
Chinese Journal OF Rice Science ›› 2020, Vol. 34 ›› Issue (4): 368-382.DOI: 10.16819/j.1001-7216.2020.9108
• Research Papers • Previous Articles
Li LI1, Xuyan MO2, Tiantian LI1, Liyuan ZHANG1,3,*( ), Hansong DONG1,2,3
), Hansong DONG1,2,3
Received:2019-10-08
Revised:2020-03-06
Online:2020-07-10
Published:2020-07-10
Contact:
Liyuan ZHANG
李丽1, 莫旭艳2, 李甜甜1, 张丽媛1,3,*( ), 董汉松1,2,3
), 董汉松1,2,3
通讯作者:
张丽媛
基金资助:CLC Number:
Li LI, Xuyan MO, Tiantian LI, Liyuan ZHANG, Hansong DONG. Effector XopN of Xanthomonas oryzae pv. oryzae Plays a Virulent Role in Rice Varieties Harboring OsSWEET11 Homologs[J]. Chinese Journal OF Rice Science, 2020, 34(4): 368-382.
李丽, 莫旭艳, 李甜甜, 张丽媛, 董汉松. 白叶枯病菌效应子XopN在拥有OsSWEET11同源基因的水稻品种上发挥毒性作用[J]. 中国水稻科学, 2020, 34(4): 368-382.
Add to citation manager EndNote|Ris|BibTeX
URL: http://www.ricesci.cn/EN/10.16819/j.1001-7216.2020.9108
| 水稻品种 Rice variety | 抗病基因 Resistance gene | 基因产物 Encoding protein | 文献 Reference |
|---|---|---|---|
| IRBB1 | Xa1 | NLR (leucine-rich repeat receptor) | 31-33 |
| IRBB3 | Xa3 | LRR-RLK (leucine-rich repeat-receptor-like kinase) | 31-33 |
| IRBB8 | xa8 | Unknown | 31, 34 |
| IRBB10 | Xa10 | Executor | 31, 33 |
| IRBB13 | xa13 | OsSWEET11 (synonym Xa13) | 31, 33 |
| IRBB14 | Xa14 | Unknown | 31 |
| IRBB203 | Xa3 | LRR-RLK | 31, 33 |
| IRBB204 | Xa4 | RLK | 31, 33 |
| IRBB205 | Xa5 | TFIIA TF | 31, 33 |
| IRBB208 | Xa8 | Putative OsSWEET11 | 本研究This study |
| IRBB211 | Xa11 | RLK | AAY32487.1 |
| IR24 | Xa18 | RLK | AAY32494 |
| Asominori | Putative Xa13 | Putative OsSWEET11 | 本研究This study |
| 日本晴 Nipponbare | Xa13 (synonym Os8N3) | OsSWEET11 (synonym XA13) | 33-36 |
Table 1 Xoo-relevant resistance gene and encoding proteins in the tested rice varieties.
| 水稻品种 Rice variety | 抗病基因 Resistance gene | 基因产物 Encoding protein | 文献 Reference |
|---|---|---|---|
| IRBB1 | Xa1 | NLR (leucine-rich repeat receptor) | 31-33 |
| IRBB3 | Xa3 | LRR-RLK (leucine-rich repeat-receptor-like kinase) | 31-33 |
| IRBB8 | xa8 | Unknown | 31, 34 |
| IRBB10 | Xa10 | Executor | 31, 33 |
| IRBB13 | xa13 | OsSWEET11 (synonym Xa13) | 31, 33 |
| IRBB14 | Xa14 | Unknown | 31 |
| IRBB203 | Xa3 | LRR-RLK | 31, 33 |
| IRBB204 | Xa4 | RLK | 31, 33 |
| IRBB205 | Xa5 | TFIIA TF | 31, 33 |
| IRBB208 | Xa8 | Putative OsSWEET11 | 本研究This study |
| IRBB211 | Xa11 | RLK | AAY32487.1 |
| IR24 | Xa18 | RLK | AAY32494 |
| Asominori | Putative Xa13 | Putative OsSWEET11 | 本研究This study |
| 日本晴 Nipponbare | Xa13 (synonym Os8N3) | OsSWEET11 (synonym XA13) | 33-36 |
| 菌株及质粒 Strains and plasmids | 相关性状 The study-relevant characteristics | 来源 Resource |
|---|---|---|
| 菌株Bacterial strains 黄单胞菌Xanthomonas oryzae pv. oryzae | ||
