
Chinese Journal OF Rice Science ›› 2020, Vol. 34 ›› Issue (5): 383-396.DOI: 10.16819/j.1001-7216.2020.0302
• Research Papers • Next Articles
Xiaohan TANG1, Shijia LIU1, Xi LIU1, Yunlu TIAN1, Yunlong WANG1, Xuan TENG1, Erchao DUAN1, Yuanyan ZHANG1, Ling JIANG1, Wenwei ZHANG1, Yihua WANG1,*( ), Jianmin WAN1,2
), Jianmin WAN1,2
Received:2020-03-01
Revised:2020-03-18
Online:2020-09-10
Published:2020-09-10
Contact:
Yihua WANG
唐小涵1, 刘世家1, 刘喜1, 田云录1, 王云龙1, 滕烜1, 段二超1, 张元燕1, 江玲1, 张文伟1, 王益华1,*( ), 万建民1,2
), 万建民1,2
通讯作者:
王益华
基金资助:CLC Number:
Xiaohan TANG, Shijia LIU, Xi LIU, Yunlu TIAN, Yunlong WANG, Xuan TENG, Erchao DUAN, Yuanyan ZHANG, Ling JIANG, Wenwei ZHANG, Yihua WANG, Jianmin WAN. Tryptophanyl-tRNA Synthetase Gene WRS1 Regulates Rice Seed Development[J]. Chinese Journal OF Rice Science, 2020, 34(5): 383-396.
唐小涵, 刘世家, 刘喜, 田云录, 王云龙, 滕烜, 段二超, 张元燕, 江玲, 张文伟, 王益华, 万建民. 色氨酰-tRNA合成酶基因WRS1调控水稻种子发育[J]. 中国水稻科学, 2020, 34(5): 383-396.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2020.0302
| 引物名称 Primer name | 正向引物序列 Forward (5′-3′) | 反向引物序列 Reserve (5′-3′) | 产物大小 Product size / bp |
|---|---|---|---|
| N4 | TCCCTGATTCCATTTCTC | TTATGCCTCCATTGTTGA | 404 |
| N10 | GGCCATCTCATCCCCTTCAT | AATTGAGTCCCGGCCACC | 633 |
| Hyg | CTATTTCTTTGCCCTCGGAC | ATGCCTGAACTCACCGCGAC | 1083 |
Table 1 Markers used for identification in wrs1 transgenic lines.
| 引物名称 Primer name | 正向引物序列 Forward (5′-3′) | 反向引物序列 Reserve (5′-3′) | 产物大小 Product size / bp |
|---|---|---|---|
| N4 | TCCCTGATTCCATTTCTC | TTATGCCTCCATTGTTGA | 404 |
| N10 | GGCCATCTCATCCCCTTCAT | AATTGAGTCCCGGCCACC | 633 |
| Hyg | CTATTTCTTTGCCCTCGGAC | ATGCCTGAACTCACCGCGAC | 1083 |
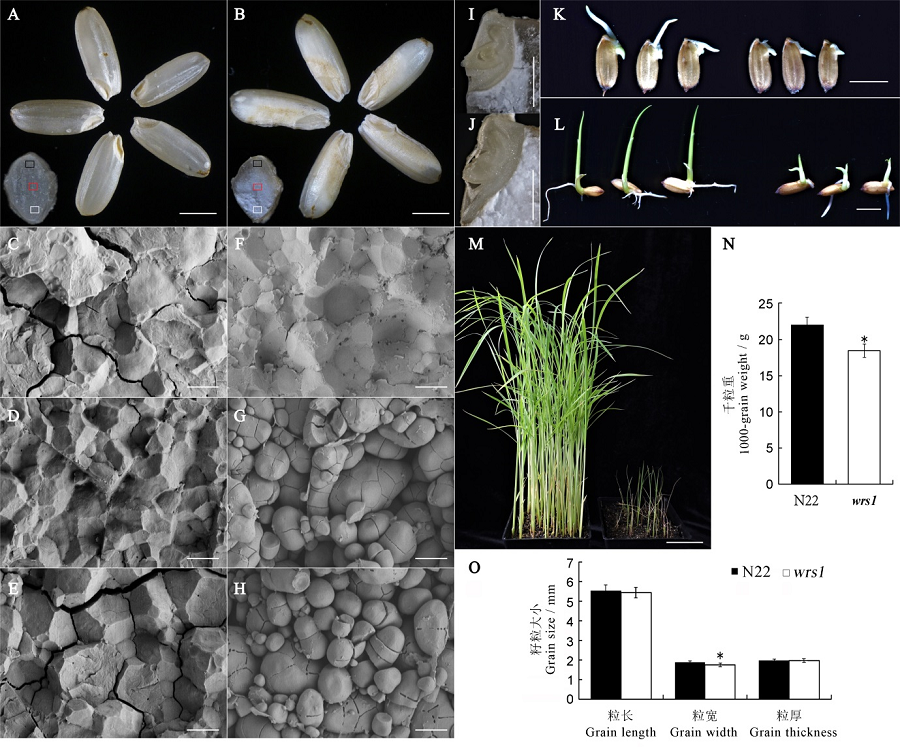
Fig. 1. Phenotypic analysis of wild-type and the wrs1 mutant. A, B, Comparison of mature seeds of wild-type (A) and the wrs1 mutant (B). Bars = 2 mm. C~H, Scanning electron microscopic (SEM) analysis of cross-sections of wild-type (C-E) and wrs1 mutant (F-H) mature seeds. C-E and F-H correspond to the black frame, red frame, and white frames in wild-type and wrs1, respectively. Bars = 5 μm. I, J, Embryos of wild-type (I) and wrs1 mutant (J) seeds after imbibition (30℃, 10 h). Bars = 1 mm. K~M, Developmental status of wild-type and the wrs1 mutant. 24 hours after seed germination (K), 3 days after germination (L), and 20 days after germination (M). Bars = 5 mm in K and L, 5 cm in M. N, 1000-grain weight, n = 3. O, Grain length, width and thickness, n = 20. All values are means ± SD. *indicates significant difference at P<0.05 level(Student’s t-test).
