
Chinese Journal OF Rice Science ›› 2016, Vol. 30 ›› Issue (2): 111-120.DOI: 10.16819/j.1001-7216.2016.5138
• Orginal Article • Next Articles
Jiao MENG1, Hai-hua WANG1,2,3,*( ), Jian-hua XIANG1, Dan JIANG1, Xi-xu PENG1,3, Huan-huan HE1
), Jian-hua XIANG1, Dan JIANG1, Xi-xu PENG1,3, Huan-huan HE1
Received:2015-09-16
Revised:2015-11-30
Online:2016-03-10
Published:2016-03-10
Contact:
Hai-hua WANG
孟姣1, 王海华1,2,3,*( ), 向建华1, 蒋丹1, 彭喜旭1,3, 贺欢欢1
), 向建华1, 蒋丹1, 彭喜旭1,3, 贺欢欢1
通讯作者:
王海华
基金资助:CLC Number:
Jiao MENG, Hai-hua WANG, Jian-hua XIANG, Dan JIANG, Xi-xu PENG, Huan-huan HE. Expression Profiles of Rice WRKY Transcription Factor Gene Family Responsive to Exogenous Nitric Oxide Application[J]. Chinese Journal OF Rice Science, 2016, 30(2): 111-120.
孟姣, 王海华, 向建华, 蒋丹, 彭喜旭, 贺欢欢. 水稻WRKY转录因子基因家族响应外源一氧化氮的表达谱分析[J]. 中国水稻科学, 2016, 30(2): 111-120.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2016.5138
| 基因 Gene | 登录号 Accession No. | 正向引物 Forward primer (5'-3') | 反向引物 Reverse primer (5'-3') |
|---|---|---|---|
| OsWRKY15 | Os01g0656400 | GTCGTCGCCCTTCGTGTC | ATGAACTCCATGTCGTCTCCC |
| OsWRKY42 | Os02g0462800 | ATGGGAGCGTGTCCAACG | GCGGCCATCAGAAGCATAA |
| OsWRKY21 | Os01g0821600 | GCTGAGCAACAGAGCAGGTA | GCGTCAGTTATGCGTCAAG |
| OsWRKY77 | Os01g0584900 | CCTCGCTGCTGCTTCCTT | TTCAGGTCGCCTTGGTCC |
| OsWRKY71 | Os02g0181300 | AGGAATTGATGATTCGCTGTA | CGCCGATGATCGTTGGTT |
| OsWRKY24 | Os01g0826400 | TGCTCCTGACCTCCAGTATCTT | TTCCTCTGCTCGTCCTTGC |
| OsWRKY69 | Os08g0386200 | TGCTTCGACTTCGACCCG | TCTTCCCTCCTGCCACCA |
| Actin 1 | Os03g0718100 | ATCGCCCTGGACTATGAC | GAAACGCTCAGCACCAAT |
Table 1 Primers of the candidate genes for qRT-PCR.