| PXO99A | PXO99的氮杂胞苷抗性克隆,也叫菲律宾小种6号 The azacytidine-resistant clone of PXO99, synonym Philippine race 6 | 本实验室This lab |
| ∆XopN | PXO99A XopN敲除突变体 PXO99A XopN-knockout unmarked mutant | 本研究This study |
| ∆XopN/XopN | 用质粒pHMXopN回补PXO99A XopN敲除突变体 PXO99A XopN mutant complemented with pHMXopN | 本研究This study |
| ∆XopQ | PXO99A XopQ敲除突变体 PXO99A XopQ-knockout unmarked mutant | 本实验室This lab |
| 质粒Plasmids | ||
| pMD19-Ts | 基因克隆T载体,含Apr, Mob+, Mob(P), LacZa+, Ampr Gene-cloning T-simple vector, Apr, Mob+, Mob(P), LacZa+, Ampr | TaKaRa |
| pMD20-T | 基因重组T载体,含Kanr, Ampr Gene-recombination T-simple vector, Kanr, Ampr | TaKaRa |
| pMD20-T::F1::Km::F2 | 重组的基因突变载体 Recombinant gene-mutagenicity vector | 本研究This study |
| pHMI | 多个pUC19重组的寄主更广的载体,含SpR Broad-host range vector with pUC19 polylinker, SpR | 本实验室This lab |
| pHMI-XopN | XopN整合到pHMI载体lacz启动子上的重组基因表达载体 Recombinant gene-expression vector containing XopN fused to lacZ promoter of pHMI | 本研究This study |
Table 2 Bacterial strains and plasmid vectors used and created in this study.
| 菌株及质粒 Strains and plasmids | 相关性状 The study-relevant characteristics | 来源 Resource |
|---|---|---|
| 菌株Bacterial strains 黄单胞菌Xanthomonas oryzae pv. oryzae | ||
| PXO99A | PXO99的氮杂胞苷抗性克隆,也叫菲律宾小种6号 The azacytidine-resistant clone of PXO99, synonym Philippine race 6 | 本实验室This lab |
| ∆XopN | PXO99A XopN敲除突变体 PXO99A XopN-knockout unmarked mutant | 本研究This study |
| ∆XopN/XopN | 用质粒pHMXopN回补PXO99A XopN敲除突变体 PXO99A XopN mutant complemented with pHMXopN | 本研究This study |
| ∆XopQ | PXO99A XopQ敲除突变体 PXO99A XopQ-knockout unmarked mutant | 本实验室This lab |
| 质粒Plasmids | ||
| pMD19-Ts | 基因克隆T载体,含Apr, Mob+, Mob(P), LacZa+, Ampr Gene-cloning T-simple vector, Apr, Mob+, Mob(P), LacZa+, Ampr | TaKaRa |
| pMD20-T | 基因重组T载体,含Kanr, Ampr Gene-recombination T-simple vector, Kanr, Ampr | TaKaRa |
| pMD20-T::F1::Km::F2 | 重组的基因突变载体 Recombinant gene-mutagenicity vector | 本研究This study |
| pHMI | 多个pUC19重组的寄主更广的载体,含SpR Broad-host range vector with pUC19 polylinker, SpR | 本实验室This lab |
| pHMI-XopN | XopN整合到pHMI载体lacz启动子上的重组基因表达载体 Recombinant gene-expression vector containing XopN fused to lacZ promoter of pHMI | 本研究This study |
| 基因或基因片段 a Gene or partial sequence a | 引物 Primer | 酶切位点 Restriction site |
|---|---|---|
| F1 | 5'-AAGCTTGCCGACATGAAGGTGTATGAGGTG-3' | Hind Ⅲ |
| 5'-ACTAGTTGCGGCACCCGATGCTGCTGC-3' | SpeⅠ | |
| F2 | 5'-GGTACCTGGCCGCTGCCGGGTAATGC-3' | KpnⅠ |
| 5'-GAATTCGCGTGCTAATCAGGTTTGCCTTTG-3' | EcoRⅠ | |
| Kan | 5'-AGTGATTGCGCCTACCCGG-3' | 无None |
| 5'-ACGTCTTGAGCGATTGTGTAGG-3' | 无None | |
| XopN | 5'-GACCATGATTACGCCAAGCTTCTGGCAACAGCATCCCTCC-3' | Hind Ⅲ |
| 5'-GACCTGCAGGCATGCAAGCTTTTACGCCGGCGGCAGTGCCCGATC-3' | Hind Ⅲ |
Table 3 Information of primers used in PCR protocols for Xoo XopN knockout and complementation.