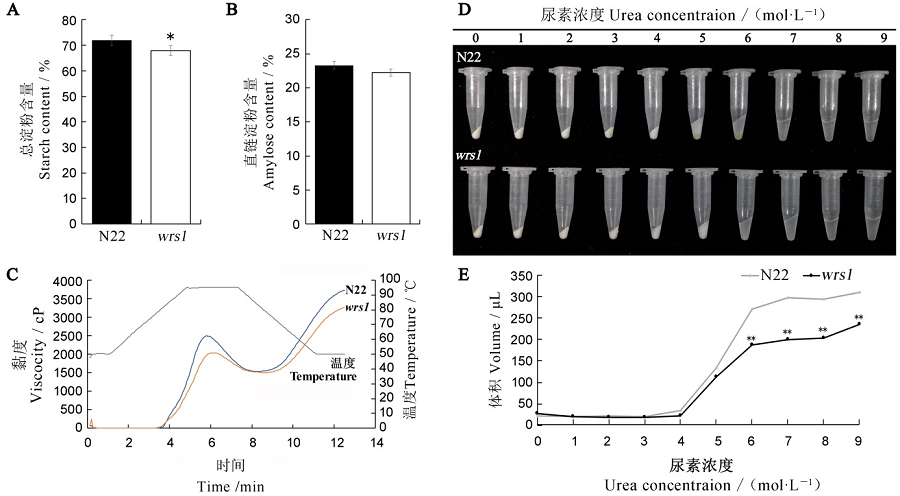
Fig. 2. Content and physicochemical characteristics of starches in mature seeds of wild-type and the wrs1 mutant. A, B, Contents of total starch (A) and amylose (B) in wild-type and the wrs1 mutant. n = 3. C, Analysis of RVA characteristic values of starch in wild-type and the wrs1 mutant. D, Swollen volume of the wild-type and wrs1 starch in urea solution. E, Comparation of the swollen volume in the wild-type and wrs1. n = 3. Values are means ± SD. * and **indicate significant difference at 0.05 and 0.01 level, respectively(Student’s t-test).
| 材料 Material | 峰值黏度 Peak viscosity / cP | 热浆黏度 Hot paste viscosity / cP | 崩解值 Breakdown viscosity / cP | 冷胶黏度 Cool paste viscosity / cP | 消减值 Setback viscosity / cP | 糊化温度 Pasting temperature /℃ | 峰值时间 Peak time / min |
|---|---|---|---|---|---|---|---|
| N22 | 2494 | 1536 | 958 | 3727 | 1233 | 81.40 | 5.80 |
| wrs1 | 2041 | 1506 | 535 | 3262 | 1221 | 80.65 | 6.13 |
Table 2 Analysis of RVA characteristic values of starch in wild-type and the wrs1 mutant.
