
中国水稻科学 ›› 2018, Vol. 32 ›› Issue (5): 427-436.DOI: 10.16819/j.1001-7216.2018.7037
张占田, 孙雅菲, 艾昊, 罗闻真, 冯冰, 孙文献, 徐国华, 孙淑斌*( )
)
收稿日期:2017-03-30
修回日期:2017-08-18
出版日期:2018-09-10
发布日期:2018-09-10
通讯作者:
孙淑斌
基金资助:
Zhantian ZHANG, Yafei SUN, Hao AI, Wenzhen LUO, Bing FENG, Wenxian SUN, Guohua XU, Shubin SUN*( )
)
Received:2017-03-30
Revised:2017-08-18
Online:2018-09-10
Published:2018-09-10
Contact:
Shubin SUN
摘要:
【目的】水稻OsSHR2(LOC_Os03g31880)基因为拟南芥AtSHR的同源基因,与OsSHR1、OsSCR1和OsSCR2 同属于水稻GRAS转录因子家族。已有研究报道,转录因子基因SHR和SCR共同调控植物根系、叶片的发育,并参与各项生命活动。本研究旨在阐明OsSHR2在水稻中的时空表达特征及其在营养生长中的调控作用。【方法】通过生物信息学分析、表达模式分析、萌发动力学分析和水培实验验证该基因的功能。【结果】生物信息学分析发现OsSHR2、OsSHR1、OsSCR1和OsSCR2与拟南芥和其他物种的SHR亚家族和SCR亚家族成员具有很高的序列一致性;表达模式和pOsSHR2::GUS材料染色分析发现,OsSHR2在整个生长发育过程中的根系、叶片、维管组织和生殖器官中表达强烈,并集中在根尖的中柱、侧根原基和叶片及茎维管组织的中心表达,在野生型的地上部和根系中,OsSHR2受缺磷影响下调表达;对获得的OsSHR2的CRISPR-Cas9突变体osshr2进行种子萌发实验和水培实验,发现与野生型相比,osshr2的萌发时间延后,萌发率降低,在正常供磷和缺磷处理下,osshr2的地上部和根系长度显著小于野生型。【结论】OsSHR2在地上部和根系的发育、维管组织形成以及营养与生殖生长中具有重要作用,这为今后OsSHR2在分子育种等领域的应用奠定理论基础。
中图分类号:
张占田, 孙雅菲, 艾昊, 罗闻真, 冯冰, 孙文献, 徐国华, 孙淑斌. 水稻转录因子基因OsSHR2的表达特征及其在营养生长中的调控作用[J]. 中国水稻科学, 2018, 32(5): 427-436.
Zhantian ZHANG, Yafei SUN, Hao AI, Wenzhen LUO, Bing FENG, Wenxian SUN, Guohua XU, Shubin SUN. Expression Patterns and Regulation of Transcription Factor Gene OsSHR2 in Vegetative Growth in Rice[J]. Chinese Journal OF Rice Science, 2018, 32(5): 427-436.
