
Chinese Journal OF Rice Science ›› 2025, Vol. 39 ›› Issue (5): 615-623.DOI: 10.16819/j.1001-7216.2025.240510
• Research Papers • Previous Articles Next Articles
WANG Jingbo, SU Chang, FENG Jing, JIANG Sixu, XU Hai, CUI Zhibo*( ), ZHAO Minghui*(
), ZHAO Minghui*( )
)
Received:2024-05-19
Revised:2024-09-27
Online:2025-09-10
Published:2025-09-10
王镜博, 苏畅, 冯晶, 姜思旭, 徐海, 崔志波*( ), 赵明辉*(
), 赵明辉*( )
)
基金资助:WANG Jingbo, SU Chang, FENG Jing, JIANG Sixu, XU Hai, CUI Zhibo, ZHAO Minghui. Functional Study on Aluminum Tolerance of OsAlR1 Gene in Rice[J]. Chinese Journal OF Rice Science, 2025, 39(5): 615-623.
王镜博, 苏畅, 冯晶, 姜思旭, 徐海, 崔志波, 赵明辉. 水稻OsAlR1基因耐铝性功能研究[J]. 中国水稻科学, 2025, 39(5): 615-623.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2025.240510
| 引物名称 Primer name | 序列 Primer sequence (5'-3') | 用途 Usage |
|---|---|---|
| OsAlR1-F | AACACGGGGGACTTTGCAACATGGGCAACTCGCTGGCGTTG | 基因扩增 Gene amplification |
| OsAlR1-R | TCCTCGCCCTTCACGATACACGCCTCATAGATGGTATCAAGGGC | |
| OsAlR1-gRNA-F | GGCAGCGTTGCGGTGCGTCTGTGG | CRISPR/Cas9载体构建 CRISPR/Cas9 vector construction |
| OsAlR1-gRNA-R | AAACCCACAGACGCACCGCAACGC | |
| OsAlR1-seq-F | GGAGAGAACACGGGGGAC | 鉴定 Identification |
| OsAlR1-seq-R | GTGCAGATGAACTTGAGG | |
| OsAlR1-OE-F | GGAGAGAACACGGGGGAC | 表达量分析 Expression analysis |
| OsAlR1-OE-R | GTGCAGATGAACTTGAGG | |
| OsAlR1-KO-F | ATGGGCAACTCGCTGGCGTTG | |
| OsAlR1-KO-R | CGCCTCATAGATGGTATCAAGGGC | |
| OsAlR1-qPCR-F | AGCTGATGATCGAGGCGC | |
| OsAlR1-qPCR-R | CTTGCTCCGCCCTTTCTTG | |
| OsPAL7-qPCR-F | CATCGGGAAGCTCATGTTCG | |
| OsPAL7-qPCR-R | CTTGAAGCCGTAGTCCAAGC | |
| OsHCT4-qPCR-F | GGTCCGGCGATCATGTACTA | |
| OsHCT4-qPCR-R | GCGAACACCTTCCTGAACTC | |
| Os02g0467600-qPCR-F | CGCAGAGCTTCGAGTACAAC | |
| Os02g0467600-qPCR-R | CACGGCACTTGTTGAGGTAG | |
| OsCCR18-qPCR-F | CTGATGACGCTGCCAAGAAC | |