| 基因或基因片段 a Gene or partial sequence a | 引物 Primer | 酶切位点 Restriction site |
|---|---|---|
| F1 | 5'-AAGCTTGCCGACATGAAGGTGTATGAGGTG-3' | Hind Ⅲ |
| 5'-ACTAGTTGCGGCACCCGATGCTGCTGC-3' | SpeⅠ | |
| F2 | 5'-GGTACCTGGCCGCTGCCGGGTAATGC-3' | KpnⅠ |
| 5'-GAATTCGCGTGCTAATCAGGTTTGCCTTTG-3' | EcoRⅠ | |
| Kan | 5'-AGTGATTGCGCCTACCCGG-3' | 无None |
| 5'-ACGTCTTGAGCGATTGTGTAGG-3' | 无None | |
| XopN | 5'-GACCATGATTACGCCAAGCTTCTGGCAACAGCATCCCTCC-3' | Hind Ⅲ |
| 5'-GACCTGCAGGCATGCAAGCTTTTACGCCGGCGGCAGTGCCCGATC-3' | Hind Ⅲ |
| 水稻品种 Rice variety | 基因 Gene | 登录号 Locus number | 文献 Reference | 引物 Primer | PCR产物长度 Product size / bp | 用途 Subject | ||
|---|---|---|---|---|---|---|---|---|
| 所有品种 All varieties | OsEF1α | AF030517 | 9-13 | 5'-CGTGCCTGTGGGTCGTGTTG-3' 5'-TCCTGGAGAGCCTCGTGGTG-3' | 120 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 日本晴Nipponbare | Xa13 (OsSWEET11) | AK070510 | 10 | 5'-TGGTTCTGCTACGGCCTCTT-3' 5'-GGTACCAGAAGTAGAGCCCCATCT-3' | 103 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| IRBB208, Asominori | 假定的OsSWEET11同源物 Hypothetic OsSWEET11 homologs | AK070510 | 本研究 This study | 5'-TGGTTCTGCTACGGCCTCTT-3' 5'-GGTACCAGAAGTAGAGCCCCATCT-3' | 103 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 日本晴, IRBB208 Asomimori Nipponbare, IRBB208 and Asominori | SWEET11与其假定同源物 OsSWEET11 and its homologs | AK070510 | 本研究 This study | 5'-CCAAGGCCAAACCACACATG-3' 5'-TAACTACTAATAATACCTG-3' | 1517 (Full length) | 通过RT-PCR 进行基因克隆,验证产物序列 Gene cloning by RT-PCR and confir mation of the product sequences | ||
| IRBB13 | xa13 (OsSWEET11) | DQ421394.1 | 33 | 5'-GGCGTGCACAAGGTCGAGGT-3' 5'-CTGGCCGTCATCGATCCGGC-3' | 205(3911-4116 in DQ421394.1) | 通过RT-qPCR进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 5'-TACCCGAACGTGGGCGGCTT-3' 5'-AGTAACTACTAATAATACCTGTC-3' | 752(3701-4453 in DQ421394.1) | 通过RT-PCR进行基因克隆,验证产物序列Gene cloning by RT-PCR and confirmation of the product sequence | ||||||
Table 4 Information on rice genes and their primers used for RT-PCR and RT-qPCR analyses.