| 基因 Gene | 登录号 Accession No. | 正向引物 Forward primer (5'-3') | 反向引物 Reverse primer (5'-3') |
|---|---|---|---|
| OsWRKY15 | Os01g0656400 | GTCGTCGCCCTTCGTGTC | ATGAACTCCATGTCGTCTCCC |
| OsWRKY42 | Os02g0462800 | ATGGGAGCGTGTCCAACG | GCGGCCATCAGAAGCATAA |
| OsWRKY21 | Os01g0821600 | GCTGAGCAACAGAGCAGGTA | GCGTCAGTTATGCGTCAAG |
| OsWRKY77 | Os01g0584900 | CCTCGCTGCTGCTTCCTT | TTCAGGTCGCCTTGGTCC |
| OsWRKY71 | Os02g0181300 | AGGAATTGATGATTCGCTGTA | CGCCGATGATCGTTGGTT |
| OsWRKY24 | Os01g0826400 | TGCTCCTGACCTCCAGTATCTT | TTCCTCTGCTCGTCCTTGC |
| OsWRKY69 | Os08g0386200 | TGCTTCGACTTCGACCCG | TCTTCCCTCCTGCCACCA |
| Actin 1 | Os03g0718100 | ATCGCCCTGGACTATGAC | GAAACGCTCAGCACCAAT |
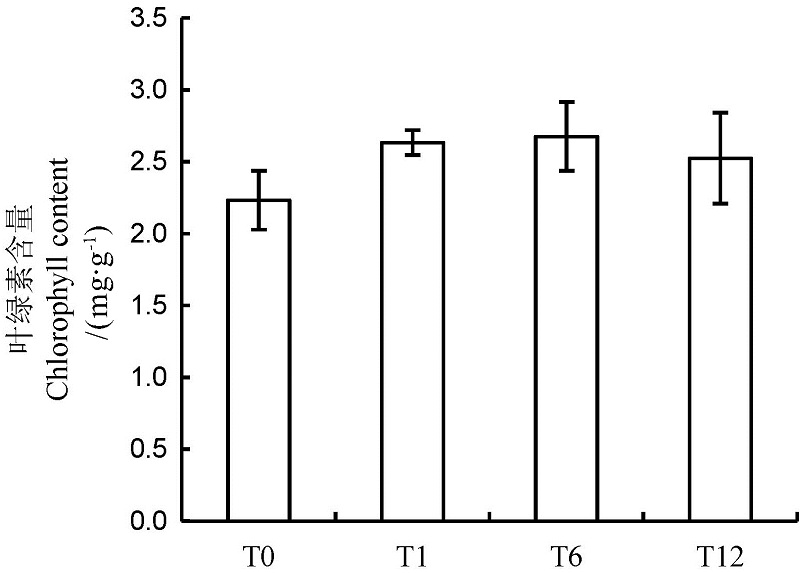
Fig. 1. Chlorophyll contents in rice leaves after NO treatment for different time. NO treatments for 0,1,6 and 12 h are indicated as T0, T1, T6 and T12, respectively. The same as below.
| 基因 Gene | 探针 Probe | 表达倍数 Expression fold change | 组别 Group | ||
|---|---|---|---|---|---|
| T1/T0 | T6/T0 | T12/T0 | |||
| OsWRKY1 | Os01g0246700|mRNA|AY676928|CDS | 1.87 | 2.06 | 1.35 | Ⅱb |
| Os01g0246700|mRNA|AK105509|CDS+3'UTR | 2.72 | 1.54 | 2.19 | ||
| OsWRKY3 | Os03g0758000|mRNA|AY341859|CDS+3'UTR | 0.33 | 0.38 | 3.70 | Ⅰb |
| OsWRKY6 | Os03g0798500|COMBINER_EST|Os03g0798500|8 | 0.59 | 0.57 | 2.25 | Ⅱc |
| OsWRKY8 | Os05g0583000|mRNA|AK109568|CDS+3'UTR | 2.95 | 1.13 | 0.44 | Ⅱd |
| OsWRKY11 | Os01g0626400|mRNA|AY341856|CDS+3'UTR | 1.91 | 2.88 | 1.60 | Ⅰb |
| Os01g0626400|mRNA|AK108745|CDS+3'UTR | 2.23 | 2.57 | 1.63 | ||
| OsWRKY12 | Os01g0624700|mRNA|AK111416|CDS+3'UTR | 3.14 | 2.74 | 1.99 | Ⅱd |
| OsWRKY13 | Os01g0750100|mRNA|AK060018|CDS+3'UTR | 1.55 | 0.67 | 0.48 | Ⅱd |
| OsWRKY15 | Os01g0656400|mRNA|AY676926|CDS | 4.98 | 1.16 | 2.70 | Ⅲa |
| OsWRKY17 | Os01g0972800|mRNA|AY341842|CDS+3'UTR | 0.09 | 0.09 | 0.32 | Ⅰb |
| OsWRKY21 | Os01g0821600|mRNA|AK108657|CDS+3'UTR | 4.69 | 4.42 | 1.31 | Ⅲb |
| OsWRKY23 | Os01g0734000|mRNA|AK108909|CDS+3'UTR | 2.24 | 1.69 | 2.11 | Ⅰb |
| OsWRKY24 | Os01g0826400|mRNA|AK107199|CDS+3'UTR | 6.48 | 0.83 | 3.86 | Ⅰa |