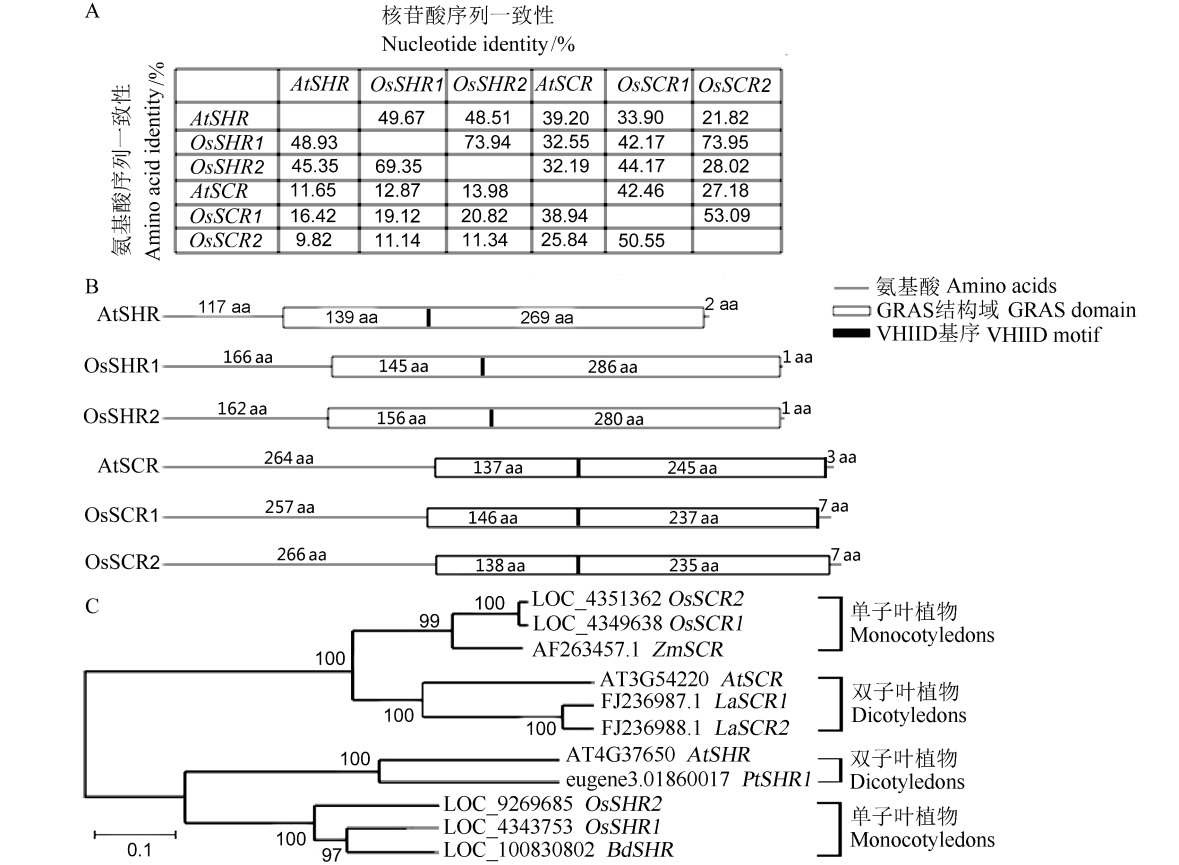
图1 不同物种SHR和SCR同源基因的生物信息学分析 A–拟南芥和水稻的SHR和SCR同源基因氨基酸和核苷酸的序列一致性分析;B–通过基因家族分析网站InterPro(http://www.ebi.ac.uk/ interpro/)进行拟南芥与水稻的SHR和SCR同源基因GRAS保守结构域和保守基序VHIID的位置预测;C–不同物种SHR和SCR同源基因的进化树分析。
Fig. 1. Bioinformatics analysis of SHR and SCR homologous genes of different species. A, Sequence homology analysis of amino acids and nucleic acid of SHR and SCR homologous genes in Arabidopsis thaliana and rice; B, Prediction of the positions of SHR and SCR homologous GRAS conserved domains and conserved motifs in Arabidopsis thaliana and rice by InterPro (http://www.ebi.ac.uk/interpro/); C, Phylogenetic tree analysis of SHR and SCR homologous genes of different species.
