
Chinese Journal OF Rice Science ›› 2026, Vol. 40 ›› Issue (2): 210-222.DOI: 10.16819/j.1001-7216.2026.250310
• Research Papers • Previous Articles Next Articles
XUE Pao1,2, WANG Youshuang1, HE Wanwan1,2, HUANG Chenbo3, ZHANG Han3, DING Zhenqian1, CHEN Qiuli1, FAN Yunxin1, DING Chengwei1, SUN Lianping3,*( ), HU Tingting1,2,*(
), HU Tingting1,2,*( )
)
Received:2025-03-12
Revised:2025-06-05
Online:2026-03-10
Published:2026-03-17
薛炮1,2, 王友霜1, 何弯弯1,2, 黄晨博3, 张涵3, 丁震乾1, 陈秋丽1, 范运新1, 丁成伟1, 孙廉平3,*( ), 胡婷婷1,2,*(
), 胡婷婷1,2,*( )
)
基金资助:XUE Pao, WANG Youshuang, HE Wanwan, HUANG Chenbo, ZHANG Han, DING Zhenqian, CHEN Qiuli, FAN Yunxin, DING Chengwei, SUN Lianping, HU Tingting. Identification and Cloning of SG5 in Rice[J]. Chinese Journal OF Rice Science, 2026, 40(2): 210-222.
薛炮, 王友霜, 何弯弯, 黄晨博, 张涵, 丁震乾, 陈秋丽, 范运新, 丁成伟, 孙廉平, 胡婷婷. 水稻颖壳不闭合基因SG5的鉴定与克隆[J]. 中国水稻科学, 2026, 40(2): 210-222.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2026.250310
| 引物名称 Primer name | 引物序列(5'→3') Primer sequence (5'→3') | 功能 Function |
|---|---|---|
| InDel 5-11F | GGAGTTCTGGTGGCTTTCC | 基因定位 Gene mapping |
| InDel 5-11R | CAGTTCCCCATTTCCCTCTA | |
| InDel 5-13F | TGAGTTTCCGGTGTTCCATA | |
| InDel 5-13R | AAGGCAAAGTCGTTCAGCTT | |
| SG5-Cas9-F | ggcaATATTGAGCCCTATATCCAG | 基因编辑 Gene editing |
| SG5- Cas9-R | aaacCTGGATATAGGGCTCAATAT | |
| SG5-1305-GUS-F | catgattacgaattcATCACCTCTGCCCAAAACCA | GUS载体构建 Construction of GUS vector |
| SG5-1305-GUS-R | tcagatctaccatggTGTAATGGTAGAATCCTGGCTTTACCA | |
| SG5-1305-GFP-F | aagtccggagctagctctagaATGACGATCTGCAGCTGTGAGG | 亚细胞定位 Subcellular localization |
| SG5-1305-GFP-R | ggtcctcgagacgtctctagaAAATCCATAGGCAGTACTGAAATAACTTTGGG | |
| Hyg-F | ACGGTGTCGTCCATCACAGTTTGCC | 潮霉素基因检测 Detection of hygromycin resistance gene |
| Hyg-R | TTCCGGAAGTGCTTGACATTGGGGA | |
| Cas9-F | CGTGGAAGATCGGTTCAACGC | 阳性株检测 Sequencing of positive plants |
| Cas9-R | CTGCCGGCCAGATTGGCA | |
| CR-SG5-F | TGATGCCAACTCCTACTGCAC | 突变位点检测 Mutation site detection |
| CR-SG5-R | ACTTGGGCTTTCCGTGTGTT | |
| UBQ10-qPCR-F | TGGTCAGTAATCAGCCAGTTTGG | qRT-PCR |
| UBQ10-qPCR-R | GCACCACAAATACTTGACGAACAG | |
| SG5-qPCR-F | GATACCTCACCAGTGGTCACTG | |
| SG5-qPCR-R | TCCTAAACGCGTAAGATGTACGG |
Table 1. Primer sequences used in this study
| 引物名称 Primer name | 引物序列(5'→3') Primer sequence (5'→3') | 功能 Function |