| 水稻品种 Rice variety | 基因 Gene | 登录号 Locus number | 文献 Reference | 引物 Primer | PCR产物长度 Product size / bp | 用途 Subject | ||
|---|---|---|---|---|---|---|---|---|
| 所有品种 All varieties | OsEF1α | AF030517 | 9-13 | 5'-CGTGCCTGTGGGTCGTGTTG-3' 5'-TCCTGGAGAGCCTCGTGGTG-3' | 120 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 日本晴Nipponbare | Xa13 (OsSWEET11) | AK070510 | 10 | 5'-TGGTTCTGCTACGGCCTCTT-3' 5'-GGTACCAGAAGTAGAGCCCCATCT-3' | 103 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| IRBB208, Asominori | 假定的OsSWEET11同源物 Hypothetic OsSWEET11 homologs | AK070510 | 本研究 This study | 5'-TGGTTCTGCTACGGCCTCTT-3' 5'-GGTACCAGAAGTAGAGCCCCATCT-3' | 103 | 通过RT-qPCR 进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 日本晴, IRBB208 Asomimori Nipponbare, IRBB208 and Asominori | SWEET11与其假定同源物 OsSWEET11 and its homologs | AK070510 | 本研究 This study | 5'-CCAAGGCCAAACCACACATG-3' 5'-TAACTACTAATAATACCTG-3' | 1517 (Full length) | 通过RT-PCR 进行基因克隆,验证产物序列 Gene cloning by RT-PCR and confir mation of the product sequences | ||
| IRBB13 | xa13 (OsSWEET11) | DQ421394.1 | 33 | 5'-GGCGTGCACAAGGTCGAGGT-3' 5'-CTGGCCGTCATCGATCCGGC-3' | 205(3911-4116 in DQ421394.1) | 通过RT-qPCR进行基因表达量分析 Gene expression analysis by RT-qPCR | ||
| 5'-TACCCGAACGTGGGCGGCTT-3' 5'-AGTAACTACTAATAATACCTGTC-3' | 752(3701-4453 in DQ421394.1) | 通过RT-PCR进行基因克隆,验证产物序列Gene cloning by RT-PCR and confirmation of the product sequence | ||||||
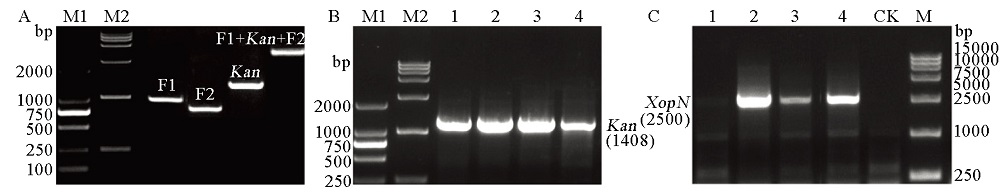
Fig. 1. XopN gene knockout from PXO99A. A, Verification of the gene-knockout recombinant vector pMD20-T::F1::Kan::F2 by PCR amplification of the indicated specific recombinant elements; M1, M2, DNA markers. B, Screening of the XopN-knockout mutant ∆XopN based on PCR amplification of Kan from genomic DNAs of the conjugate clones #1 to #4. M1, M2, DNA markers. C, Confirmation of the ∆XopN mutant by PCR amplification of the XopN from genomic DNAs of the conjugate clones #1 to #4 analyzed together with a control, which was the clone of bacteria transformed with the empty pMD20-T vector. CK, Blank control; M, DNA marker.
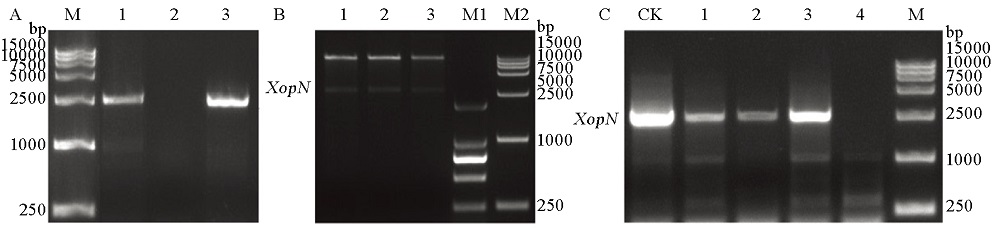
Fig. 2. Genetic complementation of the ∆XopN mutant. A, Escherichia coli colonial PCR analysis by using genomic DNA samples from the pHMI::XopN conjugate colonies #1 to #3; M, DNA marker. B, Restriction PCR analysis of the PCR products that were collected from A and digested by Hind Ⅲ. M1, M2, DNA markers. C, PCR analysis of genomic DNA samples from the pHMI::XopN conjugate colonies #1 to #3. CK, Control; M, DNA marker. In A to C, PCR was performed with the XopN-specific primers.
