 拟禾谷根结线虫江苏分离群体形态学和分子鉴定[J]. 江苏农业学报, 2017, 33(4): 794-801. |
| Feng H, Nie G A, Chen X, Zhang J F, Zhou D M, Wei L H. Morphological and molecular identification of Meloidogyne graminicola isolates from Jiangsu Province[J]. Jiangsu Journal of Agricultural Sciences, 2017, 33(4): 794-801. (in Chinese with English abstract) | |
| [22] | Wang C, Lower S, Williamson V M. Application of Pluronic gel to the study of root-knot nematode behaviour[J]. Nematology, 2009, 11(3): 453-464. |
| [23] | Cuperus J T, Fahlgren N, Carrington J C. Evolution and functional diversification of MIRNA genes[J]. The Plant Cell, 2011, 23(2): 431-442. |
| [24] | Schmittgen T D, Livak K J. Analyzing real-time PCR data by the comparative CT method[J]. Nat Protocols, 2008, 3(6): 1101-1108. |
| [25] | Koeppe S, Kawchuk L, Kalischuk M. RNA Interference past and future applications in plants[J]. International Journal of Molecular Sciences, 2023, 24(11): 9755. |
| [26] | Bilir Ö, Göl D, Hong Y, McDowell J M, Tör M. Small RNA-based plant protection against diseases[J]. Frontiers in Plant Science, 2022, 13: 951097. |
| [27] | Dalakouras A, Wassenegger M, Dadami E, Ganopoulos I, Pappas M L, Papadopoulou K. Genetically modified organism-free RNA interference: Exogenous application of RNA molecules in plants[J]. Plant Physiology, 2020, 182(1): 38-50. |
| [28] | Niu D, Hamby R, Sanchez J N, Cai Q, Yan Q, Jin H. RNAs: A new frontier in crop protection[J]. Current Opinion in Biotechnology, 2021, 70: 204-212. |
| [29] | 王帅, 魏钰洋, 张羲, 谢佳, 胡展, 孙然锋. 根结线虫趋化性研究进展[J]. 农药学学报, 2022, 24(5): 982-996. DOI: 10.16801/j.issn.1008-7303.2022.0090. |
| Wang S, Wei Y Y, Zhang X, Xie J, Hu Z, Sun R F. Research progress on chemotaxis of root-knot nematodes[J]. Chinese Journal of Pesticide Science, 2022, 24(5): 982-996. DOI: 10.16801/j.issn.1008-7303.2022.0090. (in Chinese with English abstract) | |
| [30] | Ghorbanzadeh Z, Hamid R, Jacob F, Mirzaei M, Zeinalabedini M, Abdirad S, Atwell B J, Haynes P A, Ghafari M R, Salekdeh G H. MicroRNA profiling of root meristematic zone in contrasting genotypes reveals novel insight into rice response to water deficiency[J]. Journal of Plant Growth Regulation, 2022, 42(6): 3814-3834. |
| [31] | 张静文. 普通野生稻(Oryza rufipogon Griff.)干旱胁迫相关的 microRNAs 的鉴定与功能分析[D]. 北京: 中国农业大学, 2017. |
| Zhang J W. Identification and functional analysis of drought stress-related microRNAs in Oryza rufipogon Griff.[D]. Beijing: China Agricultural University, 2017. (in Chinese) | |
| [32] | Chowdhury A T, Hasan M N, Bhuiyan F H, Islam M Q, Nayon M R W, Rahaman M M, Hoque H, Jewel N A, Ashrafuzzaman M, Prodhan S H. Identification, characterization of Apyrase (APY) gene family in rice (Oryza sativa) and analysis of the expression pattern under various stress conditions[J]. PLoS One, 2023, 18(5): e0273592. |
| [1] | YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang. Advances in Molecular Mechanism and Breeding Application of Heading Date Regulation in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 145-154. |
| [2] | NI Chen, ZHANG Jiahao, ZHU Changjin, XU Jiwei, HU Qiuqian, HUO Zhongyang, DAI Qigen, XU Ke, LI Guohui. Research Progress on the Formation and Regulation of Rice Source, Flow and Sink and Their Influencing Factors [J]. Chinese Journal OF Rice Science, 2026, 40(2): 155-170. |