|---|---|---|
| InDel 5-11F | GGAGTTCTGGTGGCTTTCC | 基因定位 Gene mapping |
| InDel 5-11R | CAGTTCCCCATTTCCCTCTA | |
| InDel 5-13F | TGAGTTTCCGGTGTTCCATA | |
| InDel 5-13R | AAGGCAAAGTCGTTCAGCTT | |
| SG5-Cas9-F | ggcaATATTGAGCCCTATATCCAG | 基因编辑 Gene editing |
| SG5- Cas9-R | aaacCTGGATATAGGGCTCAATAT | |
| SG5-1305-GUS-F | catgattacgaattcATCACCTCTGCCCAAAACCA | GUS载体构建 Construction of GUS vector |
| SG5-1305-GUS-R | tcagatctaccatggTGTAATGGTAGAATCCTGGCTTTACCA | |
| SG5-1305-GFP-F | aagtccggagctagctctagaATGACGATCTGCAGCTGTGAGG | 亚细胞定位 Subcellular localization |
| SG5-1305-GFP-R | ggtcctcgagacgtctctagaAAATCCATAGGCAGTACTGAAATAACTTTGGG | |
| Hyg-F | ACGGTGTCGTCCATCACAGTTTGCC | 潮霉素基因检测 Detection of hygromycin resistance gene |
| Hyg-R | TTCCGGAAGTGCTTGACATTGGGGA | |
| Cas9-F | CGTGGAAGATCGGTTCAACGC | 阳性株检测 Sequencing of positive plants |
| Cas9-R | CTGCCGGCCAGATTGGCA | |
| CR-SG5-F | TGATGCCAACTCCTACTGCAC | 突变位点检测 Mutation site detection |
| CR-SG5-R | ACTTGGGCTTTCCGTGTGTT | |
| UBQ10-qPCR-F | TGGTCAGTAATCAGCCAGTTTGG | qRT-PCR |
| UBQ10-qPCR-R | GCACCACAAATACTTGACGAACAG | |
| SG5-qPCR-F | GATACCTCACCAGTGGTCACTG | |
| SG5-qPCR-R | TCCTAAACGCGTAAGATGTACGG |
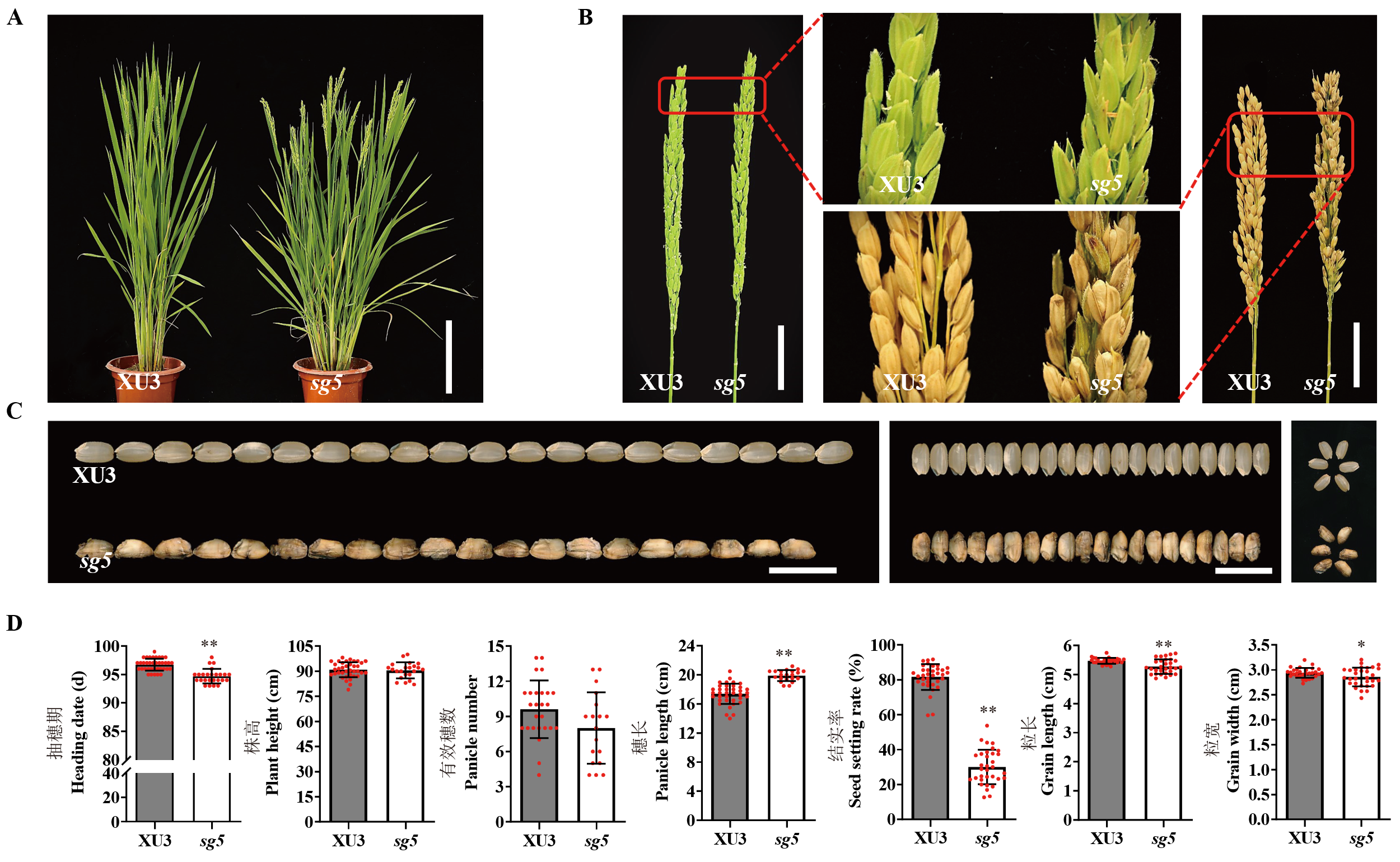
Fig. 1. Phenotypic characterization of wildtype (Xudao 3, XU3) and sg5 A, Characterization of plant type in wild type and sg5. Scale bar=20 cm. B, Glume morphology and panicle morphology in wild type and sg5. Scale bar=5 cm. C, Comparison of grain size in wild type and sg5. Scale bar=10 mm. D, Comparison of heading date, plant height, panicle number, panicle length, seed setting rate, grain length and grain width in wild type and sg5. Data are given as Mean ± SD (n≥20). Student’s t-test was used to generate P value. *, P < 0.05; **, P < 0.01.

Fig. 2. Developmental morphological changes of glumes and lodicules in wild type (XU3) and sg5 at various time points around flowering A, Characterization of panicle before flowering(A), glume before flowering(B), panicle after flowering(C), glume after flowering(D) in the wild type and sg5(Scale bar=5 cm). E, Florets of consistent early growth from wild type and sg5, showing morphological changes of glumes and lodicules at various time points around flowering. Scale bar = 20 μm. The white boxes indicate the corresponding florets.