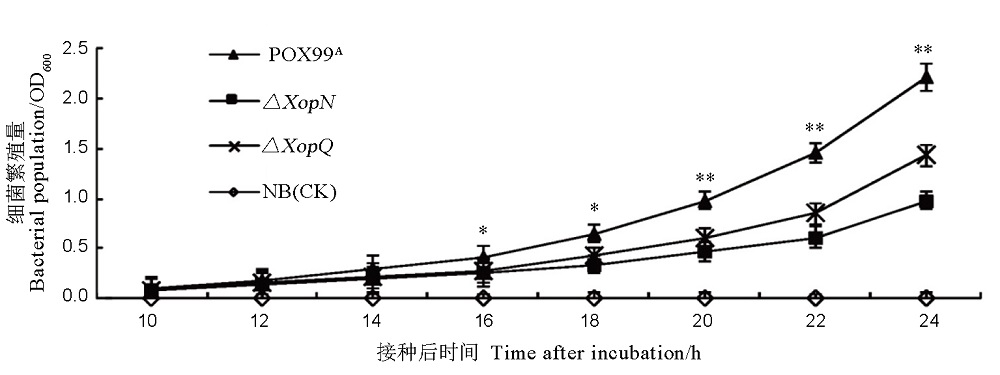
Fig. 3. Bacterial population of the indicated Xoo strains in NB medium. Data are shown as mean values ± SD. Significant differences exist between PXO99A and both mutants 16, 18, 20, 22 and 24 hours after inoculation based on ANOVA and Duncan’s test (n = 6, P < 0.05 or 0.01).
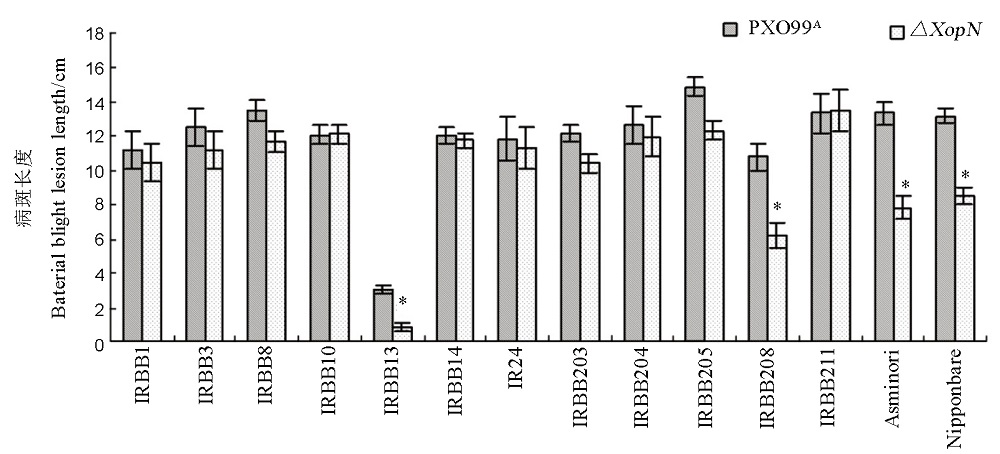
Fig. 4. Lesion Length of leaf blight caused by the Xoo strains in different rice varieties. Plants of various rice varieties at the bottom were inoculated by leaf-top clipping with bacterial suspensions of ∆XopN and PXO99A, respectively. Lesion length of leaf blight was measured two weeks later. Values are shown as mean ± standard deviation (SD) estimates of 3 independent experiments each involving 12 leaves of 6 plants. Asterisks indicate significant differences in the corresponding pair comparisons based on analysis of variance (ANOVA) and Fisher’s test at P < 0.05 (n = 12).