| Os01g0826400|mRNA|AY341849|CDS+3'UTR | 5.39 | 0.63 | 3.33 | ||
| Os01g0826400|mRNA|AY676925|CDS | 5.92 | 0.75 | 3.65 | ||
| OsWRKY28 | Os06g0649000|mRNA|AK106282|CDS+3'UTR | 7.87 | 1.32 | 2.28 | Ⅱa |
| Os06g0649000|mRNA|AK119644|CDS+3'UTR | 6.77 | 1.05 | 1.91 | ||
| OsWRKY29 | Os07g0111400|mRNA|AY341858|CDS+3'UTR | 0.73 | 1.32 | 2.70 | Ⅰb |
| OsWRKY36 | Os04g0545000|mRNA|AK073695|CDS+3'UTR | 0.47 | 0.73 | 0.61 | Ⅰb |
| OsWRKY39 | Os02g0265200|mRNA|AK066775|CDS+3'UTR | 1.43 | 2.00 | 2.62 | Ⅱd |
| OsWRKY42 | Os02g0265200|mRNA|AK119593|CDS+3'UTR | 1.42 | 2.27 | 2.34 | Ⅱc |
| Os02g0462800|mRNA|AK110587|CDS+3'UTR | 5.51 | 3.80 | 2.01 | ||
| Os02g0462800|mRNA|AY676923|CDS | 5.01 | 3.14 | 2.05 | ||
| OsWRKY50 | Os11g0117600|COMBINER_EST|Os11g0117600|8 | 0.65 | 7.31 | 1.11 | Ⅲb |
| OsWRKY53 | Os05g0343400|mRNA|AY676929|CDS | 2.10 | 0.89 | 1.02 | Ⅰa |
| OsWRKY56 | Os12g0116600|mRNA|AK102093|CDS+3'UTR | 1.31 | 0.42 | 1.03 | Ⅲb |
| OsWRKY60 | Os03g0657400|COMBINER_EST|AU057193|6 | 1.01 | 0.58 | 0.44 | Ⅰb |
| OsWRKY62 | Os09g0417800|mRNA|AK067834|CDS+3'UTR | 1.48 | 1.65 | 2.03 | Ⅱa |
| OsWRKY67 | Os05g0183100|mRNA|AK066252|CDS+3'UTR | 1.13 | 0.66 | 0.28 | Ⅰb |
| OsWRKY69 | Os08g0386200|mRNA|AK111606|CDS+3'UTR | 3.05 | 0.98 | 2.77 | Ⅲa |
| OsWRKY71 | Os08g0386200|mRNA|AY676931|CDS | 3.33 | 0.88 | 2.80 | Ⅱa |
| Os02g0181300|mRNA|AK058773|CDS+3'UTR | 3.79 | 1.72 | 1.67 | ||
| Os02g0181300|mRNA|AY676927|CDS | 3.41 | 1.45 | 1.64 | ||
| OsWRKY72 | Os11g0490900|mRNA|AK108860|CDS+3'UTR | 2.32 | 0.89 | 1.75 | Ⅰb |
| OsWRKY77 | Os01g0584900|mRNA|AK108522|CDS+3'UTR | 1.10 | 1.80 | 2.31 | Ⅰb |
| Os01g0584900|mRNA|AY341846|CDS+3'UTR | 1.24 | 1.88 | 2.48 | ||
| OsWRKY79 | Os03g0335200|mRNA|AK105244|CDS+3'UTR | 0.66 | 1.83 | 2.53 | Ⅲb |
| OsWRKY88 | Os07g0596900|mRNA|AY341847|CDS+3'UTR | 0.94 | 1.53 | 2.13 | Ⅲa |
| OsWRKY104 | Os11g0117400|COMBINER_EST|CI444468|6 | 1.40 | 0.32 | 0.93 | Ⅲb |
| OsWRKY109 | Os05g0129800|COMBINER_EST|Os05g0129800|8 | 0.73 | 0.82 | 2.02 | Ⅱd |
| OsWRKY113 | Os06g0158100|mRNA|AY676932|CDS | 2.35 | 0.96 | 3.71 | Ⅲa |
Table 2 Microarray analysis of differentially expressed WRKY genes in rice under NO treatment.