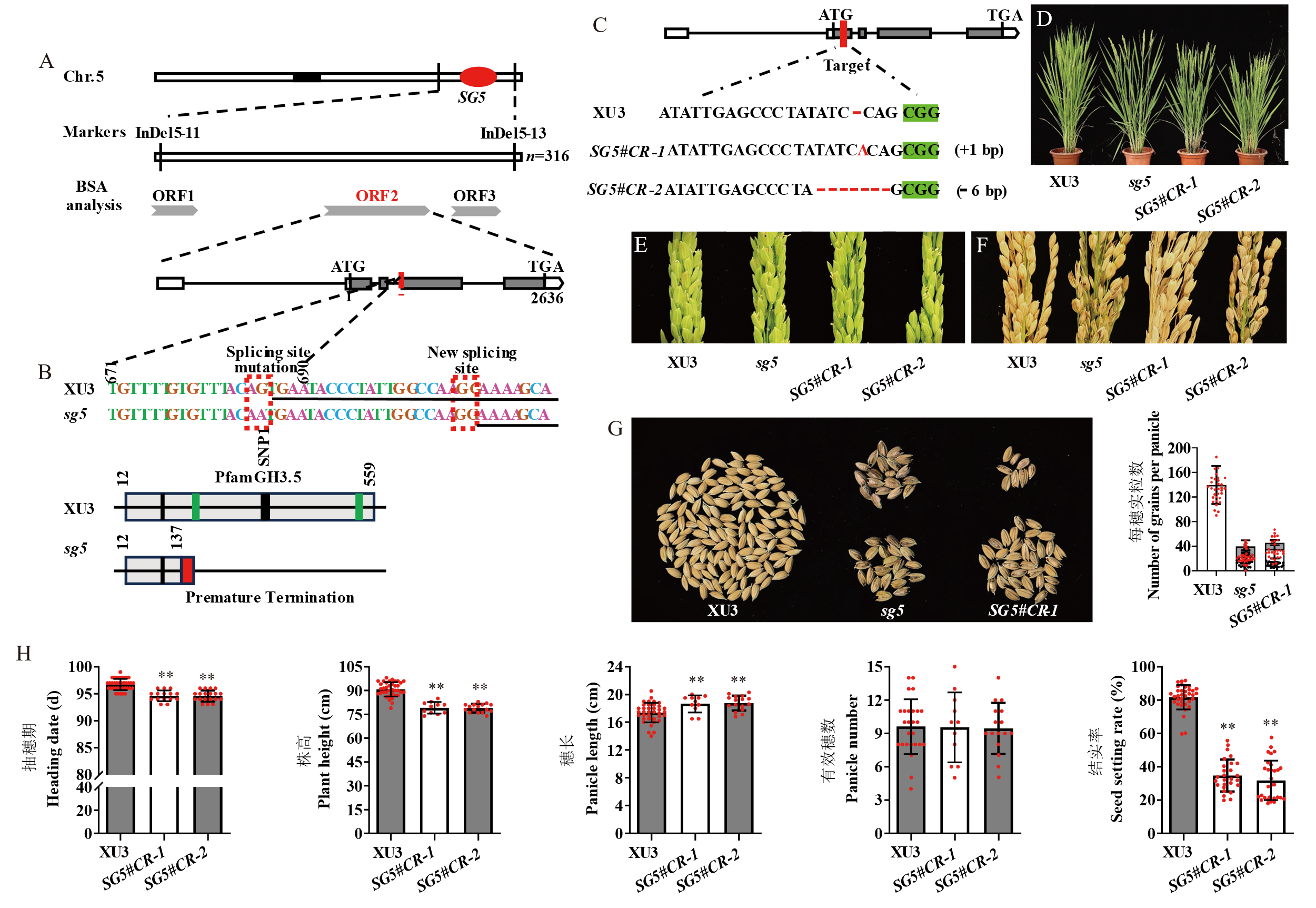
Fig. 3. Map-based cloning and functional verification of SG5 A, Map-based cloning of SG5. B, Characterization of DNA and protein sequences in Xudao 3(XU3) and sg5. Underlined nucleotides are the sequence of the first half of exons 4 in XU3, others are at the terminal of the intron 3. Red boxes are the splicing site mutation and new splicing sites. Black vertical line, the site of jasmonate. Green vertical line, the site of L-alpha-amino acid. C, Gene editing and homozygous mutant sequences. D, Characterization of plant type in XU3, sg5 mutant, SG5#CR-1, and SG5#CR-2. Scale bar=20 cm. E, Glume morphology of XU3, sg5 mutant, SG5#CR-1, and SG5#CR-2; F, Panicle morphology of wild type, sg5 mutant, SG5#CR-1, and SG5#CR-2. Scale bar=5 cm. G, Number of grains per panicle in XU3, sg5 mutant, and SG5#CR-1; Red dot, Normal seeds. Black dot, Moldy seeds. H, Comparison of heading date, plant height, panicle length, panicle number, and seed setting rate in XU3, SG5#CR-1, and SG5#CR-2. Red dot, Number of plants investigated. Data are given as Mean ± SD (n≥10). Student’s t-test was used to generate P value. * P < 0.05; ** P < 0.01.
| 差异种类 Category | 差异基因数目 Number of differential genes |
|---|---|
| 基因上游 Upstream | 638 |
| 功能获得型 Stop gain | 6 |
| 功能缺失型 Stop loss | 2 |
| 5' UTR | 10 |
| 3' UTR | 14 |
| 保守结构缺失/插入 Conservative inframe deletion / insertion | 4 |
| 散乱结构缺失/插入 Disruptive inframe deletion / insertion | 3 |
| 移码突变 Frameshift variant | 23 |
| 错义突变 Missense variant | 119 |
| 同义突变 Synonymous variant | 98 |
| 剪接受体变异 Splice acceptor variant & intron variant | 4 |
| 内含子变异 Intronic | 60 |
| 基因间变异 Intergenic region | 72 |
| 基因下游 Downstream | 212 |
| 合计 Total | 1265 |
Table 2. Summary of differentially expressed genes detected using BSA analysis
| 差异种类 Category | 差异基因数目 Number of differential genes |
|---|---|
| 基因上游 Upstream | 638 |
| 功能获得型 Stop gain | 6 |
| 功能缺失型 Stop loss | 2 |
| 5' UTR | 10 |
| 3' UTR | 14 |
| 保守结构缺失/插入 Conservative inframe deletion / insertion | 4 |
| 散乱结构缺失/插入 Disruptive inframe deletion / insertion | 3 |
| 移码突变 Frameshift variant | 23 |
| 错义突变 Missense variant | 119 |
| 同义突变 Synonymous variant | 98 |
| 剪接受体变异 Splice acceptor variant & intron variant | 4 |
| 内含子变异 Intronic | 60 |
| 基因间变异 Intergenic region | 72 |
| 基因下游 Downstream | 212 |
| 合计 Total | 1265 |
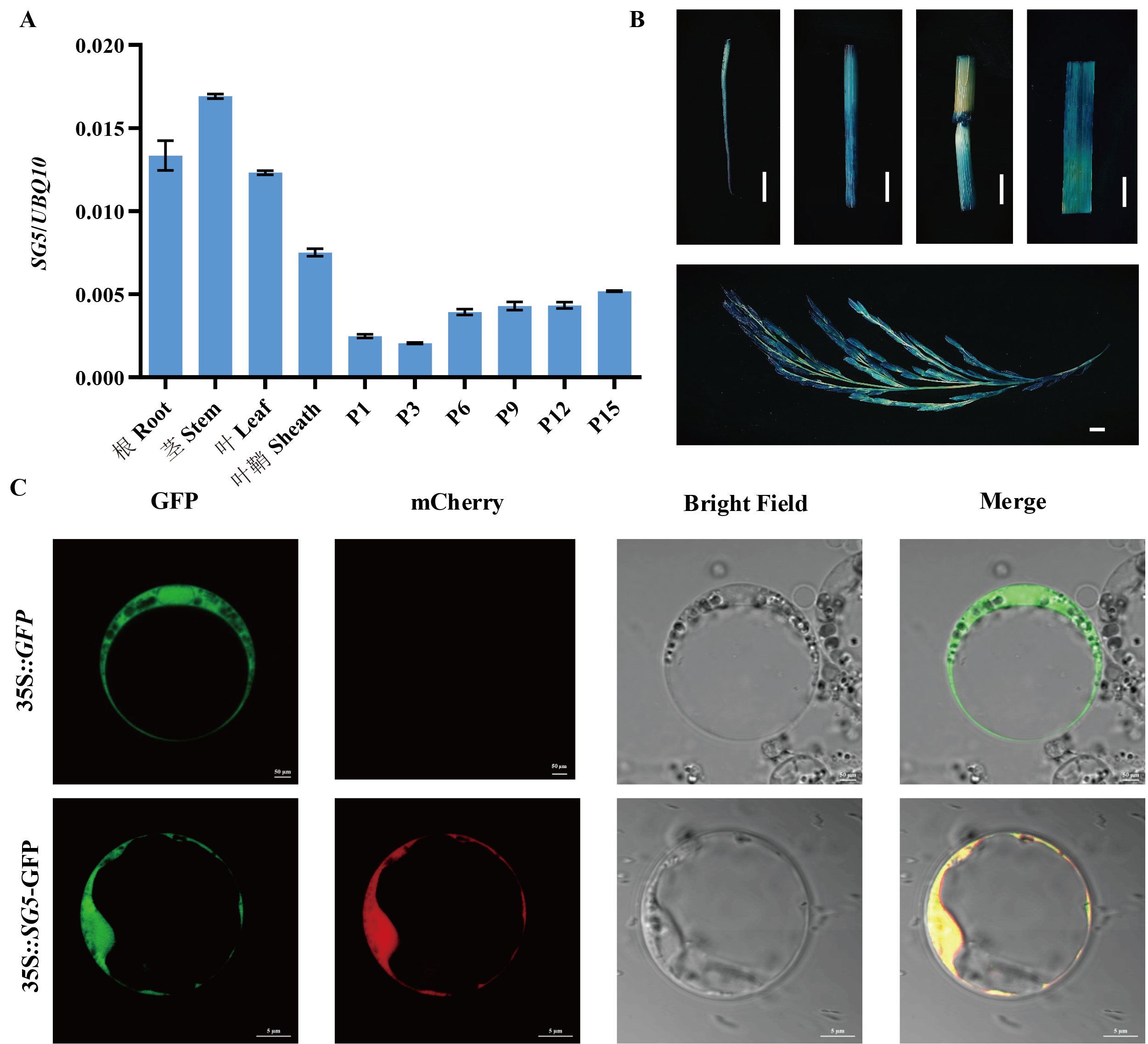
Fig. 4. Expression pattern analysis of SG5 and subcellular localization A, qRT-PCR analysis of SG5 gene expression in various tissues. Data are given as Mean ± SD (n=3). B, Histochemical analysis of GUS activity in various tissues.( Scale bar=10 mm). C, Subcellular localization of SG5 in rice protoplast. 35S::GFP(scale bar=50 μm). 35S::SG5-GFP(scale bar=5 μm).
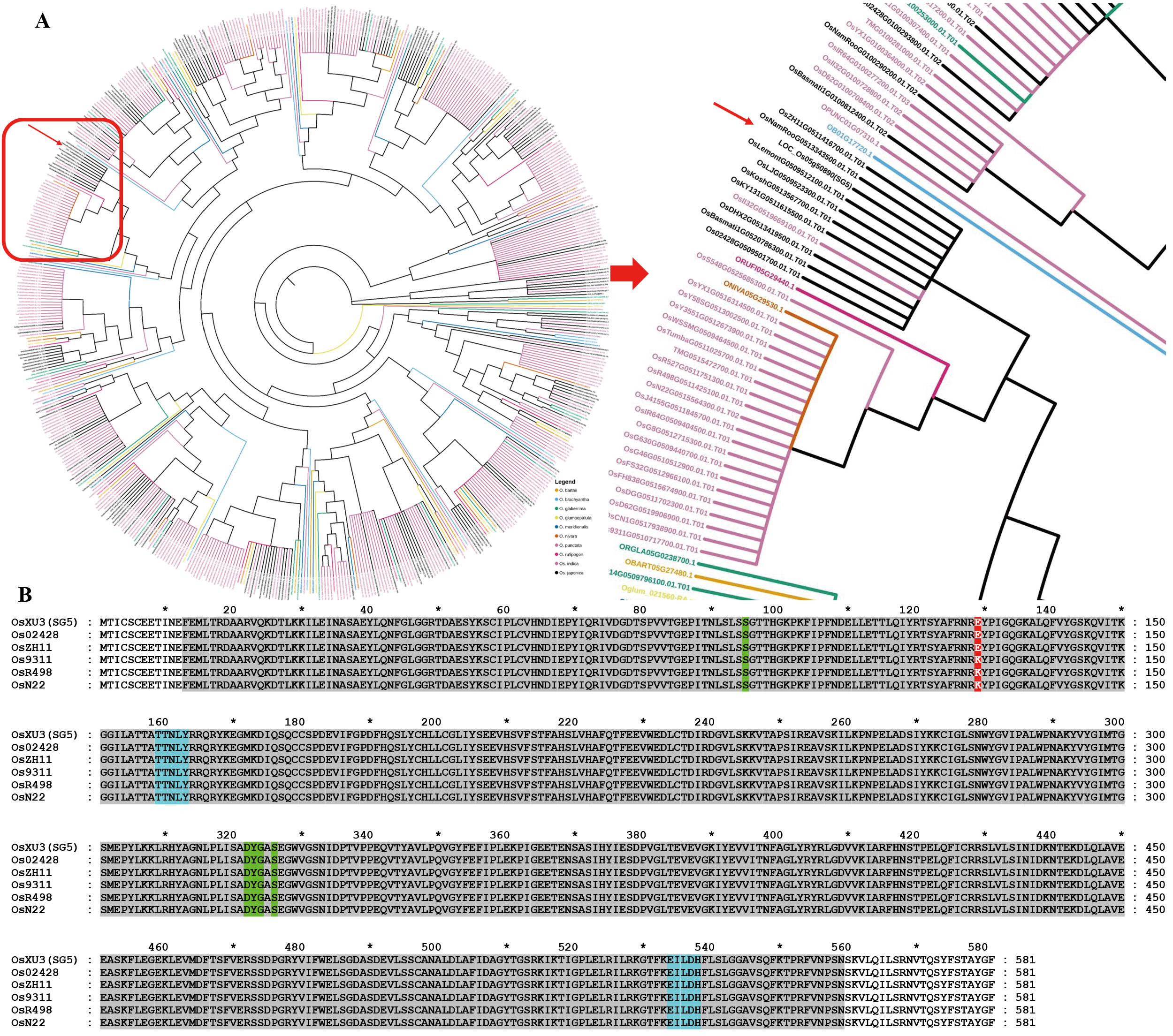
Fig. 5. Phylogenetic tree and multiple sequence alignment analysis of SG5 in rice A, Phylogenetic tree analysis of SG5 in rice. Red frame, Rice resources with the GH3-5 domain. Red arrow, SG5 protein. B, Multiple sequence alignment of SG5 in rice. Gray frame, The domain of GH3-5. Green frame, The site of jasmonate. Blue frame, The site of L-alpha-amino acid.
| [1] | Xing Y Z, Zhang Q F. Genetic and molecular bases of rice yield[J]. Annual Review of Plant Biology, 2010, 61: 421-442. |
| [2] | Ren D Y, Ding C Q, Qian Q. Molecular bases of rice grain size and quality for optimized productivity[J]. Science Bulletin, 2023, 68(3): 314-350. |