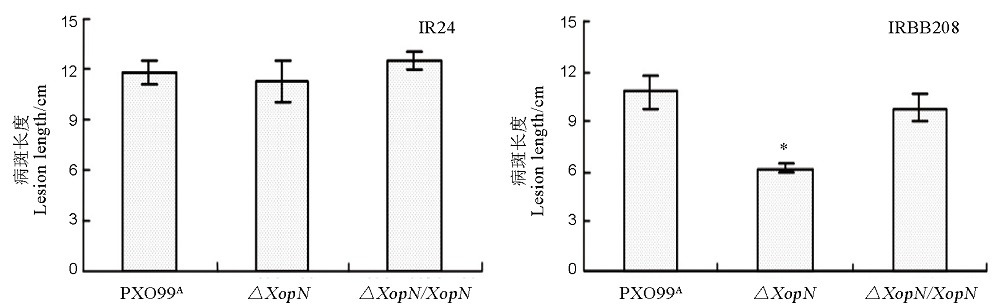
Fig. 5. Lesion length of leaf blight caused by the three Xoo strains in the two rice varieties. Plants of the rice varieties were inoculated by leaf-top clipping with bacterial suspensions of PXO99A, ∆XopN and ∆XopN/XopN, respectively. Lesion length of leaf blight was measured two weeks later. Data are mean values ± SD. Asterisks indicate significant differences in the multiple comparisons by ANOVA and Fisher’s test (P < 0.05, n = 12).
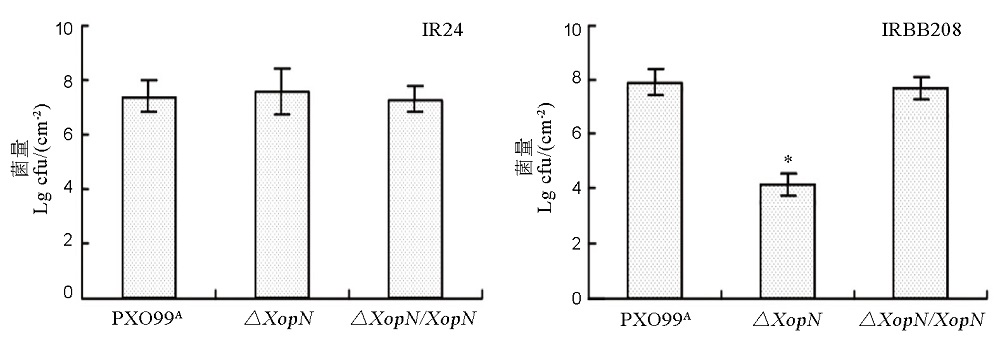
Fig. 6. Bacterial population of PXO99A, ∆XopN and ∆XopN/XopN in leaves of IR24 and IRBB208. Plants of rice varieties IR24 and IRBB208 were inoculated by leaf infiltration with bacterial suspensions prepared from each of the bacterial strains indicated at the bottom. Three days later, bacteria were recovered from leaves and scored as colony formation unit (cfu) in the given leaf size. Data are mean values±SD. Different letters indicate significant differences of multiple comparisons of data from different bacterial strains based on ANOVA and Duncan’s multiple-range test(P<0.01, n= 6).
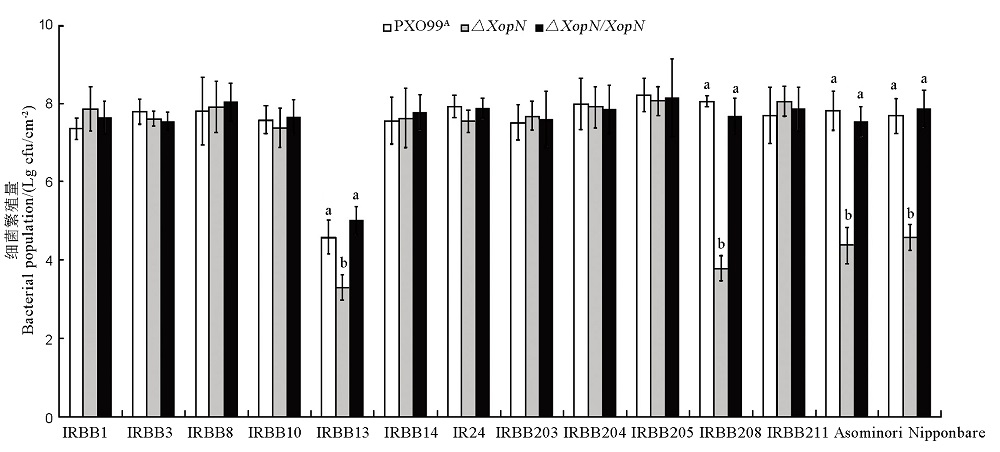
Fig. 7. Populations of PXO99A, ∆XopN and ∆XopN/XopN bacteria multiplied in leaves of the 14 rice varieties. Plants of different rice varieties at the bottom were inoculated by leaf infiltration with bacterial suspensions prepared from each of the indicated bacterial strains on the top. Three days later, bacteria were recovered from leaves and scored as cfu in the given leaf size. Data shown are mean values ± SD. Different letters on the graph indicate that ANOVA and Duncan’s multiple-range test disclose significant differences of multiple comparisons of data from the corresponding rice varieties inoculated differently (P < 0.01, n = 3).