| 基因 Gene | 探针 Probe | 表达倍数 Expression fold change | 组别 Group | ||
|---|---|---|---|---|---|
| T1/T0 | T6/T0 | T12/T0 | |||
| OsWRKY1 | Os01g0246700|mRNA|AY676928|CDS | 1.87 | 2.06 | 1.35 | Ⅱb |
| Os01g0246700|mRNA|AK105509|CDS+3'UTR | 2.72 | 1.54 | 2.19 | ||
| OsWRKY3 | Os03g0758000|mRNA|AY341859|CDS+3'UTR | 0.33 | 0.38 | 3.70 | Ⅰb |
| OsWRKY6 | Os03g0798500|COMBINER_EST|Os03g0798500|8 | 0.59 | 0.57 | 2.25 | Ⅱc |
| OsWRKY8 | Os05g0583000|mRNA|AK109568|CDS+3'UTR | 2.95 | 1.13 | 0.44 | Ⅱd |
| OsWRKY11 | Os01g0626400|mRNA|AY341856|CDS+3'UTR | 1.91 | 2.88 | 1.60 | Ⅰb |
| Os01g0626400|mRNA|AK108745|CDS+3'UTR | 2.23 | 2.57 | 1.63 | ||
| OsWRKY12 | Os01g0624700|mRNA|AK111416|CDS+3'UTR | 3.14 | 2.74 | 1.99 | Ⅱd |
| OsWRKY13 | Os01g0750100|mRNA|AK060018|CDS+3'UTR | 1.55 | 0.67 | 0.48 | Ⅱd |
| OsWRKY15 | Os01g0656400|mRNA|AY676926|CDS | 4.98 | 1.16 | 2.70 | Ⅲa |
| OsWRKY17 | Os01g0972800|mRNA|AY341842|CDS+3'UTR | 0.09 | 0.09 | 0.32 | Ⅰb |
| OsWRKY21 | Os01g0821600|mRNA|AK108657|CDS+3'UTR | 4.69 | 4.42 | 1.31 | Ⅲb |
| OsWRKY23 | Os01g0734000|mRNA|AK108909|CDS+3'UTR | 2.24 | 1.69 | 2.11 | Ⅰb |
| OsWRKY24 | Os01g0826400|mRNA|AK107199|CDS+3'UTR | 6.48 | 0.83 | 3.86 | Ⅰa |
| Os01g0826400|mRNA|AY341849|CDS+3'UTR | 5.39 | 0.63 | 3.33 | ||
| Os01g0826400|mRNA|AY676925|CDS | 5.92 | 0.75 | 3.65 | ||
| OsWRKY28 | Os06g0649000|mRNA|AK106282|CDS+3'UTR | 7.87 | 1.32 | 2.28 | Ⅱa |
| Os06g0649000|mRNA|AK119644|CDS+3'UTR | 6.77 | 1.05 | 1.91 | ||
| OsWRKY29 | Os07g0111400|mRNA|AY341858|CDS+3'UTR | 0.73 | 1.32 | 2.70 | Ⅰb |
| OsWRKY36 | Os04g0545000|mRNA|AK073695|CDS+3'UTR | 0.47 | 0.73 | 0.61 | Ⅰb |
| OsWRKY39 | Os02g0265200|mRNA|AK066775|CDS+3'UTR | 1.43 | 2.00 | 2.62 | Ⅱd |
| OsWRKY42 | Os02g0265200|mRNA|AK119593|CDS+3'UTR | 1.42 | 2.27 | 2.34 | Ⅱc |
| Os02g0462800|mRNA|AK110587|CDS+3'UTR | 5.51 | 3.80 | 2.01 | ||