| [3] | Theissen G, Melzer R. Molecular mechanisms underlying origin and diversification of the Angiosperm flower[J]. Annals of Botany, 2007, 100(3): 603-619. |
| [4] | Soltis D E, Chanderbali A S, Kim S, Buzgo M, Soltis P S. The ABC model and its applicability to basal angiosperms[J]. Annals of Botany, 2007, 100(2): 155-163. |
| [5] | Maoileidigh D S O, Graciet E, Wellmer F. Gene networks controlling Arabidopsis thaliana flower development[J]. New Phytologist, 2014, 201: 16-30. |
| [6] | Krizek B A, Fletcher J C. Molecular mechanisms of flower development: An armchair guide[J]. Nature Reviews Genetics, 2005, 6(9): 688-698. |
| [7] | Causier B, Schwarz-Sommer Z, Davies B. Floral organ identity: 20 years of ABCs[J]. Seminars in Cell & Developmental Biology, 2010, 21(1): 73-79. |
| [8] | 郭爽, 李云峰, 任德勇, 王增, 杜青, 何光华. 水稻内稃扭曲突变体palea distortion 1 (pd1) 的鉴定与基因定位[J]. 分子植物育种, 2011, 9(3): 256-260. |
| Guo S, Li Y F, Ren D Y, Wang Z, Du Q, He G H. Identification and gene mapping of a palea distortion 1 (pd1) mutant in rice (Oryza sativa L.)[J]. Molecular Plant Breeding, 2011, 9(3): 256-260. (in Chinese with English abstract) | |
| [9] | Jin Y, Luo Q, Tong H N, Wang A J, Cheng Z J, Tang J F, Li D Y, Zhao X F, Li X B, Wan J M, Jiao Y L, Chu C C, Zhu L H. An AT-hook gene is required for palea formation and floral organ number control in rice[J]. Developmental Biology, 2011, 359(2): 277-288. |
| [10] | 陈东, 毛毕刚, 潘银林, 彭彦, 韶也, 吴天昊, 赵炳然. 水稻颖花畸形突变体(gm(t))的遗传分析与基因定位[J]. 分子植物育种, 2019, 17(15): 5026-5031. |
| Chen D, Mao B G, Pan Y L, Peng Y, Shao Y, Wu T H, Zhao B R. Genetic analysis and gene mapping of glume malformation mutant(gm(t)) in rice[J]. Molecular Plant Breeding, 2019, 17(15): 5026-5031. (in Chinese with English abstract) | |
| [11] | 薛大伟. 两个水稻花器官突变体的遗传分析[D]. 北京: 中国农业科学院, 2006. |
| Xue D W. The genetic analysis of two floral organ mutants from rice[D]. Chinese Academy of Agricultural Sciences, 2006. Beijing: Chinese Academy of Agricultural Sciences, 2006. (in Chinese with English abstract) | |
| [12] | Jeon J S, Jang S, Lee S, Nam J, Kim C, Lee S H, Chung Y Y, Kim S R, Lee Y H, Cho Y G, An G. leafy hull sterile1 is a homeotic mutation in a rice MADS box gene affecting rice flower development[J]. The Plant Cell, 2000, 12(6): 871-884. |
| [13] | Agrawal G K, Abe K, Yamazaki M, Miyao A, Hirochika H. Conservation of the E-function for floral organ identity in rice revealed by the analysis of tissue culture-induced loss-of-function mutants of the OsMADS1 gene[J]. Plant Molecular Biology, 2005, 59(1): 125-135. |
| [14] | Prasad K, Parameswaran S, Vijayraghavan U. OsMADS1, a rice MADS‐box factor, controls differentiation of specific cell types in the lemma and palea and is an early‐acting regulator of inner floral organs[J]. The Plant Journal, 2005, 43(6): 915-928. |
| [15] | Hu Y, Liang W Q, Yin C S, Yang X L, Ping B Z, Li A X, Jia R, Chen M J, Luo Z J, Cai Q, Zhao X X, Zhang D B, Yuan Z. Interactions of OsMADS1 with floral homeotic genes in rice flower development[J]. Molecular plant, 2015, 8(9): 1366-1384. |
| [16] | Pelucchi N, Fornara F, Favalli C, Masiero S, Lago C, P E M, Colombo L, Kater M M. Comparative analysis of rice MADS-box genes expressed during flower development[J]. Sexual Plant Reproduction, 2002, 15(3): 113-122. |