| [3] | WANG Mengning, XIE Keran, GAO Ti, WANG Zhenmei, XIONG Dongliang, CUI Kehui. Research Progress on Effects of High Temperature on Rice Grain Weight Formation and Cultivation Strategies [J]. Chinese Journal OF Rice Science, 2026, 40(2): 171-180. |
| [4] | LUO Xiaoyun, ZHENG Xingfei, PENG Xuanguo, YU Qizhi, DONG Hualin, YIN Desuo, WANG Hongbo, HU Jianlin, XUE Lian, HU Peng, XU Deze. Research on Rice Lodging Resistance: Current Status, Challenges, and Future Directions [J]. Chinese Journal OF Rice Science, 2026, 40(2): 181-195. |
| [5] | XUE Pao, WANG Youshuang, HE Wanwan, HUANG Chenbo, ZHANG Han, DING Zhenqian, CHEN Qiuli, FAN Yunxin, DING Chengwei, SUN Lianping, HU Tingting. Identification and Cloning of SG5 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 210-222. |
| [6] | DUAN Min, XIE Liujie, YUE Yani, HUANG Shanjun. Development of High-quality Aromatic japonica Rice Lines by Using CRISPR/Cas9 Technology [J]. Chinese Journal OF Rice Science, 2026, 40(2): 235-243. |
| [7] | ZHANG Mengke, LU Jiayu, HE Jin, XU Xue, WU Shuang, WANG Peiran, CHEN Ruofan, JIN Qing, WANG Xiufeng. Development and Application of Functional Markers for Purity Identification of Wild-abortive Three-line Hybrid Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 244-252. |
| [8] | WANG Weiying, ZHAO Hong, CHEN Yu, YAO Xiaoming, LU Jianfei, GUO Qianshuang, DU Yongjun. Disruption Efficacy of Intelligent Active Aerosol Sex Pheromone on Mating Behavior of Major Lepidopteran Pests in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 264-272. |
| [9] | ZHENG Xusong, XU Hongxing, LÜ Haitao, LI Jianzhong, LÜ Zhongxian, LU Yanhui. Feeding and Damage Characteristics of Pomacea canaliculata on Direct-seeded Rice Seedlings of Different Varieties [J]. Chinese Journal OF Rice Science, 2026, 40(2): 273-280. |
| [10] | LI Xingyi, CHEN Ling, SHAO Jiantao, XIAO Suqin, LI Jinlu, FU Huixian, YIN Fuyou, ZHANG Jianhong, CHENG Zaiquan, LIU Li. Progress in Molecular Genetic Research on Co-regulation of Grain Yield and Starch Quality in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 1-17. |
| [11] | MIAO Zhening, CHEN Jijin, LI Ziming, LIU Yi, LUO Lijun. Advances and Prospects in Breeding Bacterial Blight-resistant Rice in South China [J]. Chinese Journal OF Rice Science, 2026, 40(1): 18-26. |
| [12] | ZHOU Huacheng, BAO Xiuhao, WANG Xiaole, LU Zhenfei, MA Rongrong, LU Yongfa, WANG Xiaoyan, CAI Kefeng, TANG Zhiming, ZHOU Zhuoqi, CHEN Zhixin, MA Weiwei. Research Progress in the Phase Separation of RNA-binding Proteins in Plant Cells [J]. Chinese Journal OF Rice Science, 2026, 40(1): 27-36. |
| [13] | YUE Xuanyu, XIE Wenya, FENG Zhiming, CHEN Zongxiang, HU Keming, ZUO Shimin. Function of OsERF93 in Regulating Resistance to Sheath Blight in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 37-50. |
| [14] | WANG Yixin, LIN Shen, MA Liuyang, CHEN Long, FENG Baohua, NI Shen, WEI Xiangjin, HE Jiwai, CHEN Tianxiao. Regulation of Nitrogen Uptake and Yield in Rice by the Alanine Aminotransferase Gene OsAlaAT4 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 51-60. |
| [15] | HUANG Qina, JIANG Hongrui, YANG Jie, YU Kunyu, YANG Changdeng, LIANG Yan. Bioinformatics Analysis, Development and Application of Molecular Markers for Seed Dormancy Gene Sdr4 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 61-71. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||