| 材料 Material | 峰值黏度 Peak viscosity / cP | 热浆黏度 Hot paste viscosity / cP | 崩解值 Breakdown viscosity / cP | 冷胶黏度 Cool paste viscosity / cP | 消减值 Setback viscosity / cP | 糊化温度 Pasting temperature /℃ | 峰值时间 Peak time / min |
|---|---|---|---|---|---|---|---|
| N22 | 2494 | 1536 | 958 | 3727 | 1233 | 81.40 | 5.80 |
| wrs1 | 2041 | 1506 | 535 | 3262 | 1221 | 80.65 | 6.13 |
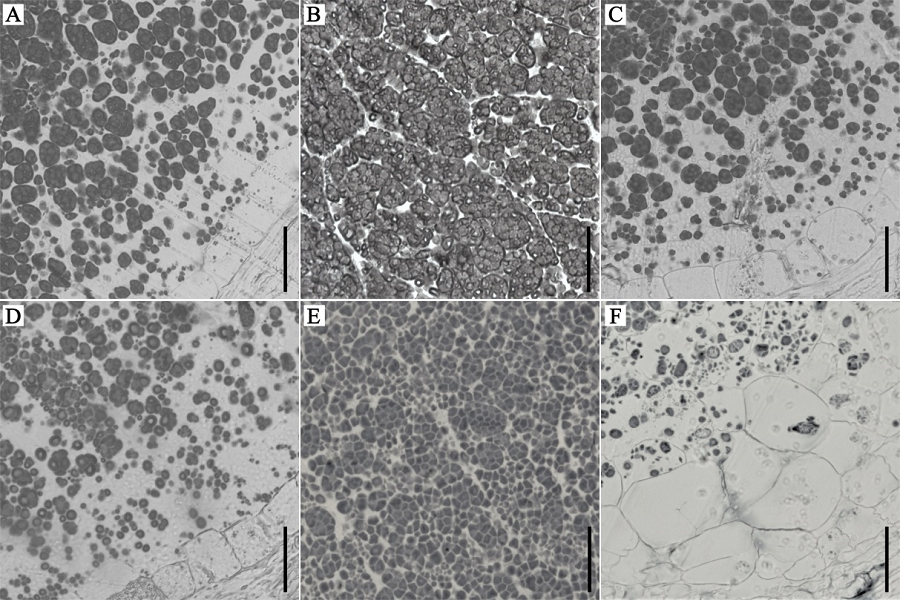
Fig. 3. Semi-thin section analysis of developing endosperm of wild-type and the wrs1 mutant. A, D—Semi-thin section analysis of the compound starch granules of back area of wild-type (A) and wrs1 (D) mutant endosperm at 8 DAF. B, E—Semi-thin section analysis of compound starch granules of central part of wild-type (B) and wrs1 mutant (E) endosperm at 8 DAF. C, F—Semi-thin section analysis of compound starch granules of the ventral side of wild-type (C) and wrs1 mutant (F) endosperm at 8 DAF. Bars = 50 μm.
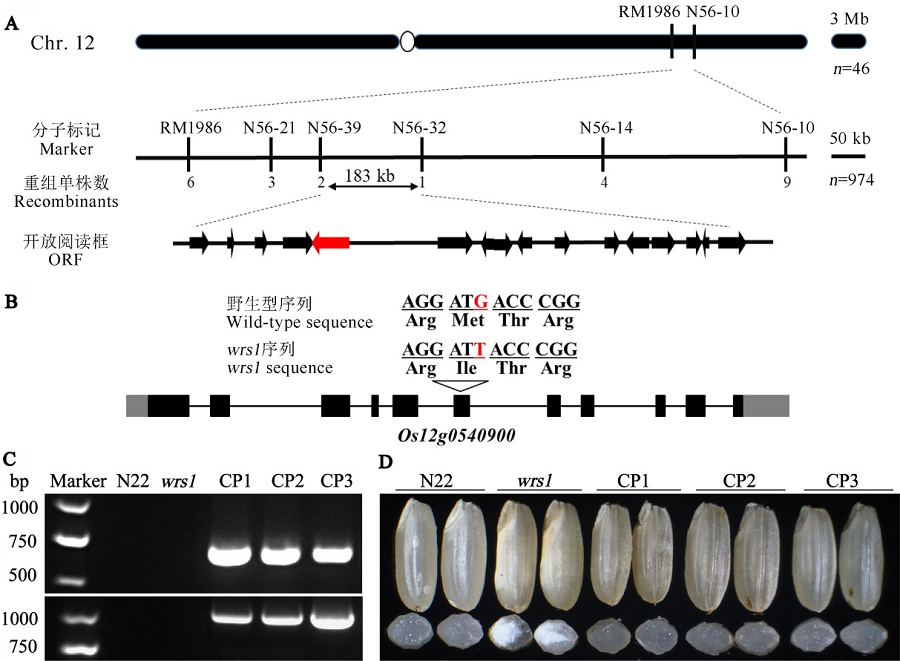
Fig. 4. Map-based cloning of WRS1. A, Fine mapping of the WRS1 locus. The WRS1 locus was mapped to a 183 kb region with markers N56-39 and N56-32 on the long arm of chromosome 12, which contains 16 predicted open reading frames (ORFs). B, The structure of Os12g0540900 and the mutation site. One nucleotide substitution (red) in the sequence is indicated, which results in a substitution of Met-256 to Ile. C, PCR analyses of wrs1 complemented lines (CP1, CP2 and CP3). The upper and lower panels in turn show the amplification of the transgenic positive identification primers N10 and Hyg. D, Mature seed appearance of wild-type, the wrs1 mutant and three positive transgenic lines (CP1, CP2 and CP3). The upside shows the mature seed phenotype of each positive transgenic line, and the downside shows cross-section of the seeds in corresponding transgenic line.