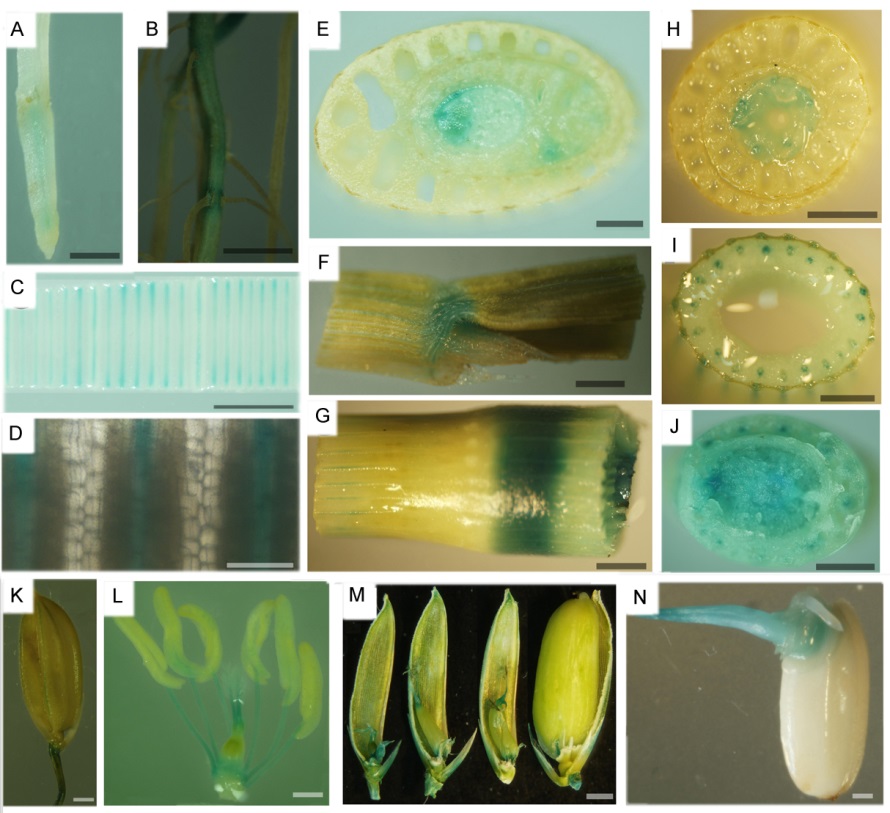
图3 pOsSHR2::GUS材料的GUS染色鉴定 A–种子根;B–种子根及侧根;C–新叶;D–新叶的放大图;E–叶原基;F–叶舌;G–茎节;H–茎及叶鞘;I–茎;J–茎基部;K–颖壳及小穗轴;L–颖花;M–子房和柱头;N–胚(萌发后3 d)。A~E,J:水稻苗期;F,G,I,K~N:水稻灌浆期;H:分蘖期。A~J中标尺为2 mm;K~N中标尺为0.5 mm。
Fig. 3. Identification of OsSHR2 promoter-driven tissue-specific GUS staining. A, Seed root; B, Seed root and lateral root; C, Young leaf; D, The enlarged view of the young leaf; E, Leaf primordium; F, Ligule; G, Node; H, Stem and leaf sheath; I, Stem; J, Basal stem; K, Husk and rachilla; L, Spikelet; M, Ovary and stigma; N, Embryo(three days after germination). A-E, J: Seedling stage; F, G, I, K-N: Grain filling stage; H, Tilling stage. A-J, Bar=2 mm; K-N, Bar=0.5 mm.
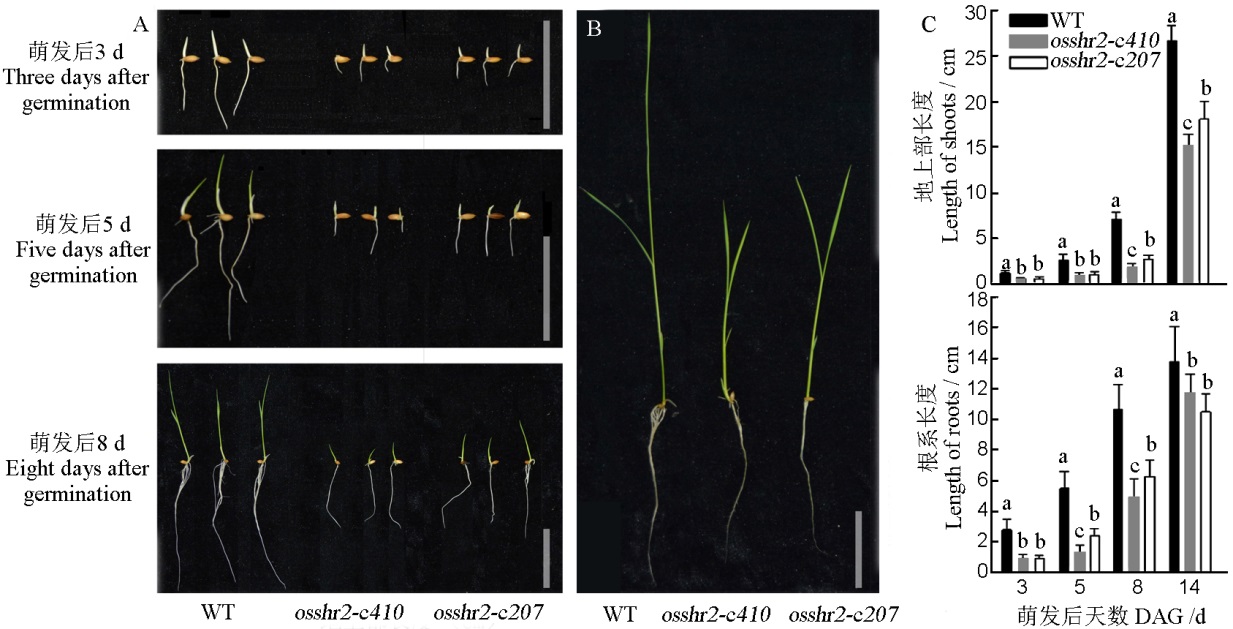