| OsCCR18-qPCR-R | ATCATCTTCTCCGGGTCGTC | |
| PRX82-qPCR-F | CGAGGAGTTCACGGAGAGTT | |
| PRX82-qPCR-R | AGCAGTCGTGGAAGAAGAGG | |
| Os06g0522300-qPCR-F | CTGACCAGGAGCTCTACACC | |
| Os06g0522300-qPCR-R | CTCTTGTGAAGTCGGCGAAG |
Table 1. Primer sequences used in this study
| 引物名称 Primer name | 序列 Primer sequence (5'-3') | 用途 Usage |
|---|---|---|
| OsAlR1-F | AACACGGGGGACTTTGCAACATGGGCAACTCGCTGGCGTTG | 基因扩增 Gene amplification |
| OsAlR1-R | TCCTCGCCCTTCACGATACACGCCTCATAGATGGTATCAAGGGC | |
| OsAlR1-gRNA-F | GGCAGCGTTGCGGTGCGTCTGTGG | CRISPR/Cas9载体构建 CRISPR/Cas9 vector construction |
| OsAlR1-gRNA-R | AAACCCACAGACGCACCGCAACGC | |
| OsAlR1-seq-F | GGAGAGAACACGGGGGAC | 鉴定 Identification |
| OsAlR1-seq-R | GTGCAGATGAACTTGAGG | |
| OsAlR1-OE-F | GGAGAGAACACGGGGGAC | 表达量分析 Expression analysis |
| OsAlR1-OE-R | GTGCAGATGAACTTGAGG | |
| OsAlR1-KO-F | ATGGGCAACTCGCTGGCGTTG | |
| OsAlR1-KO-R | CGCCTCATAGATGGTATCAAGGGC | |
| OsAlR1-qPCR-F | AGCTGATGATCGAGGCGC | |
| OsAlR1-qPCR-R | CTTGCTCCGCCCTTTCTTG | |
| OsPAL7-qPCR-F | CATCGGGAAGCTCATGTTCG | |
| OsPAL7-qPCR-R | CTTGAAGCCGTAGTCCAAGC | |
| OsHCT4-qPCR-F | GGTCCGGCGATCATGTACTA | |
| OsHCT4-qPCR-R | GCGAACACCTTCCTGAACTC | |
| Os02g0467600-qPCR-F | CGCAGAGCTTCGAGTACAAC | |
| Os02g0467600-qPCR-R | CACGGCACTTGTTGAGGTAG | |
| OsCCR18-qPCR-F | CTGATGACGCTGCCAAGAAC | |
| OsCCR18-qPCR-R | ATCATCTTCTCCGGGTCGTC | |
| PRX82-qPCR-F | CGAGGAGTTCACGGAGAGTT | |
| PRX82-qPCR-R | AGCAGTCGTGGAAGAAGAGG | |
| Os06g0522300-qPCR-F | CTGACCAGGAGCTCTACACC | |
| Os06g0522300-qPCR-R | CTCTTGTGAAGTCGGCGAAG |
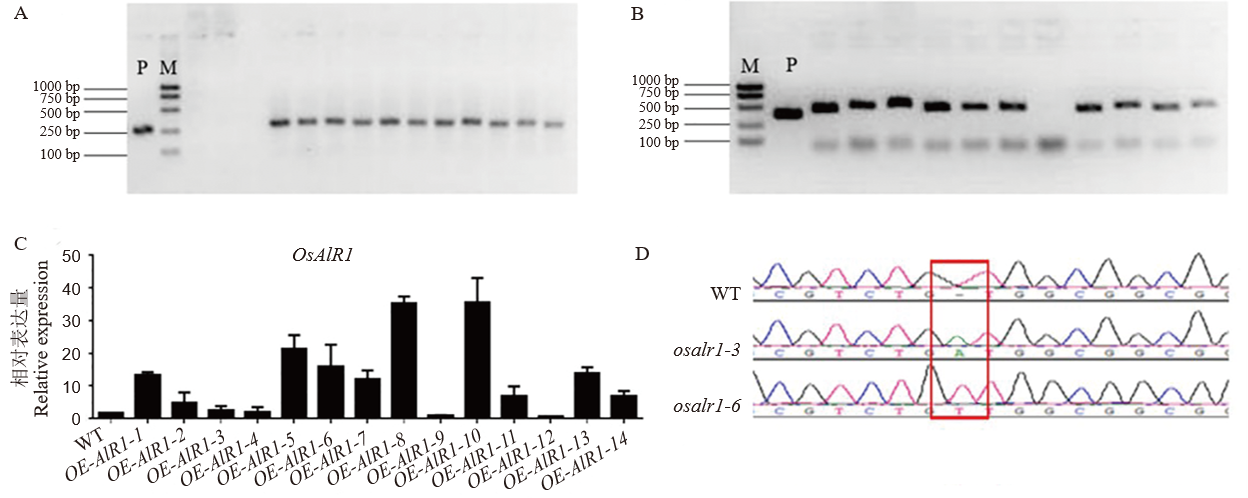
Fig. 2. Identification of transgenic plants A, Identification of overexpressing lines; B, Identification of knockout lines; C, Expression analysis of the overexpression lines; D, Target site analysis of knockout lines. M, Trans 1K; P, Positive control.