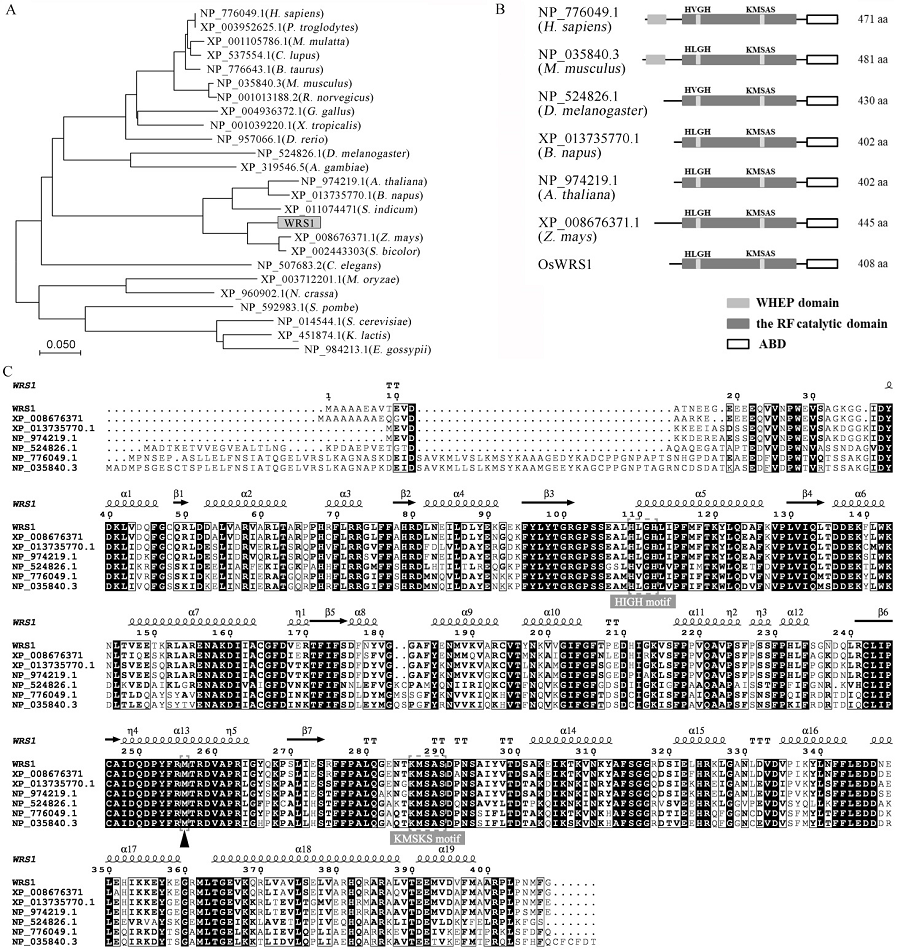
Fig. 5. Structure and phylogenetic analysis of WRS1 protein and its homologs. A, A neighbor-joining tree of WRS1 and its homologs. The tree was constructed using MEGA7 and bootstrapped with 1000 replicates. The proteins are named after their NCBI accession numbers. B, Domain analysis of WRS1 and its homologous proteins. The gray boxes represent the WHEP domain that is conserved in animals, the black boxes represent the Rossmann fold catalytic domain, the white boxes represent the anti-codon binding domain, aa represents the amino acid. C, Protein sequence alignment of WRS1 and its homologs. Two gray boxes indicate the catalytic site of Class I aaRSs; The arrowhead indicates the mutation site. The secondary structure is obtained from the PHYRE2 website by homology modeling (normal mode) with a known yeast tryptophanyl-tRNA synthetase crystal model c3kt3D as the pattern; the amino acid sequences are compared by the ESPript 3.0 website (default).