图4 WT和突变体osshr2的萌发及水培下的表型 A–萌发表型(萌发后3 d、5 d和8 d),标尺为5 cm;B–水培表型(萌发后14 d),标尺为5 cm;C–材料地上部和根系长度统计(萌发后3 d、5 d、8 d和14 d);DAG–萌发后天数。
Fig. 4. Phenotype and statistics of seed germination and hydroponics experiments of WT and osshr2. A, Germination phenotype(3, 5 and 8 days after germination), bar=5 cm; B, Hydroponics phenotype(14 days after germination), bar=5 cm; C, Shoot and root length (3, 5, 8 and 14 days after germination); DAG, Day after germination.
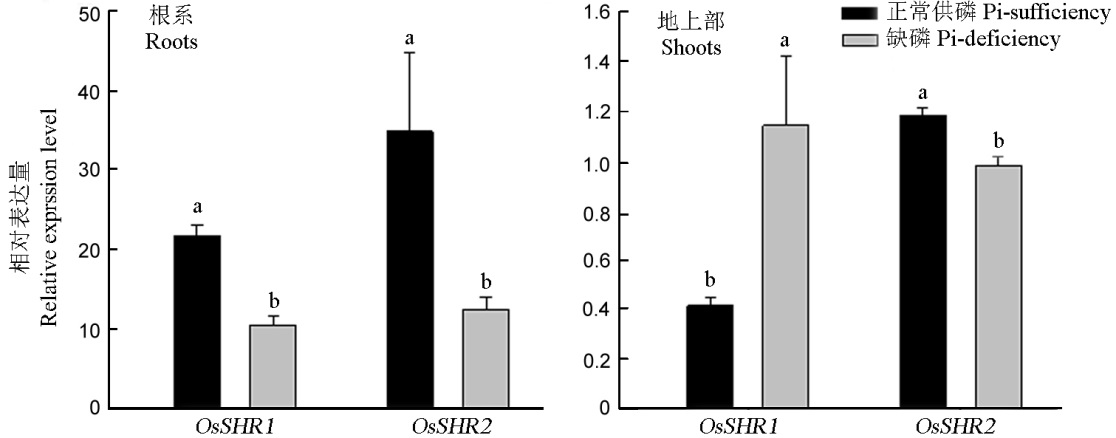
图5 OsSHR1和OsSHR2在水稻正常供磷与缺磷条件下的相对表达量 A–OsSHR1和OsSHR2在水稻地下部的相对表达量;B–OsSHR1和OsSHR2在水稻地上部的相对表达量。
Fig. 5. Relative expression level of OsSHR1 and OsSHR2 under Pi-sufficient and Pi-deficient conditions in rice. A, The relative expression of OsSHR1 and OsSHR2 in the roots; B, The relative expression of OsSHR1 and OsSHR2 in the shoots.