| Os02g0462800|mRNA|AY676923|CDS | 5.01 | 3.14 | 2.05 | ||
| OsWRKY50 | Os11g0117600|COMBINER_EST|Os11g0117600|8 | 0.65 | 7.31 | 1.11 | Ⅲb |
| OsWRKY53 | Os05g0343400|mRNA|AY676929|CDS | 2.10 | 0.89 | 1.02 | Ⅰa |
| OsWRKY56 | Os12g0116600|mRNA|AK102093|CDS+3'UTR | 1.31 | 0.42 | 1.03 | Ⅲb |
| OsWRKY60 | Os03g0657400|COMBINER_EST|AU057193|6 | 1.01 | 0.58 | 0.44 | Ⅰb |
| OsWRKY62 | Os09g0417800|mRNA|AK067834|CDS+3'UTR | 1.48 | 1.65 | 2.03 | Ⅱa |
| OsWRKY67 | Os05g0183100|mRNA|AK066252|CDS+3'UTR | 1.13 | 0.66 | 0.28 | Ⅰb |
| OsWRKY69 | Os08g0386200|mRNA|AK111606|CDS+3'UTR | 3.05 | 0.98 | 2.77 | Ⅲa |
| OsWRKY71 | Os08g0386200|mRNA|AY676931|CDS | 3.33 | 0.88 | 2.80 | Ⅱa |
| Os02g0181300|mRNA|AK058773|CDS+3'UTR | 3.79 | 1.72 | 1.67 | ||
| Os02g0181300|mRNA|AY676927|CDS | 3.41 | 1.45 | 1.64 | ||
| OsWRKY72 | Os11g0490900|mRNA|AK108860|CDS+3'UTR | 2.32 | 0.89 | 1.75 | Ⅰb |
| OsWRKY77 | Os01g0584900|mRNA|AK108522|CDS+3'UTR | 1.10 | 1.80 | 2.31 | Ⅰb |
| Os01g0584900|mRNA|AY341846|CDS+3'UTR | 1.24 | 1.88 | 2.48 | ||
| OsWRKY79 | Os03g0335200|mRNA|AK105244|CDS+3'UTR | 0.66 | 1.83 | 2.53 | Ⅲb |
| OsWRKY88 | Os07g0596900|mRNA|AY341847|CDS+3'UTR | 0.94 | 1.53 | 2.13 | Ⅲa |
| OsWRKY104 | Os11g0117400|COMBINER_EST|CI444468|6 | 1.40 | 0.32 | 0.93 | Ⅲb |
| OsWRKY109 | Os05g0129800|COMBINER_EST|Os05g0129800|8 | 0.73 | 0.82 | 2.02 | Ⅱd |
| OsWRKY113 | Os06g0158100|mRNA|AY676932|CDS | 2.35 | 0.96 | 3.71 | Ⅲa |
| 组/亚组 Group/ subgroup | 差异表达基因数 Number of differentially expressed genes | 组/亚组的基因总数 Total number of genes in group/subgroup | 比例 Ratio /% |
|---|---|---|---|
| Ⅰ | 12 | 34 | 35.3 |
| Ⅱ | 11 | 30 | 36.7 |
| a | 3 | 4 | 75.0 |
| b | 1 | 8 | 12.5 |
| c | 2 | 7 | 28.6 |
| d | 5 | 11 | 45.6 |
| Ⅲ | 9 | 36 | 25.0 |
Table 3 Group distribution of differentially expressed WRKY genes of rice responsive to nitric oxide.