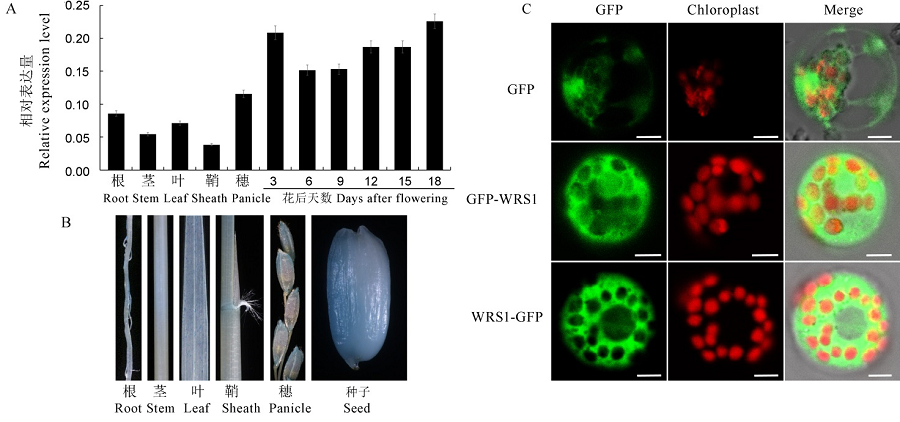
Fig. 6. Expression pattern and subcellular localization of WRS1. A, Relative expression levels of WRS1 in root, stem, leaf, sheath, panicle and developing seeds of wild-type. B, GUS staining patterns in root, stem, leaf, sheath, panicle and seed in transgenic plants. C, Fluorescence micrographs of the WRS1-GFP and GFP-WRS1 fusion constructs in rice protoplasts, with the empty pAN580-GFP served as a control. Bars, 5 μm.
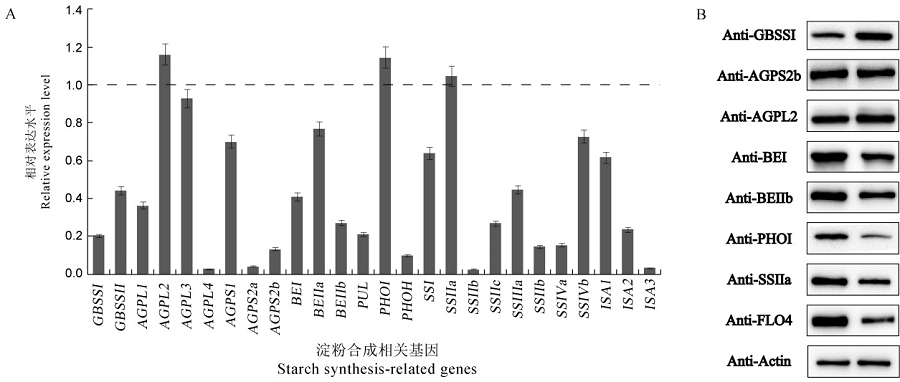
Fig. 7. Expression and immunoblotting analyses of starch synthesis-related genes. A, Expression analysis of starch synthesis-related genes. The relative expression levels of the genes in the wrs1 mutant were based on the corresponding genes in N22. Total RNA was extracted from seeds at 12 days after flowering. Ubiquitin was used as control. n = 3. All values are means ± SD. Student’s t-test, *P<0.05, **P<0.01. B, Western blotting analysis of starch synthesis related proteins in wild-type and mutant mature seeds. GBSSⅠ, Granule bound starch synthaseⅠ; AGPS2b, ADP-glucose pyrophosphorylase small subunit 2b; AGPL2, ADP-glucose pyrophosphorylase large subunit 2; BEⅠ, Starch-branching enzyme; BEⅡb, Starch-branching enzymeⅡb; PHOⅠ, Plastid phosphorylaseⅠ; SSⅡa, Starch synthase Ⅱa; FLO4(FLOURY ENDOSPERM4), cytosolic pyruvate orthophosphate dikinase B; Actin was used as control.

Fig. 8. Contents of protein and free amino acids in wrs1. A, Protein contents in wild-type and wrs1 grains. B, Free amino acid contents in the seeds of wild-type and the wrs1 mutant. C, Total free amino acids contents in the seeds of wild-type and the wrs1 mutant; D, Relative contents of free amino acids in mature seeds of wild-type and the wrs1 mutant. n = 3. All values are means ± SD. Student’s t-test, *P<0.05, **P<0.01.