| [17] | Wang K J, Tang D, Hong L L, Xu W Y, Huang J, Li M, Gu M H, Xue Y B, Cheng Z K. DEP and AFO regulate reproductive habit in rice[J]. PLoS Genetics, 2010, 6(1): e1000818. |
| [18] | Kobayashi K, Yasuno N, Sato Y, Yoda M, Yamazaki R, Kimizu M, Yoshida H, Nagamura Y, Kyozuka J. Inflorescence meristem identity in rice is specified by overlapping functions of three AP1/FUL-like MADS box genes and PAP2 a SEPALLATA MADS box gene[J]. The Plant Cell, 2012, 24(5): 1848-1859. |
| [19] | Yadav S R, Prasad K, Vijayraghavan U. Divergent regulatory OsMADS2 functions control size, shape and differentiation of the highly derived rice floret second-whorl organ[J]. Genetics, 2007, 176(1): 283-294. |
| [20] | Yao S G, Ohmori S, Kimizu M, Yoshida H. Unequal genetic redundancy of rice PISTILLATA orthologs, OsMADS2 and OsMADS4, in lodicule and stamen development[J]. Plant & Cell Physiology, 2008, 49(5): 853-857. |
| [21] | Yu X Q, Xia S S, Xu Q K, Cui Y J, Gong M, Zeng D L, Zhang Q, Shen L, Jiao G A, Gao Z Y, Hu J, Zhang G H, Zhu L, Guo L B, Ren D Y, Qian Q. ABNORMAL FLOWER AND GRAIN 1 encodes OsMADS6 and determines palea identity and affects rice grain yield and quality[J]. Science China Life Sciences, 2020, 63(2): 228-238. |
| [22] | Ohmori S, Kimizu M, Sugita M, Miyao A, Hirochika H, Uchida E, Nagato Y, Yoshida H. MOSAIC FLORAL ORGANS1, an AGL6 -like MADS box gene, regulates floral organ identity and meristem fate in rice[J]. The Plant Cell, 2009, 21(10): 3008-3025. |
| [23] | Zhang J, Nallamilli B R, Mujahid H, Peng Z H. OsMADS6 plays an essential role in endosperm nutrient accumulation and is subject to epigenetic regulation in rice (Oryza sativa)[J]. The Plant Journal, 2010, 64(4): 604-617. |
| [24] | Duan Y L, Xing Z, Diao Z J, Xu W Y, Li S P, Du X Q, Wu G H, Wang C L, Lan T, Meng Z, Liu H Q, Wang F, Wu W R, Xue Y B. Characterization of Osmads6-5, a null allele, reveals that OsMADS6 is a critical regulator for early flower development in rice (Oryza sativa L.)[J]. Plant Molecular Biology, 2012, 80(4-5): 429-442. |
| [25] | Wang H H, Zhang L, Cai Q, Hu Y, Jin Z M, Zhao X X, Fan W, Huang Q M, Luo Z J, Chen M J, Zhang D B, Yuan Z. OsMADS32 interacts with PI-like proteins and regulates rice flower development[J]. Journal of Integrative Plant Biology, 2015, 57(5): 504-513. |
| [26] | Luo Q, Zhou K D, Zhao X F, Zeng Q C, Xia H A, Zhai W X, Xu J C, Wu X J, Yang H S, Zhu L H. Identification and fine mapping of a mutant gene for palealess spikelet in rice[J]. Planta, 2005, 221(2): 222-230. |
| [27] | Xiao H, Tang J F, Li Y F, Wang W M, Li X B, Jin L, Xie R, Luo H F, Zhao X F, Meng Z, He G H, Zhu L H. STAMENLESS 1, encoding a single C2H2 zinc finger protein, regulates floral organ identity in rice[J]. The Plant Journal, 2009, 59(5): 789-801. |
| [28] | Yan D W, Zhang X M, Zhang L, Ye S H, Zeng L J, Liu J Y, Li Q, He Z H. CURVED CHIMERIC PALEA 1 encoding an EMF1-like protein maintains epigenetic repression of OsMADS58in rice palea development[J]. The Plant Journal, 2015, 82(1): 12-24. |
| [29] | Zheng M, Wang Y H, Wang Y L, Wang C M, Ren Y L, Lv J, Peng C, Wu T, Liu K, Zhao S L, Liu X, Guo X P, Jiang L, Terzaghi W, Wan J M. DEFORMED FLORAL ORGAN1 (DFO1) regulates floral organ identity by epigenetically repressing the expression of OsMADS58 in rice (Oryza sativa)[J]. The New Phytologist, 2015, 206(4): 1476-1490. |