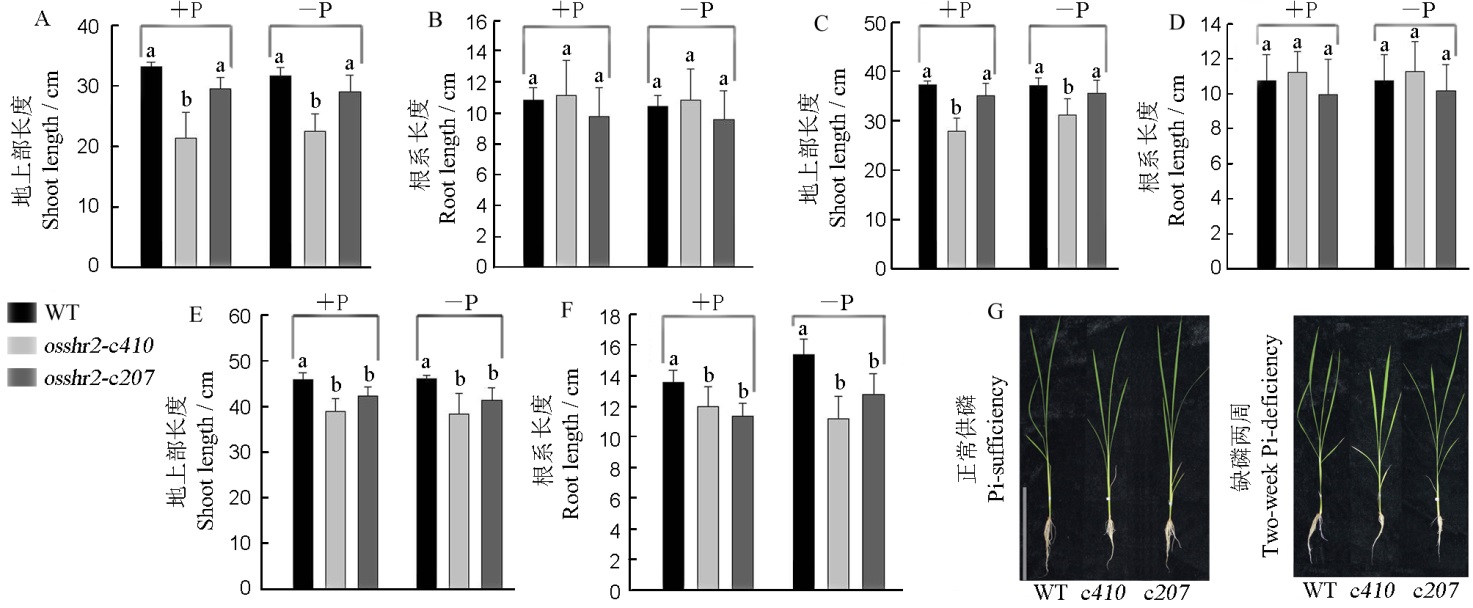
图6 WT和突变体osshr2在正常供磷和缺磷条件下的表型 A和B–缺磷3 d;C和D–缺磷1周;E和F–缺磷2周;G–缺磷2周的表型, 标尺为20 cm。
Fig. 6. Phenotype of the wild type(WT) and osshr2 under Pi-sufficient and Pi-deficient conditions. A and B, Pi-deficient lasting three days; C and D, Pi-deficient lasting one week; E and F, Pi-deficient lasting two weeks; G, The phenotype of material under Pi-sufficient and Pi-deficient lasting two weeks, bar=20 cm. c410, osshr2-c410; c207, osshr2-c207.