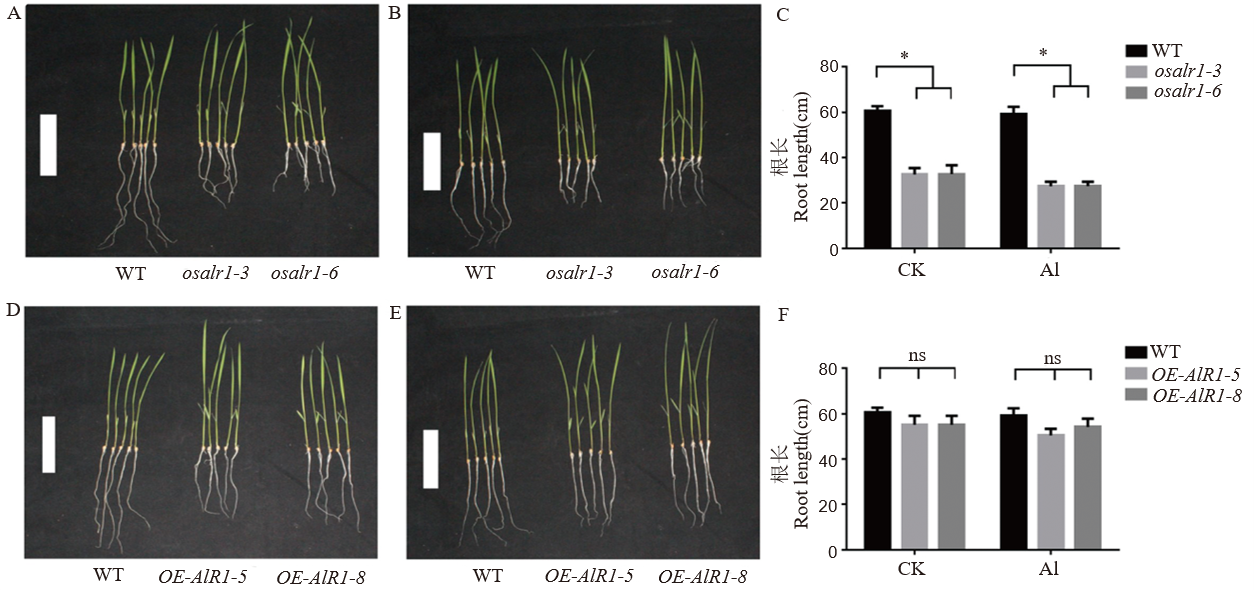
Fig. 3. Root length identification of transgenic plants (Scale Bar=10 cm) A, Root length of control groups of knockout lines; B, Root length of experimental groups of knockout lines; C, Comparison of root length of knockout lines before and after Al treatment; D, Root length of control groups of overexpression lines; E, Root length of experimental groups of overexpression lines; F, Comparison of root length of overexpression lines before and after Al treatment; CK, Before Al treatment; Al, After Al treatment. * indicates significant difference at the 0.05 probability level. The same below.
| [1] | Liu H B, Zhu R, Shu K, Lü W X, Wang S, Wang C L. Aluminum stress signaling, response, and adaptive mechanisms in plants[J]. Plant Signaling & Behavior, 2022, 17(1): 2057060. |
| [2] | Ma J F. Syndrome of aluminum toxicity and diversity of aluminum resistance in higher plants[J]. International Review of Cytology, 2007, 264: 225-252. |
| [3] | Liu J, Piñeros M A, Kochian L V. The role of aluminum sensing and signaling in plant aluminum resistance[J]. Journal of Integrative Plant Biology, 2014, 56(3): 221-230. |
| [4] | Sharma P, Dubey R S. Involvement of oxidative stress and role of antioxidative defense system in growing rice seedlings exposed to toxic concentrations of aluminum[J]. Plant Cell Reports, 2007, 26(11): 2027-2038. |
| [5] | Kanazawa S, Sano S, Koshiba T, Ushimaru T. Changes in antioxidative enzymes in cucumber cotyledons during natural senescence: Comparison with those during dark-induced senescence[J]. Physiologia Plantarum, 2000, 109(2): 211-216. |
| [6] | Mittler R. Oxidative stress, antioxidants and stress tolerance[J]. Trends in Plant Science, 2002, 7(9): 405-410. |
| [7] | Liu Q Q, Luo L, Zheng L Q. Lignins: Biosynthesis and biological functions in plants[J]. International Journal of Molecular Sciences, 2018, 19(2): 335. |
| [8] | Dong N Q, Lin H X. Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions[J]. Journal of Integrative Plant Biology, 2021, 63(1): 180-209. |
| [9] | Wang M L, Wang Y D, Li X Z, Zhang Y, Chen X L, Liu J Y, Qiu Y W, Wang A X. Integration of metabolomics and transcriptomics reveals the regulation mechanism of the phenylpropanoid biosynthesis pathway in insect resistance traits in Solanum habrochaites[J]. Horticulture Research, 2024, 11(2): uhad277. |
| [10] | Fritz R R, Hodgins D S, Abell C W. Phenylalanine ammonia-lyase. Induction and purification from yeast and clearance in mammals[J]. Journal of Biological Chemistry, 1976, 251(15): 4646-4650. |
| [11] | Ohl S, Hedrick S A, Chory J, Lamb C J. Functional properties of a phenylalanine ammonia-lyase promoter from Arabidopsis[J]. The Plant Cell, 1990, 2(9): 837-848. |
| [12] | Zhang X B, Liu C J. Multifaceted regulations of gateway enzyme phenylalanine ammonia-lyase in the biosynthesis of phenylpropanoids[J]. Molecular Plant, 2015, 8(1): 17-27. |
| [13] | Lacombe E, Hawkins S, van Doorsselaere J, Piquemal J, Goffner D, Poeydomenge O, Boudet A M, Grima-Pettenati J. Cinnamoyl CoA reductase, the first committed enzyme of the lignin branch biosynthetic pathway: Cloning, expression and phylogenetic relationships[J]. The Plant Journal, 1997, 11(3): 429-441. |