| [30] | Yuan Z, Gao S, Xue D W, Luo D, Li L T, Ding S Y, Yao X, Wilson Z A, Qian Q, Zhang D B. RETARDED PALEA1 controls palea development and floral zygomorphy in rice[J]. Plant Physiology, 2009, 149(1): 235-244. |
| [31] | Zeng D D, Qin R, Alamin M, Liang R, Yang C C, Jin X L, Shi C H. DBOP specifies palea development by suppressing the expansion of the margin of palea in rice[J]. Genes and Genomics, 2016, 38(11): 1095-1103. |
| [32] | Lee D Y, Lee J, Moon S, Park S Y, An G. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem[J]. The Plant Journal, 2007, 49(1): 64-78. |
| [33] | Ren D Y, Li Y F, Zhao F M, Sang X C, Shi J Q, Wang N, Guo S, Ling Y H, Zhang C W, Yang Z L, He G H. MULTI-FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice[J]. Plant Physiology, 2013, 162(2): 872-884. |
| [34] | Ren D Y, Rao Y C, Wu L W, Xu Q K, Li Z Z, Yu H P, Zhang Y, Leng Y J, Hu J, Zhu L, Gao Z Y, Dong G J, Zhang G H, Guo L B, Zeng D L, Qian Q. The pleiotropic ABNORMAL FLOWER AND DWARF1 affects plant height, floral development and grain yield in rice[J]. Journal of Integrative Plant Biology, 2016, 58(6): 529-539. |
| [35] | Yan D W, Zhou Y, Ye S H, Zeng L J, Zhang X M, He Z H. BEAK-SHAPED GRAIN 1/TRIANGULAR HULL 1, a DUF640 gene, is associated with grain shape, size and weight in rice[J]. Science China Life Sciences, 2013, 56(3): 275-283. |
| [36] | Livak K J, Schmittgen T D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCTmethod[J]. Methods, 2001, 25(4): 402-408. |
| [37] | 王忠, 顾蕴洁, 高煜珠. 水稻开颖机理的探讨: Ⅲ.浆片的结构及其在开颖过程中内含物的变化[J]. 作物学报, 1991, 17(2): 96-101, 161-162. |
| Wang Z, Gu Y J, Gao Y Z. Studies on the mechanism of the anthesis of rice Ⅲ.Structure of the lodicule and changes of its contents during flowering[J]. Acta Agronomica Sinica, 1991, 17(2): 96-101, 161-162. (in Chinese with English abstract) | |
| [38] | 秦鹏. 水稻颖壳发育基因hzp和ug1的研究[D]. 长沙: 湖南农业大学, 2021. |
| Qin P. The study on genes hzp and ug1 controlling rice glume development[D]. Changsha: Hunan Agricultural University, 2021. (in Chinese with English abstract) | |
| [39] | Terol J, Domingo C, Talón M. The GH3 family in plants: Genome wide analysis in rice and evolutionary history based on EST analysis[J]. Gene, 2006, 371(2): 279-290. |
| [40] | Riemann M, Riemann M, Takano M. Rice JASMONATE RESISTANT 1 is involved in phytochrome and jasmonate signalling[J]. Plant, Cell & Environment, 2008, 31(6): 783-792. |
| [41] | Xiao Y G, Chen Y, Charnikhova T, Mulder P P J, Heijmans J, Hoogenboom A, Agalou A, Michel C, Morel J B, Dreni L, Kater M M, Bouwmeester H, Wang M, Zhu Z, Ouwerkerk P B F. OsJAR1 is required for JA-regulated floret opening and anther dehiscence in rice[J]. Plant Molecular Biology, 2014, 86(1/2): 19-33. |
| [42] | Jain M, Kaur N, Tyagi A K, Khurana J P. The auxin-responsive GH3 gene family in rice (Oryza sativa)[J]. Functional & Integrative Genomics, 2006, 6(1): 36-46. |
| [43] | Zhao S Q, Xiang J J, Xue H W. Studies on the rice LEAF INCLINATION1 (LC1), an IAA-amido synthetase, reveal the effects of auxin in leaf inclination control[J]. Molecular Plant, 2013, 6(1): 174-187. |
| [44] | Du H, Wu N, Fu J, Wang S P, Li X H, Xiao J H, Xiong L Z. A GH3 family member, OsGH3-2, modulates auxin and abscisic acid levels and differentially affects drought and cold tolerance in rice[J]. Journal of Experimental Botany, 2012, 63(18): 6467-6480. |