| [1] | 朱义旺, 林雅容, 陈亮. 我国水稻分子育种研究进展. 厦门大学学报, 2016, 55(5): 661-671. |
| Zhu Y W, Lin Y R, Chen L.Research progress of rice molecular breeding in China.J Xiamen Univ, 2016, 55(5): 661-671. (in Chinese with English abstract) | |
| [2] | Riechmann J L, Heard J, Martin G, Reuber L, Jiang C Z, Keddie J, Adam L, Pineda O, Ratcliffe O J, Samaha R R, Creelman R, Pilgrim M, Broun P, Zhang J Z, Ghandehari D, Sherman B K, Yu G L.Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science, 2000, 290(5499): 2105-2110. |
| [3] | Pysh L D, Wysocka-Diller J W, Christine C, David B, Benfey P N. The GRAS gene family in Arabidopsis: Sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J Cell & Mol Biol, 1999, 18(1): 111. |
| [4] | Cui H, Levesque M P, Vernoux T, Jung J W, Paquette A J, Gallagher K L, Wang J Y, Blilou I, Scheres B, Benfey P N.An evolutionarily conserved mechanism delimiting SHR movement defines a single layer of endodermis in plants.Science, 2007, 316(5823): 421-425. |
| [5] | Dolan L.SCARECROWs at the Border.Science, 2007, 316(5823): 377-378. |
| [6] | Wu S, Lee C M, Hayashi T, Pricea S, Divolb F, Henryb S, Pauluzzib G, Perinb C, Gallaghera K L.A plausible mechanism, based upon short-root movement, for regulating the number of cortex cell layers in roots.Proc Natl Acad Sci USA, 2014, 111(45): 16184-16189. |
| [7] | Benfey P N, Scheres B.Root development.Curr Biol, 2000, 10(22): 813-815. |
| [8] | Benfey P N, Linstead P J, Roberts K, Schiefelbein J W, Hauser M T, Aeschbacher R A.Root development in Arabidopsis: Four mutants with dramatically altered root morphogenesis.Development, 1993, 119(1): 57-70. |
| [9] | Laurenzio L D, Wysockadiller J, Malamy J E, Pysh L, Helariutta Y, Freshour G, Hahn M G, Feldmann K A, Benfey P N.The SCARECROW gene regulates an asymmetric cell division that is essential for generating the radial organization of the Arabidopsis root. Cell, 1996, 86(3): 423-433. |
| [10] | 倪君. OsIAA23介导的生长素信号胚后维持水稻根静止中心. 杭州: 浙江大学, 2011. |
| Ni J.OsIAA23-mediated auxin signaling defines postembryonic maintenance of QC in primary in rice. Hangzhou: Zhejiang University, 2011. (in Chinese with English abstract) | |
| [11] | Lim J, Benfey P N.Molecular analysis of the SCARECROW gene in maize reveals a common basis for radial patterning in diverse meristems.Discuss Pap, 2000, 12(8): 1307-1318. |
| [12] | Sbabou L, Bucciarelli B, Miller S, Liu J, Berhada F, Filali-Maltouf A, Allan D, Vance C.Molecular analysis of SCARECROW genes expressed in white lupin cluster roots. J Exp Bot, 2010, 61(5): 1351-1363. |
| [13] | Wang J, Anderssongunneras S, Gaboreanu I, Hertzberg M, Tucker M R, Zheng B, Lesniewska J, Mellerowicz E J, Laux T, Sandberg G, Jones B.Reduced expression of the SHORT-ROOT gene increases the rates of growth and development in hybrid poplar and Arabidopsis. PloS ONE, 2011, 6(12): e28878. |
| [14] | Wysockadiller J W, Helariutta Y, Fukaki H, Malamy J E, Benfey P N.Molecular analysis of SCARECROW function reveals a radial patterning mechanism common to root and shoot. Development, 2000, 127(3): 595-603. |