| 组/亚组 Group/ subgroup | 差异表达基因数 Number of differentially expressed genes | 组/亚组的基因总数 Total number of genes in group/subgroup | 比例 Ratio /% |
|---|---|---|---|
| Ⅰ | 12 | 34 | 35.3 |
| Ⅱ | 11 | 30 | 36.7 |
| a | 3 | 4 | 75.0 |
| b | 1 | 8 | 12.5 |
| c | 2 | 7 | 28.6 |
| d | 5 | 11 | 45.6 |
| Ⅲ | 9 | 36 | 25.0 |
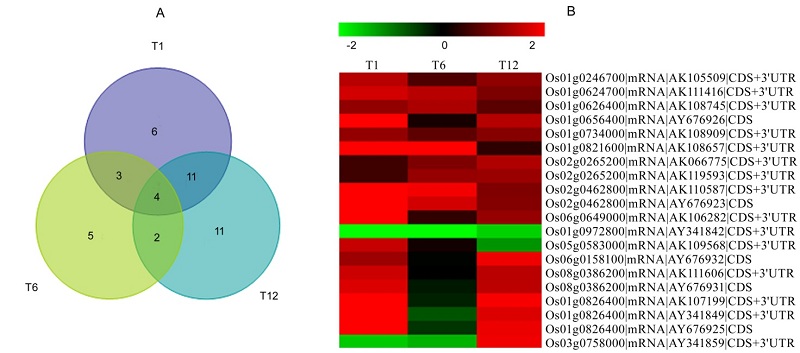
Fig.3. Venny analysis (A) and heatmap clustering (B) of NO-responsive rice WRKY gene. The color scale is shown on the top. Red, Up-regulated; Green, Down-regulated.
| [1] | Riaño-Pachón D M, Ruzicic S, Dreyer I, et al. PlnTFDB: An integrative plant transcription factor database.BMC Bioinform, 2007, 8: 42. |
| [2] | Eulgem T, Rushton P J, Robatzek S, et al.The WRKY superfamily of plant transcription factors.Trends Plant Sci, 2000, 5: 199-206. |
| [3] | Pandey S P, Somssich I E.The role of WRKY transcription factors in plant immunity.Plant Physiol, 2009, 150: 1648-1655. |
| [4] | Agarwal P, Reddy M P, Chikara J.WRKY: Its structure, evolutionary relationship, DNA-binding selectivity, role in stress tolerance and development of plants.Mol Biol Rep, 2011, 38: 3883-3896. |
| [5] | Chen L, Song Y, Li S, et al.The role of WRKY transcription factors in plant abiotic stresses.Biochim Biophys Acta, 2012, 1819: 120-128. |
| [6] | Li J, Brader G, Palva E T.The WRKY70 transcription factor: A node of convergence for jasmonate-mediated and salicylate-mediated signals in plant defense. Plant Cell, 2004, 16: 319-331. |
| [7] | Qiu D, Xiao J, Ding X, et al.OsWRKY13 mediates rice disease resistance by regulating defense-related genes in salicylate- and jasmonate-dependent signaling.Mol Plant Microbe Interact, 2007, 20: 492-499. |
| [8] | Wang H, Meng J, Peng X, et al.Rice WRKY4 acts as a transcriptional activator mediating defense responses toward Rhizoctonia solani, the causing agent of rice sheath blight.Plant Mol Biol, 2005, 89(1/2): 157-171. |
| [9] | Shang Y, Yan L, Liu Z Q, et al.The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition.Plant Cell, 2010, 22, 1909-1935. |
| [10] | Ryu H S, Han M, Lee S K, et al.A comprehensive expression analysis of the WRKY gene superfamily in rice plants during defense response.Plant Cell Rep, 2006, 25: 836-847. |
| [11] | Ramamoorthy R, Jiang S Y, Kumar N, et al.A comprehensive transcriptional profiling of the WRKY gene family in rice under various abiotic and phytohormone treatments.Plant Cell Physiol, 2008, 49: 865-879. |
| [12] | Lamotte O, Courtois C, Pugin L B A, et al. Nitric oxide in plants: The biosynthesis and cell signalling properties of a fascinating molecule.Planta, 2005, 221: 1-4. |
| [13] | Gould K, Lamotte O, Klinguer A, et al.Nitric oxide production in tobacco leaf cells: A generalized stress response?Plant Cell Environ, 2003, 26: 1851-1862. |
| [14] | Delledonne M.NO news is good news for plants.Curr Opin Plant Biol, 2005, 8: 390-396. |