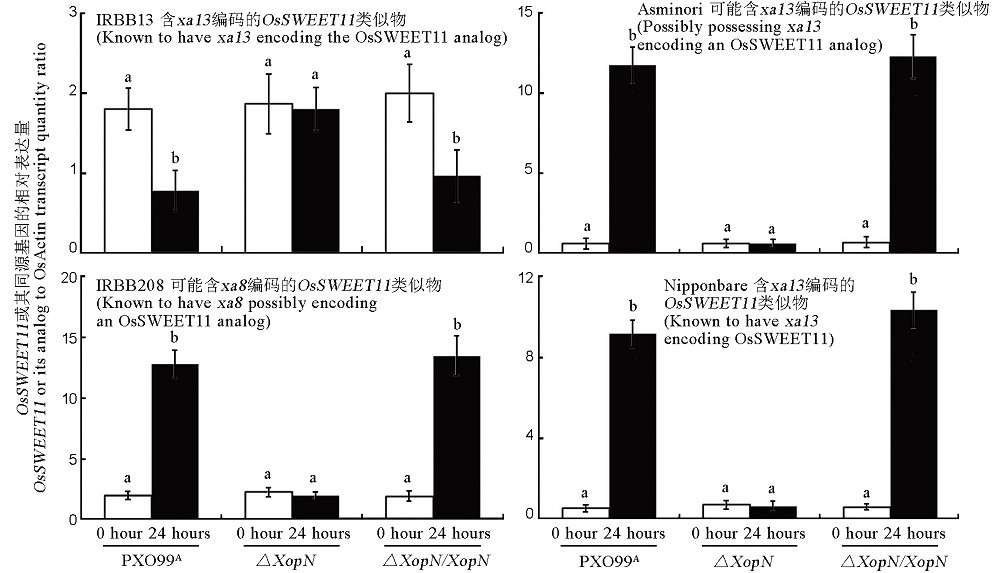
Fig. 8. Effects of XopN on expression levels of OsSWEET11 or its analogs in leaves of the 4 rice varieties. Plants were inoculated by leaf infiltration with bacterial suspensions prepared from each of the indicated bacterial strains. Immediately before inoculation (0 hour) and 24 hours after inoculation, inoculated leaves were excised and used to isolate RNAs. RNA samples were subjected to RT-qPCR in two combinations. One was IRBB13 RNA samples and a pair of primers specific to the recessive resistance gene OsSWEET11/xa13 of IRBB13. The other was RNA samples from additional 3 rice varieties and a pair of primers specific to the dominant disease-susceptibility gene OsSWEET11/Xa13 of Nipponbare. Relative levels of the genes were quantified as ratios of their transcript amounts to the transcript amount of OsEF1α, a constitutively expressed gene used as a reference. Data are mean values ± SD estimates. Different letters on graphs indicate significant differences of multiple comparisons of data from differently inoculated plants based on ANOVA and Duncan’s test (P < 0.05, n = 6).