| [15] | 霍胜楠. 水稻胚胎发生相关基因的表达及其功能鉴定. 济南: 山东农业大学, 2008. |
| Huo S N.Isolation and characterization of rice genes involved in embryo development. Jinan: Shandong Agricultural University, 2008. (in Chinese with English abstract) | |
| [16] | Cui H, Kong D, Liu X, Hao Y.SCARECROW, SCR-LIKE 23 and SHORT-ROOT control bundle sheath cell fate and function in Arabidopsis thaliana. Plant J Cell & Mol Biol, 2014, 78(2): 319-327. |
| [17] | Gao X R, Wang C L, Cui H C.Identification of bundle sheath cell fate factors provides new tools for C3-to-C4 engineering.Plant Signal & Behav, 2014, 9(6): e29163. |
| [18] | Morikami A.The SCARECROW gene’s role in asymmetric cell divisions in rice plants. Plant J, 2003, 36(1): 45-54. |
| [19] | Lucas M, Swarup R, Paponov I A, Swarup K, Casimiro I, Lake D, Peret B, Zappala S, Mairhofer S, Whitworth M, Wang J H, Ljung K, Marchant A, Sandberg G, Holdsworth M J, Palme K, Pridmore T, Mooney S, Bennett M J.Short-Root regulates primary, lateral, and adventitious root development in Arabidopsis. Plant Physiol, 2011, 155(1): 384-398. |
| [20] | Tian H, Jia Y, Niu T, Yu Q, Ding Z.The key players of the primary root growth and development also function in lateral roots in Arabidopsis. Plant Cell Rep, 2014, 33(5): 745-753. |
| [21] | Goh T, Toyokura K, Wells D M, Swarup K, Yamamoto M, Mimura T, Weijers D, Fukaki H, Laplaze L, Bennett M J, Guyomarc’h S.Quiescent center initiation in the Arabidopsis lateral root primordia is dependent on the SCARECROW transcription factor. Development, 1991, 143(18): 3363. |
| [22] | Lavenus J, Goh T, Guyomarc’h S, Hill K, Lucas M, Voß U, Kenobi K, Wilson M H, Farcot E, Hagen G, Guilfoyle T J, Fukaki H, Laplaze L, Bennettb M J.Inference of the Arabidopsis lateral root gene regulatory network suggests a bifurcation mechanism that defines primordia flanking and central zones. Plant Cell, 2015, 27(5): 1368-1388. |
| [23] | Bieleski R.Phosphate pools, phosphate transport, and phosphate availability.Ann Rev Plant Physiol, 1973, 24(1): 225-252. |
| [24] | Muchhal U S, Pardo J M, Raghothama K G.Phosphate transporters from the higher plant Arabidopsis thaliana.Proc Natl Acad Sci USA, 1996, 93(19): 10519-105123. |
| [25] | Wang L, Shan L, Ye Z, Li Z, Du X, Liu D. Comparative genetic analysis of Arabidopsis purple acid phosphatases AtPAP10, AtPAP12,AtPAP26 provides new insights into their roles in plant adaptation to phosphate deprivation. J Integr Plant Biol, 2014, 56(3): 299-314. |
| [26] | Rausch C, Bucher M.Molecular mechanisms of phosphate transport in plants.Planta, 2002, 216(1): 23-37. |
| [27] | Paszkowski U, Kroken S, Roux C, Briggs S P.Rice phosphate transporters include an evolutionarily divergent gene specifically activated in arbuscular mycorrhizal symbiosis.Proc Natl Acad Sci USA, 2002, 99(20): 13324-13329. |
| [28] | Liu F, Chang X J, Ye Y, Xie W B, Wu P, Lian X M.Comprehensive sequence and whole-life-cycle expression profile analysis of the phosphate transporter gene family in rice.Mol Plant, 2011, 4(6): 1105-1122. |