| [45] | Fu J, Yu H H, Li X H, Xiao J H, Wang S P. Rice GH3 gene family: Regulators of growth and development[J]. Plant Signaling Behavior, 2011, 6(4): 570-574. |
| [46] | Kong W L, Zhong H, Deng X X, Gautam M, Gong Z Y, Zhang Y, Zhao G Q, Liu C, Li Y S. Evolutionary analysis of GH3 genes in six Oryza Species/Subspecies and their expression under salinity stress in Oryza sativa ssp. japonica[J]. Plants, 2019, 8(2): 30. |
| [47] | Mao C J, He J M, Liu L, Deng Q M, Yao X F, Liu C M, Qiao Y L, Li P, Ming F. OsNAC2 integrates auxin and cytokinin pathways to modulate rice root development[J]. Plant Biotechnology Journal, 2020, 18(2): 429-442. |
| [48] | Ding X H, Cao Y L, Huang L L, Zhao J, Xu C G, Li X H, Wang S P. Activation of the Indole-3-Acetic Acid-Amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice[J]. The Plant Cell, 2008, 20(1): 228-240. |
| [49] | Yadav S R, Khanday I, Majhi B B, Veluthambi K, Vijayraghavan U. Auxin-responsive OsMGH3, a common downstream target of OsMADS1 and OsMADS6, controls rice floret fertility[J]. Plant and Cell Physiology, 2011, 52(12): 2123-2135. |
| [50] | Fukumoto K, Alamgir K M, Yamashita Y, Mori I C, Matsuura H, Galis I. Response of rice to insect elicitors and the role of OsJAR1 in wound and herbivory-induced JA-Ile accumulation[J]. Journal of Integrative Plant Biology, 2013, 55(8): 775-784. |
| [51] | Song S S, Qi T C, Huang H, Ren Q C, Wu D W, Chang C Q, Peng W, Liu Y, Peng J R, Xie D X. The jasmonate-ZIM domain proteins interact with the R2R3-MYB transcription factors MYB21 and MYB24 to affect jasmonate-regulated stamen development in Arabidopsis[J]. The Plant Cell, 2011, 23(3): 1000-1013. |
| [52] | Wakuta S, Suzuki E, Saburi W, Matsuura H, Nabeta K, Imai R, Matsui H. OsJAR1 and OsJAR2 are jasmonyl-L-isoleucine synthases involved in wound- and pathogen-induced Jasmonic acid signalling[J]. Biochemical and Biophysical Research Communications, 2011, 409: 634-639. |
| [53] | Zhang S N, Wang S K, Xu Y X, Yu C L, Shen C J, Qian Q, Geisler M, Jiang D A, Qi Y H. The auxin response factor, OsARF19, controls rice leaf angles through positively regulating OsGH3-5 and OsBRI1[J]. Plant, Cell and Environment, 2015, 38: 638-654. |
| [54] | Ding W Y, Gou Y J, Li Y J, Li J J, Fang Y D, Liu X P, Zhu X Y, Ye R J, Heng Y Q, Wang H Y, Shen R X. A jasmonate-mediated regulatory network modulates diurnal floret opening time in rice[J]. New phytologist, 2024, 244(1): 176-191. |
| [55] | 王忠, 顾蕴洁, 高煜珠. 水稻开颖机理的探讨: V.不育系与可育系浆片和花丝结构的比较[J]. 作物学报, 1994, 20(1): 13-17, 129-131. |
| Wang Z, Gu Y J, Gao Y Z. Studies on the mechanism of the anthesis of rice V. comparison of lodicule and filament structure between sterile line and fertile line[J]. Acta Agronomica Sinica, 1994, 20(1): 13-17, 129-131. (in Chinese with English abstract) |
| [1] | YANG Dabing, DU Xueshu, LI Jinbo, XIA Mingyuan, HU Liang, SHI Huan, WAN Bingliang. Advances in Molecular Mechanism and Breeding Application of Heading Date Regulation in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 145-154. |
| [2] | NI Chen, ZHANG Jiahao, ZHU Changjin, XU Jiwei, HU Qiuqian, HUO Zhongyang, DAI Qigen, XU Ke, LI Guohui. Research Progress on the Formation and Regulation of Rice Source, Flow and Sink and Their Influencing Factors [J]. Chinese Journal OF Rice Science, 2026, 40(2): 155-170. |