| [14] | Schoch G, Goepfert S, Morant M, Hehn A, Meyer D, Ullmann P, Werck-Reichhart D. CYP98A3 from Arabidopsis thaliana is a 3'-hydroxylase of phenolic esters, a missing link in the phenylpropanoid pathway[J]. Journal of Biological Chemistry, 2001, 276(39): 36566-36574. |
| [15] | Hoffmann L, Maury S, Martz F, Geoffroy P, Legrand M. Purification, cloning, and properties of an acyltransferase controlling shikimate and quinate ester intermediates in phenylpropanoid metabolism[J]. Journal of Biological Chemistry, 2003, 278(1): 95-103. |
| [16] | Wang J B, Su C, Cui Z B, Huang L X, Gu S, Jiang S X, Feng J, Xu H, Zhang W Z, Jiang L L, Zhao M H. Transcriptomics and metabolomics reveal tolerance new mechanism of rice roots to Al stress[J]. Frontiers in Genetics, 2022, 13: 1063984. |
| [17] | Zhao M H, Song J Y, Wu A T, Hu T, Li J Q. Mining beneficial genes for aluminum tolerance within a core collection of rice landraces through genome-wide association mapping with high density SNPs from specific-locus amplified fragment sequencing[J]. Frontiers in Plant Science, 2018, 9: 1838. |
| [18] | Guo J H, Liu X J, Zhang Y, Shen J L, Han W X, Zhang W F, Christie P, Goulding K W T, Vitousek P M, Zhang F S. Significant acidification in major Chinese croplands[J]. Science, 2010, 327(5968): 1008-1010. |
| [19] | Chu X Q, Wang J G, Li M Z, Zhang S J, Gao Y Y, Fan M, Han C, Xiang F N, Li G Y, Wang Y, Yu X, Xiang C B, Bai M Y. HBI transcription factor-mediated ROS homeostasis regulates nitrate signal transduction[J]. The Plant Cell, 2021, 33(9): 3004-3021. |
| [20] | Choudhury F K, Rivero R M, Blumwald E, Mittler R. Reactive oxygen species, abiotic stress and stress combination[J]. The Plant Journal, 2017, 90(5): 856-867. |
| [21] | Ito T, Ohkama-Ohtsu N. Degradation of glutathione and glutathione conjugates in plants[J]. Journal of Experimental Botany, 2023, 74(11): 3313-3327. |
| [22] | de Storme N, Geelen D. The impact of environmental stress on male reproductive development in plants: Biological processes and molecular mechanisms[J]. Plant, Cell & Environment, 2014, 37(1): 1-18. |
| [1] | YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang. Advances in Molecular Mechanism and Breeding Application of Heading Date Regulation in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 145-154. |
| [2] | NI Chen, ZHANG Jiahao, ZHU Changjin, XU Jiwei, HU Qiuqian, HUO Zhongyang, DAI Qigen, XU Ke, LI Guohui. Research Progress on the Formation and Regulation of Rice Source, Flow and Sink and Their Influencing Factors [J]. Chinese Journal OF Rice Science, 2026, 40(2): 155-170. |
| [3] | WANG Mengning, XIE Keran, GAO Ti, WANG Zhenmei, XIONG Dongliang, CUI Kehui. Research Progress on Effects of High Temperature on Rice Grain Weight Formation and Cultivation Strategies [J]. Chinese Journal OF Rice Science, 2026, 40(2): 171-180. |