| [1] | Fitzgerald M A, McCouch S R, Hall R D. Not just a grain of rice: The quest for quality[J]. Trends in Plant Science, 2009, 14(3): 133-139. |
| [2] | Crofts N, Nakamura Y, Fujita N.Critical and speculative review of the roles of multi-protein complexes in starch biosynthesis in cereals[J]. Plant Science, 2017, 262: 1-8. |
| [3] | Feike D, Seung D, Graf A, Bischof S, Ellick T, Coiro M, Soyk S, Eicke S, Mettler-Altmann T, Lu K J, Trick M, Zeeman S C, Smith A M.The starch granule-associated protein EARLY STARVATION1 is required for the control of starch degradation in Arabidopsis thaliana leaves[J]. Plant Cell, 2016, 28(6): 1472-1489. |
| [4] | Silver D M, Kotting O, Moorhead G B.Phosphoglucan phosphatase function sheds light on starch degradation[J]. Trends in Plant Science, 2014, 19(7): 471-478. |
| [5] | Hejazi M, Mahlow S, Fettke J.The glucan phosphorylation mediated by α-glucan, water dikinase (GWD) is also essential in the light phase for a functional transitory starch turn-over[J/OL].Plant Signaling & Behavior, 2014, 9(7): e28892. |
| [6] | Hwang S K, Nishi A, Satoh H, Okita T W.Rice endosperm-specific plastidial alpha-glucan phosphorylase is important for synthesis of short-chain malto- oligosaccharides[J]. Archives of Biochemistry and Biophysics, 2010, 495(1): 82-92. |
| [7] | Higgins J E, Kosar-Hashemi B, Li Z, Howitt C A, Larroque O, Flanagan B, Morell M K, Rahman S.Characterization of starch phosphorylases in barley grains[J]. Journal of the Science of Food and Agriculture, 2013, 93(9): 2137-2145. |
| [8] | Peng C, Wang Y, Liu F, Ren Y, Zhou K, Lv J, Zheng M, Zhao S, Zhang L, Wang C, Jiang L, Zhang X, Guo X, Bao Y, Wan J.FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm[J]. Plant Journal, 2014, 77(6): 917-930. |
| [9] | Seung D, Soyk S, Coiro M, Maier B A, Eicke S, Zeeman S C.PROTEIN TARGETING TO STARCH is required for localizing GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis[J/OL].PLoS Biology, 2015, 13(2): e1002080. |
| [10] | Pfister B, Zeeman S C.Formation of starch in plant cells[J]. Cellular and Molecular Life Sciences, 2016, 73(14): 2781-2807. |
| [11] | Hunter B G, Beatty M K, Singletary G W, Hamaker B R, Dilkes B P, Larkins B A, Jung R.Maize opaque endosperm mutations create extensive changes in patterns of gene expression[J]. Plant Cell, 2002, 14(10): 2591-2612. |
| [12] | Wakasa Y, Yasuda H, Oono Y, Kawakatsu T, Hirose S, Takahashi H, Hayashi S, Yang L, Takaiwa F.Expression of ER quality control-related genes in response to changes in BiP1 levels in developing rice endosperm[J]. Plant Journal, 2011, 65(5): 675-689. |
| [13] | Han X, Wang Y, Liu X, Jiang L, Ren Y, Liu F, Peng C, Li J, Jin X, Wu F, Wang J, Guo X, Zhang X, Cheng Z, Wan J.The failure to express a protein disulphide isomerase-like protein results in a floury endosperm and an endoplasmic reticulum stress response in rice[J]. Journal of Experiment Botany, 2012, 63(1): 121-130. |
| [14] | Wang Y, Ren Y, Liu X, Jiang L, Chen L, Han X, Jin M, Liu S, Liu F, Lv J, Zhou K, Su N, Bao Y, Wan J.OsRab5a regulates endomembrane organization and storage protein trafficking in rice endosperm cells[J]. Plant Journal, 2010, 64(5): 812-824. |
| [15] | Liu F, Ren Y, Wang Y, Peng C, Zhou K, Lv J, Guo X, Zhang X, Zhong M, Zhao S, Jiang L, Wang H, Bao Y, Wan J.OsVPS9A functions cooperatively with OsRAB5A to regulate post-Golgi dense vesicle-mediated storage protein trafficking to the protein storage vacuole in rice endosperm cells[J]. Molecular Plant, 2013, 6(6): 1918-1932. |
| [16] | Ren Y, Wang Y, Liu F, Zhou K, Ding Y, Zhou F, Wang Y, Liu K, Gan L, Ma W, Han X, Zhang X, Guo X, Wu F, Cheng Z, Wang J, Lei C, Lin Q, Jiang L, Wu C, Bao Y, Wang H, Wan J.GLUTELIN PRECURSOR ACCUMULATION3 encodes a regulator of post-Golgi vesicular traffic essential for vacuolar protein sorting in rice endosperm[J]. Plant Cell, 2014, 26(1): 410-425. |