| [3] | WANG Mengning, XIE Keran, GAO Ti, WANG Zhenmei, XIONG Dongliang, CUI Kehui. Research Progress on Effects of High Temperature on Rice Grain Weight Formation and Cultivation Strategies [J]. Chinese Journal OF Rice Science, 2026, 40(2): 171-180. |
| [4] | LUO Xiaoyun, ZHENG Xingfei, PENG Xuanguo, YU Qizhi, DONG Hualin, YIN Desuo, WANG Hongbo, HU Jianlin, XUE Lian, HU Peng, XU Deze. Research on Rice Lodging Resistance: Current Status, Challenges, and Future Directions [J]. Chinese Journal OF Rice Science, 2026, 40(2): 181-195. |
| [5] | DUAN Min, XIE Liujie, YUE Yani, HUANG Shanjun. Development of High-quality Aromatic japonica Rice Lines by Using CRISPR/Cas9 Technology [J]. Chinese Journal OF Rice Science, 2026, 40(2): 235-243. |
| [6] | ZHANG Mengke, LU Jiayu, HE Jin, XU Xue, WU Shuang, WANG Peiran, CHEN Ruofan, JIN Qing, WANG Xiufeng. Development and Application of Functional Markers for Purity Identification of Wild-abortive Three-line Hybrid Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 244-252. |
| [7] | WANG Weiying, ZHAO Hong, CHEN Yu, YAO Xiaoming, LU Jianfei, GUO Qianshuang, DU Yongjun. Disruption Efficacy of Intelligent Active Aerosol Sex Pheromone on Mating Behavior of Major Lepidopteran Pests in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(2): 264-272. |
| [8] | ZHENG Xusong, XU Hongxing, LÜ Haitao, LI Jianzhong, LÜ Zhongxian, LU Yanhui. Feeding and Damage Characteristics of Pomacea canaliculata on Direct-seeded Rice Seedlings of Different Varieties [J]. Chinese Journal OF Rice Science, 2026, 40(2): 273-280. |
| [9] | LI Xingyi, CHEN Ling, SHAO Jiantao, XIAO Suqin, LI Jinlu, FU Huixian, YIN Fuyou, ZHANG Jianhong, CHENG Zaiquan, LIU Li. Progress in Molecular Genetic Research on Co-regulation of Grain Yield and Starch Quality in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 1-17. |
| [10] | MIAO Zhening, CHEN Jijin, LI Ziming, LIU Yi, LUO Lijun. Advances and Prospects in Breeding Bacterial Blight-resistant Rice in South China [J]. Chinese Journal OF Rice Science, 2026, 40(1): 18-26. |
| [11] | YUE Xuanyu, XIE Wenya, FENG Zhiming, CHEN Zongxiang, HU Keming, ZUO Shimin. Function of OsERF93 in Regulating Resistance to Sheath Blight in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 37-50. |
| [12] | WANG Yixin, LIN Shen, MA Liuyang, CHEN Long, FENG Baohua, NI Shen, WEI Xiangjin, HE Jiwai, CHEN Tianxiao. Regulation of Nitrogen Uptake and Yield in Rice by the Alanine Aminotransferase Gene OsAlaAT4 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 51-60. |
| [13] | HUANG Qina, JIANG Hongrui, YANG Jie, YU Kunyu, YANG Changdeng, LIANG Yan. Bioinformatics Analysis, Development and Application of Molecular Markers for Seed Dormancy Gene Sdr4 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 61-71. |
| [14] | CHENG Zhaoping, HE Niqing, BAI Kangcheng, LIN Shaojun, HUANG Fenghuang, LIU Junhua, CHENG Zuxin, HUANG Chengzhi, YANG Dewei. Breeding and Utilization of the Three-Line Male Sterile Line Fumeng A with Pyramided Rice Blast Resistance Genes Pigm-1 and Pid2 [J]. Chinese Journal OF Rice Science, 2026, 40(1): 72-84. |
| [15] | LIU Yaping, DONG Yici, ZHENG Junyan, QIU Xuan, LIU Pengcheng, YE Yafeng, LIU Binmei, CHEN Xifeng, MA Bojun. Phenotypic Identification and Gene Fine Mapping of the Lesion Mimic and Early Senescence Mutant lmes7 in Rice [J]. Chinese Journal OF Rice Science, 2026, 40(1): 85-94. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||