| [15] | Palmieri M C, Sell S, Huang X, et al.Nitric oxide-responsive genes and promoters in Arabidopsis thaliana: A bioinformatics approach.J Exp Bot, 2008, 59:177-186. |
| [16] | Mukhtar M S, Deslandes L, Auriac M-C, et al.The Arabidopsis transcription factor WRKY27 influences wilt disease symptom development caused by Ralstonia solanacearum.Plant J, 2008, 56: 935-947. |
| [17] | Arnon D I.Copper enzymes in isolated chloroplasts: Polyphenoloxidase in Beta vulgaris. Plant Physiol, 1949, 24:1-15. |
| [18] | Rice WRKY Working Group. Nomenclature report on rice WRKY's-Conflict regarding gene names and its solution.Rice, 2012, 5: 3. |
| [19] | Wu K L, Guo Z J, Wang H H, et al.The WRKY family of transcription factors in rice and Arabidopsis and their origins.DNA Res, 2005, 12: 9-26. |
| [20] | Dong J, Chen C, Chen Z.Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response.Plant Mol Biol, 2003, 51: 21-37. |
| [21] | Kalde M, Barth M, Somssich I E, et al.Members of the Arabidopsis WRKY group Ⅲ transcription factors are part of different plant defense signaling pathways.Mol Plant Microbe Interact, 2003, 16: 295-305. |
| [22] | Chujo T, Miyamoto K, Shimogawa T, et al.OsWRKY28, a PAMP-responsive transrepressor, negatively regulates innate immune responses in rice against rice blast fungus.Plant Mol Biol, 2013, 82: 23-37. |
| [23] | Peng Y, Bartley L E, Chen X, et al.OsWRKY62 is a negative regulator of basal and Xa21-mediated defense against Xanthomonas oryzae pv. oryzae in rice.Mol Plant, 2008, 1: 446-458. |
| [24] | Yokotani N, Sato Y, Tanabe S, et al.WRKY76 is a rice transcriptional repressor playing opposite roles in blast disease resistance and cold stress tolerance.J Exp Bot, 2013, 64(16): 5085-5097. |
| [25] | Liu X, Bai X, Wang X, et al.OsWRKY71, a rice transcription factor, is involved in rice defense response.J Plant Physiol, 2007, 164: 969-979. |
| [26] | Park C Y, Lee J H, Yoo J H, et al.WRKY group IId transcription factors interact with calmodulin.FEBS Lett, 2005, 579: 1545-1550. |
| [27] | Durner J, Wendehenne D, Klessig D F.Defense gene induction in tobacco by nitric oxide, cyclic GMP, and cyclic ADP-ribose.Proc Natl Acad Sci USA, 1998, 95: 10328-10333. |
| [28] | Liu X Q, Bai X Q, Qian Q, et al.OsWRKY03, a rice transcriptional activator that functions in defense signaling pathway upstream of OsNPR1.Cell Res, 2005, 15(8): 593-603. |
| [29] | Jing S, Zhou X, Song Y, et al.Heterologous expression of OsWRKY23 gene enhances pathogen defense and dark-induced leaf senescence in Arabidopsis. Plant Growth Regul, 2009, 58:181-190. |
| [30] | Song Y, Jing S, Yu D.Overexpression of the stress-induced OsWRKY08 improves osmotic stress tolerance inArabidopsis. Chin Sci Bull, 2009, 54: 4671-4678. |
| [31] | Cheng H, Liu H, Deng Y, et al.The WRKY45-2 WRKY13 WRKY42 transcriptional regulatory cascade is required for rice resistance to fungal pathogen.Plant Physiol, 2015, 167(3): 1087-1099. |