| [29] | Zhang F, Sun Y, Pei W, Jain A, Sun R, Cao Y, Wu X N, Jiang T T, Zhang L, Fan X R, Chen A Q, Shen Q R, Xu G H, Sun S B.Involvement of OsPht1;4 in phosphate acquisition and mobilization facilitates embryo development in rice. Plant J Cell & Mol Biol, 2015, 82(4): 556. |
| [30] | Rubio V, Linhares F, Solano R, Martín A C, Iglesias J, Leyva A, Paz-Ares J.A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae.Genes & Dev, 2001, 15(16): 2122-2133. |
| [31] | Bustos R, Castrillo G, Linhares F, Puga M I, Rubio V, Perez-Perez J, Solano R, Leyva A, Paz-Ares J.A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PloS Genet, 2010, 6(9): e1001102. |
| [32] | Wu P, Wang X.Role of OsPHR2 on phosphorus homeostasis and root hairs development in rice(Oryza sativa L.). Plant Signal & Behav, 2008, 3(9): 674-675. |
| [33] | Zhou J, Jiao F, Wu Z C, Li Y Y, Wang X M, He X W, Zhong W Q, Wu P.OsPHR2 is involved in phosphate- starvation signaling and excessive phosphate accumulation in shoots of plants. Plant Physiol, 2008, 146(4): 1673-1686. |
| [34] | Raghothama K G, Maggio A, Narasimhan M L, Kononowicz A K, Wang G, D’Urzo M P, Hasegawa P M, Bressanl R A. Tissue-specific activation of the OsMotin gene by ABA, C2H4 and NaCl involves the same promoter region. Plant Mol Biol, 1997, 34(3): 393-402. |
| [35] | Liao H, Rubio G, Yan X, Cao A, Brown K M, Lynch J P.Effect of phosphorus availability on basal root shallowness in common bean.Plant & Soil, 2001, 232(1): 69-79. |
| [1] | 任志奇, 薛可欣, 董铮, 李小湘, 黎用朝, 郭玉静, 刘文强, 郭梁, 盛新年, 刘之熙, 潘孝武. 水稻外卷叶突变体ocl1的鉴定及基因定位[J]. 中国水稻科学, 2023, 37(4): 337-346. |
| [2] | 肖乐铨, 李雷, 戴伟民, 强胜, 宋小玲. 转cry2A*/bar基因水稻与杂草稻杂交后代的苗期生长特性[J]. 中国水稻科学, 2023, 37(4): 347-358. |
| [3] | 李刚, 高清松, 李伟, 张雯霞, 王健, 程保山, 王迪, 高浩, 徐卫军, 陈红旗, 纪剑辉. 定向敲除SD1基因提高水稻的抗倒性和稻瘟病抗性[J]. 中国水稻科学, 2023, 37(4): 359-367. |
| [4] | 汪胜勇, 陈宇航, 陈会丽, 黄钰杰, 张啸天, 丁双成, 王宏伟. 水稻减数分裂期高温对苯丙烷类代谢及下游分支代谢途径的影响[J]. 中国水稻科学, 2023, 37(4): 368-378. |
| [5] | 董立强, 杨铁鑫, 李睿, 商文奇, 马亮, 李跃东, 隋国民. 株行距配置对超高产田水稻产量及根系形态生理特性的影响[J]. 中国水稻科学, 2023, 37(4): 392-404. |
| [6] | 韩聪, 何禹畅, 吴丽娟, 郏丽丽, 王磊, 鄂志国. 水稻碱性亮氨酸拉链(bZIP)蛋白家族功能研究进展[J]. 中国水稻科学, 2023, 37(4): 436-448. |
| [7] | 沈雨民, 陈明亮, 熊焕金, 熊文涛, 吴小燕, 肖叶青. 水稻内外稃异常发育突变体blg1 (beak like grain 1)的表型分析与精细定位[J]. 中国水稻科学, 2023, 37(3): 225-232. |
| [8] | 段敏, 谢留杰, 高秀莹, 唐海娟, 黄善军, 潘晓飚. 利用CRISPR/Cas9技术创制广亲和水稻温敏雄性不育系[J]. 中国水稻科学, 2023, 37(3): 233-243. |
| [9] | 程玲, 黄福钢, 邱一埔, 王心怡, 舒宛, 邱永福, 李发活. 籼稻材料570011抗褐飞虱基因的遗传分析及鉴定[J]. 中国水稻科学, 2023, 37(3): 244-252. |
| [10] | 王文婷, 马佳颖, 李光彦, 符卫蒙, 李沪波, 林洁, 陈婷婷, 奉保华, 陶龙兴, 符冠富, 秦叶波. 高温下不同施肥量对水稻产量品质形成的影响及其与能量代谢的关系分析[J]. 中国水稻科学, 2023, 37(3): 253-264. |
| [11] | 刘嫒桦, 李小坤. 不同肥料施用与稻米品质关系的整合分析[J]. 中国水稻科学, 2023, 37(3): 276-284. |
| [12] | 杨晓龙, 王彪, 汪本福, 张枝盛, 张作林, 杨蓝天, 程建平, 李阳. 不同水分管理方式对旱直播水稻产量和稻米品质的影响[J]. 中国水稻科学, 2023, 37(3): 285-294. |
| [13] | 魏晓东, 宋雪梅, 赵凌, 赵庆勇, 陈涛, 路凯, 朱镇, 黄胜东, 王才林, 张亚东. 硅锌肥及其施用方式对南粳46产量和稻米品质的影响[J]. 中国水稻科学, 2023, 37(3): 295-306. |
| [14] | 林聃, 江敏, 苗波, 郭萌, 石春林. 水稻高温热害模型研究及其在福建省的应用[J]. 中国水稻科学, 2023, 37(3): 307-320. |
| [15] | 郑承梅, 孙金秋, 刘梦杰, 杨永杰, 陆永良, 郭怡卿, 唐伟. 水稻田糠稷种子萌发和出苗特性及化学防除药剂筛选[J]. 中国水稻科学, 2023, 37(3): 321-328. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||