| [17] | Wang Y, Liu F, Ren Y, Wang Y, Liu X, Long W, Wang D, Zhu J, Zhu X, Jing R, Wu M, Hao Y, Jiang L, Wang C, Wang H, Bao Y, Wan J.GOLGI TRANSPORT 1B regulates protein export from the endoplasmic reticulum in rice endosperm cells[J]. Plant Cell, 2016, 28(11): 2850-2865. |
| [18] | Zhu J, Ren Y, Wang Y, Liu F, Teng X, Zhang Y, Duan E, Wu M, Zhong M, Hao Y, Zhu X, Lei J, Wang Y, Yu Y, Pan T, Bao Y, Wang Y, Wan J.OsNHX5-mediated pH homeostasis is required for post-Golgi trafficking of seed storage proteins in rice endosperm cells[J/OL].BMC Plant Biology, 2019, 19(1): 295. |
| [19] | Perona J J, Gruic-Sovulj I.Synthetic and Editing Mechanisms of Aminoacyl-tRNA Synthetases[M]//Kim S. Topics in Current Chemistry-Series, 2014: 1-41. |
| [20] | Guo M, Yang X L, Schimmel P.New functions of aminoacyl-tRNA synthetases beyond translation[J]. Nature Reviews Molecular Cell Biology, 2010, 11(9): 668-674. |
| [21] | Park S G, Ewalt K L, Kim S.Functional expansion of aminoacyl-tRNA synthetases and their interacting factors: New perspectives on housekeepers[J]. Trends in Biochemical Sciences, 2005, 30(10): 569-574. |
| [22] | Duchêne A, Giritch A, Hoffmann B, Cognat V, Lancelin D, Peeters N M, Zaepfel M, Maréchal-Drouard L, Small I D.Dual targeting is the rule for organellar aminoacyl-tRNA synthetases in Arabidopsis thaliana[J]. Proceedings of the National Academy of Sciences of the United States of America, 2005, 102(45): 16484-16489. |
| [23] | Steiner R E, Ibba M. Regulation of tRNA-dependent translational quality control[J/OL]. IUBMB Life, 2019, 69: DOI: 10.1002/iub.2080. |
| [24] | Jin M. Unique roles of tryptophanyl-tRNA synthetase in immune control and its therapeutic implications[J].Experimental & Molecular Medicine, 2019, 51(1): . |
| [25] | Yakobov N, Debard S, Fischer F, Senger B, Becker H D.Cytosolic aminoacyl-tRNA synthetases: Unanticipated relocations for unexpected functions[J]. Biochimica et Biophysica Acta, 2018, 1861(4): 387-400. |
| [26] | Ray P, Arif A, Fox P.Macromolecular complexes as depots for releasable regulatory proteins[J]. Trends in Biochemical Sciences, 2007, 32(4): 158-164. |
| [27] | Berg M, Rogers R, Muralla R, Meinke D.Requirement of aminoacyl-tRNA synthetases for gametogenesis and embryo development in Arabidopsis[J]. Plant Journal, 2005, 44(5): 866-878. |
| [28] | Yang X, Li G, Tian Y, Song Y, Liang W, Zhang D.A rice glutamyl-tRNA synthetase modulates early anther cell division and patterning[J]. Plant Physiology, 2018, 177(2): 728-744. |
| [29] | Kagi C, Baumann N, Nielsen N, Stierhof Y D, Gross-Hardt R.The gametic central cell of Arabidopsis determines the lifespan of adjacent accessory cells[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(51): 22 350-22 355. |
| [30] | Kim Y K, Lee J Y, Cho H S, Lee S S, Ha H J, Kim S, Choi D, Pai H S.Inactivation of organellar glutamyl- and seryl-tRNA synthetases leads to developmental arrest of chloroplasts and mitochondria in higher plants[J]. Journal of Biological Chemistry, 2005, 280(44): 37098-37106. |
| [31] | Wang Y, Wang C, Zheng M, Lyu J, Xu Y, Li X, Niu M, Long W, Wang D, Wang H, Terzaghi W, Wang Y, Wan J.WHITE PANICLE1, a val-tRNA synthetase regulating chloroplast ribosome biogenesis in rice, is essential for early chloroplast development[J]. Plant Physiology, 2016, 170(4): 2110-2123. |
| [32] | Zhang Y, Hao Y, Wang Y, Wang C, Wang Y, Long W, Wang D, Liu X, Jiang L, Wan J.Lethal albinic seedling, encoding a threonyl-tRNA synthetase, is involved in development of plastid protein synthesis system in rice[J]. Plant Cell Reports, 2017, 36(7): 1053-1064. |
| [33] | Kitagawa M, Balkunde R, Bui H, Jackson D.An aminoacyl tRNA Synthetase, OKI1, is required for proper shoot meristem size in Arabidopsis[J]. Plant Cell Physiology, 2019, 60(11): 2597-2608. |