| [32] | Zhang Z L, Shin M, Zou X, et al.A negative regulator encoded by a rice WRKY gene represses both abscisic acid and gibberellins signaling in aleurone cells.Plant Mol Biol, 2009, 70: 139-151. |
| [33] | Sperotto R A, Boff T, Duarte G L, et al.Increased senescence-associated gene expression and lipid peroxidation induced by iron deficiency in rice roots.Plant Cell Rep, 2008, 27: 183-195. |
| [34] | 黄盼盼. 18个水稻OsWRKY基因的克隆和OsWRKY21功能分析. 广州: 中山大学, 2010. |
| Huang P P.Cloning 18 OsWRKY genes and functional analysis of OsWRKY21. GuangZhou:Sun Yat-sen University, 2010. (in Chinese with English abstract) |
| [1] | ZHU Chunquan, WEI Qianqian, DANG Caixia, HUANG Jing, XU Qingshan, PAN Lin, ZHU Lianfeng, CAO Xiaochuang, KONG Yali, XIANG Xingjia, LIU Jia, JIN Qianyu, ZHANG Junhua. Salicylic Acid Alleviates Low Phosphorus Stress in Rice via a Nitric Oxide-dependent Manner [J]. Chinese Journal OF Rice Science, 2022, 36(5): 476-486. |
| [2] | YAO Shu, ZHANG Yadong, LU Kai, WANG Cailin. Progress in Functions, Allelic Variations and Interactions of Soluble Starch Synthases Genes SSⅡa and SSⅢa in Rice [J]. Chinese Journal OF Rice Science, 2022, 36(3): 227-236. |
| [3] | Yuxiang LI, Hairong LIN, Qian LIANG, Guodong WANG. Effects of Dopamine Priming on Seed Germination and Seedling Growth of Rice Under Salt Stress [J]. Chinese Journal OF Rice Science, 2021, 35(5): 487-494. |
| [4] | Shilin DING, Chaolei LIU, Qian QIAN, Zhenyu GAO. Research Advances on Molecular Genetic Mechanism for Cadmium Absorption and Transportation in Rice [J]. Chinese Journal OF Rice Science, 2019, 33(5): 383-390. |
| [5] | Kunneng ZHOU, Jiafa XIA, Tingchen MA, Yuanlei WANG, Zefu LI. Mapping and Mutation Analysis of Stripe Leaf and White Panicle Gene SLWP in Rice [J]. Chinese Journal OF Rice Science, 2018, 32(4): 325-334. |
| [6] | Xixu PENG, Ningning BAI, Haihua WANG. Isolation and Expression Profiles of Cadmium Stress-Responsive Rice WRKY15 Transcription Factor Gene [J]. Chinese Journal OF Rice Science, 2018, 32(2): 103-110. |
| [7] | Dao-ming WU, Hua-ping CAO, Xiao-li YU, Hong SHEN. Aluminium Tolerance of OsPIN2 Overexpressed Rice Seedlings under Pot Culture [J]. Chinese Journal OF Rice Science, 2015, 29(3): 250-258. |
| [8] | Yong-jie YANG, Xue-qin YANG, Cai-xia ZHANG, Guan-fu FU, Ting-ting CHEN, Long-xing TAO. Effects of Nitric Oxide on Drought Stress-induced Physiological Characteristics in Leaves of Nipponbare (Oryza sativa L) [J]. Chinese Journal OF Rice Science, 2015, 29(1): 65-72. |
| [9] | LIU Kai-li ,HAN Hang-ru ,XU Ying-jie ,LING Teng-Fang ,LIU Zhi-bing ,SUN Yong-gang ,HUA Rong ,SHEN Wen-biao. Exogenous Nitric Oxide Alleviates Salt Stress-induced Membrane Lipid Peroxidation in Rice Seedling Roots [J]. Chinese Journal of Rice Science, 2005, 19(4): 333-337 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||