| [4] | LUO Xiaoyun, ZHENG Xingfei, PENG Xuanguo, YU Qizhi, DONG Hualin, YIN Desuo, WANG Hongbo, HU Jianlin, XUE Lian, HU Peng, XU Deze. Research on Rice Lodging Resistance: Current Status, Challenges, and Future Directions [J]. Chinese Journal OF Rice Science, 2026, 40(2): 181-195. |
| [5] | XUE Pao, WANG Youshuang, HE Wanwan, HUANG Chenbo, ZHANG Han, DING Zhenqian, CHEN Qiuli, FAN Yunxin, DING Chengwei, SUN Lianping, HU Tingting. Identification and Cloning of SG5 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 210-222. |
| [6] | DUAN Min, XIE Liujie, YUE Yani, HUANG Shanjun. Development of High-quality Aromatic japonica Rice Lines by Using CRISPR/Cas9 Technology [J]. Chinese Journal OF Rice Science, 2026, 40(2): 235-243. |
| [7] | ZHANG Mengke, LU Jiayu, HE Jin, XU Xue, WU Shuang, WANG Peiran, CHEN Ruofan, JIN Qing, WANG Xiufeng. Development and Application of Functional Markers for Purity Identification of Wild-abortive Three-line Hybrid Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 244-252. |
| [8] | WANG Weiying, ZHAO Hong, CHEN Yu, YAO Xiaoming, LU Jianfei, GUO Qianshuang, DU Yongjun. Disruption Efficacy of Intelligent Active Aerosol Sex Pheromone on Mating Behavior of Major Lepidopteran Pests in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 264-272. |
| [9] | ZHENG Xusong, XU Hongxing, LÜ Haitao, LI Jianzhong, LÜ Zhongxian, LU Yanhui. Feeding and Damage Characteristics of Pomacea canaliculata on Direct-seeded Rice Seedlings of Different Varieties [J]. Chinese Journal OF Rice Science, 2026, 40(2): 273-280. |
| [10] | LI Xingyi, CHEN Ling, SHAO Jiantao, XIAO Suqin, LI Jinlu, FU Huixian, YIN Fuyou, ZHANG Jianhong, CHENG Zaiquan, LIU Li. Progress in Molecular Genetic Research on Co-regulation of Grain Yield and Starch Quality in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 1-17. |
| [11] | MIAO Zhening, CHEN Jijin, LI Ziming, LIU Yi, LUO Lijun. Advances and Prospects in Breeding Bacterial Blight-resistant Rice in South China [J]. Chinese Journal OF Rice Science, 2026, 40(1): 18-26. |
| [12] | YUE Xuanyu, XIE Wenya, FENG Zhiming, CHEN Zongxiang, HU Keming, ZUO Shimin. Function of OsERF93 in Regulating Resistance to Sheath Blight in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 37-50. |
| [13] | WANG Yixin, LIN Shen, MA Liuyang, CHEN Long, FENG Baohua, NI Shen, WEI Xiangjin, HE Jiwai, CHEN Tianxiao. Regulation of Nitrogen Uptake and Yield in Rice by the Alanine Aminotransferase Gene OsAlaAT4 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 51-60. |
| [14] | HUANG Qina, JIANG Hongrui, YANG Jie, YU Kunyu, YANG Changdeng, LIANG Yan. Bioinformatics Analysis, Development and Application of Molecular Markers for Seed Dormancy Gene Sdr4 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 61-71. |
| [15] | CHENG Zhaoping, HE Niqing, BAI Kangcheng, LIN Shaojun, HUANG Fenghuang, LIU Junhua, CHENG Zuxin, HUANG Chengzhi, YANG Dewei. Breeding and Utilization of the Three-Line Male Sterile Line Fumeng A with Pyramided Rice Blast Resistance Genes Pigm-1 and Pid2 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 72-84. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||