| [34] | Zuo D, Yi S, Liu R, Qu B, Huang T, He W, Li C, Li H, Liao Y.A deoxynivalenol-activated methionyl-tRNA synthetase gene from wheat encodes a nuclear localized protein and protects plants against fusarium pathogens and mycotoxins[J]. Phytopathology, 2016, 106(6): 614-623. |
| [35] | Luna E, van Hulten M, Zhang Y, Berkowitz O, López A, Pétriacq P, Sellwood M A, Chen B, Burrell M, van de Meene A, Pieterse C M J, Flors V, Ton J. Plant perception of β-aminobutyric acid is mediated by an aspartyl-tRNA synthetase[J]. Nature Chemical Biology, 2014, 10(6): 450-456. |
| [36] | Long W, Wang Y, Zhu S, Jing W, Wang Y, Ren Y, Tian Y, Liu S, Liu X, Chen L, Wang D, Zhong M, Zhang Y, Hu T, Zhu J, Hao Y, Zhu X, Zhang W, Wang C, Zhang W, Wan J.FLOURY SHRUNKEN ENDOSPERM1 connects phospholipid metabolism and amyloplast development in rice[J]. Plant Physiology, 2018, 177(2): 698-712. |
| [37] | Robert X, Gouet P.Deciphering key features in protein structures with the new ENDscript server[J]. Nucleic Acids Research, 2014, 42: W320-W324. |
| [38] | Ries F, Herkt C, Willmund F.Co-translational protein folding and sorting in chloroplasts[J]. Plants (Basel), 2020, 9(2): 214. doi: 10.3390/plants9020214. |
| [39] | Zheng M, Wang Y, Liu X, Sun J, Wang Y, Xu Y, Lv J, Long W, Zhu X, Guo X, Jiang L, Wang C, Wan J.The RICE MINUTE-LIKE1 (RML1) gene, encoding a ribosomal large subunit protein L3B, regulates leaf morphology and plant architecture in rice[J]. Journal of Experimental Botany, 2016, 67(11): 3457-3469. |
| [40] | Wang S, Lei C, Wang J, Ma J, Tang S, Wang C, Zhao K, Tian P, Zhang H, Qi C, Cheng Z, Zhang X, Guo X, Liu L, Wu C, Wan J.SPL33, encoding an eEF1A-like protein, negatively regulates cell death and defense responses in rice[J]. Journal of Experimental Botany, 2017, 68(5): 899-913. |
| [41] | Jia J, Arif A, Ray P S, Fox P L.WHEP domains direct noncanonical function of glutamyl-prolyl tRNA synthetase in translational control of gene expression[J]. Molecular Cell, 2008, 29(6): 679-690. |
| [42] | Sajish M, Zhou Q, Kishi S, Valdez D J, Kapoor M, Guo M, Lee S, Kim S, Yang X L, Schimmel P.Trp-tRNA synthetase bridges DNA-PKcs to PARP-1 to link IFN-gamma and p53 signaling[J]. Nature Chemical Biology, 2012, 8(6): 547-554. |
| [1] | DU Yimo, PAN Tian, TIAN Yunlu, LIU Shijia, LIU Xi, JIANG Ling, ZHANG Wenwei, WANG Yihua*,WAN Jianmin . Phenotypic Analysis and Gene Cloning of Rice Floury Endosperm Mutant fse4 [J]. Chinese Journal of Rice Science, 2019, 33(6): 499-512. |
| [2] | Peng-yi PAN, Jian-ping ZHU, Yun-long WANG, Yuan-yuan HAO, Yue CAI, Wen-wei ZHANG, Ling JIANG, Yi-hua WANG, Jian-min WAN. Phenotyping and Gene Cloning of a Floury Endosperm Mutant ws in Rice [J]. Chinese Journal OF Rice Science, 2016, 30(5): 447-457. |
| [3] | FANG Pengfei, LI Sanfeng, JIAO Guiai, XIE Lihong, HU Peisong, WEI Xiangjin*, TANG Shaoqing*. Physicochemical Property Analysis and Gene Mapping of a Floury Endosperm Mutant flo7 in Rice [J]. Chinese Journal of Rice Science, 2014, 28(5): 447-457. |
| [4] | WEI Cun-xu,ZHANG Jun,ZHOU Wei-dong,CHEN Yi-fang,LIU Qiao-quan*. Degradation of Amyloplast Envelope and Discussion on the Concept of Compound Starch Granule in Rice Endosperm [J]. Chinese Journal of Rice Science, 2008, 22(4): 377-384 . |
| [5] | XIONG Fei ,WANG Zhong ,GU Yun-jie ,CHEN Gang ,ZHOU Peng. Effects of Nitrogen Application Time on Caryopsis Development and Grain Quality of Rice Variety Yangdao 6 [J]. Chinese Journal of Rice Science, 2007, 21(6): 637-642 . |
| [6] | YANG Jian-chang*,CHANG Er-hua,TANG Cheng,ZHANG Hao,WANG Zhi-qin. Relationships of Ethylene Evolution Rate and 1-Aminocylopropane-1-Carboxylic Acid Concentration in Grains During Grain Filling with Appearance Quality of Rice [J]. Chinese Journal of Rice Science, 2007, 21(1): 77-83 . |
| [7] | XIONG Fei ,WANG Zhong ,CHEN Gang ,WANG Jue. Caryopsis Development and Main Grain Quality Characteristics in Different indica Rice Varieties [J]. Chinese Journal of Rice Science, 2006, 20(1): 43-46 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||