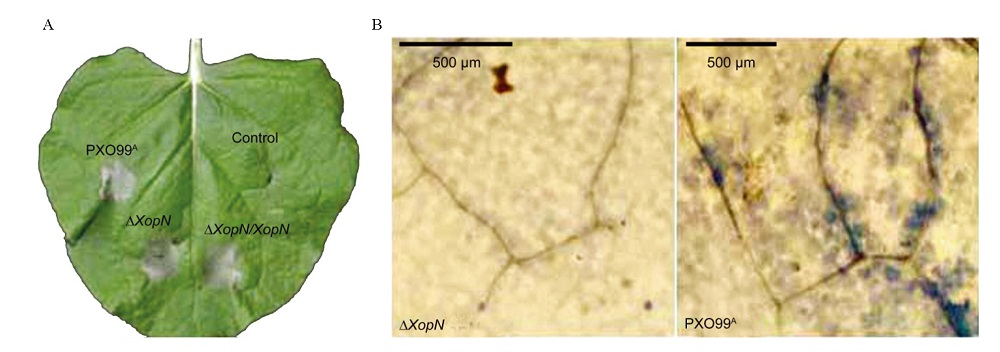
Fig. 9. Effect of XopN on the ability of Xoo to induce hypersensitive response in tobacco leaves. A, A tobacco (Nicotiana benthamiana) leaf photographed at 36 h after infiltration with pure water (control) or with a bacterial suspension of every Xoo strain shown on the leaf. This photo represents 6 leaves of 3 plants that were treated similarly. B, Parts of leaves 24 h after infiltration with the indicated bacterial suspensions and stained with trypan blue to show micro-HR. Micro-HR was visualized with a microscopy and is present in the right image but absent in the left one. Both microscopic images represent 6 infiltrated leaves excised from 3 plants.
| [1] | Mew T W.Current status and future prospects of research on bacterial blight of rice[J]. Annual Review of Phytopathology, 1987, 25: 359-382. |
| [2] | Ou S H.Rice Diseases[M]. Kew, Surrey: Commonwealth Mycological Institute, 1972: 368. |
| [3] | Buttner D, Bonas U.Regulation and secretion of Xanthomonas virulence factors[J]. FEMS Microbiology Reviews, 2010, 34(2): 107-133. |
| [4] | Mansfield J, Genin S, Magori S, Citovsky V, Sriariyanum M, Ronald P, Dow M, Verdier V, Beer S V, Machado M A, Toth I, Salmond G, Foster G D.Top 10 plant pathogenic bacteria in molecular plant pathology[J]. Molecular Plant Pathology, 2012, 13(6): 614-629. |
| [5] | Niño-Liu D O, Ronald P C, Bogdanove A J. Xanthomonas oryzae pathovars: Model pathogens of a model crop[J]. Molecular Plant Pathology, 2006, 7: 303-324. |
| [6] | Noda T, Kaku H.Growth of Xanthomonas oryzae pv. oryzae in planta and in guttation fluid of rice[J]. Annals of the Phytopathological Society of Japan, 1999, 65: 9-14. |
| [7] | Ou S H.Rice Diseases[M]. Kew, Surrey: Commonwealth Agricultural Bureaux, 1985: 109-201. |
| [8] | Park C J, Kazunari N, Ronald P C.Quantitative measurements of Xanthomonas oryzae pv. oryzae distribution in rice using fluorescent-labelling[J]. Journal of Plant Biology, 2010, 15: 595-599. |
| [9] | Bian H J, Zhang L Y, Chen L, Wang W Z, Ji H T, Dong H S.Real-time monitoring of translocation of selected type-III effectors from Xanthomonas oryzae pv. oryzae into rice cells[J]. Journal of Biosciences, 2019, 44: 4. |
| [10] | Li P, Zhang L Y, Mo X Y, Ji H T, Bian H J, Hu Y Q, Majid T, Long J Y, Pang H, Tao Y, Ma J B, Dong H S.Rice aquaporin PIP1;3 and harpin Hpa1 of bacterial blight pathogen cooperate in a type III effector translocation[J]. Journal of Experimental Botany, 2019, 70(12): 3057-3073. |
| [11] | Zhang L Y, Chen L, Dong H S.Plant aquaporins in infection by and immunity against pathogens: A critical review[J]. Frontiers in Plant Science, 2019, 10: 632. |
| [12] | Wang X, Zhang L Y, Ji H T, Mo X Y, Li P, Wang J Z, Dong H S.Hpa1 is a type III translocator in Xanthomonas oryzae pv. oryzae[J]. BMC Microbiology, 2018, 18: 105. |
| [13] | Zhang L Y, Hu Y Q, Li P, Wang X B, Dong H S.Silencing of an aquaporin gene diminishes bacterial blight disease in rice[J]. Australasian Plant Pathology, 2019, 48(2): 143-158. |
| [14] | Zhu P L, Zhao S, Tang J L, Feng J X.The rsmA-like gene rsmAxoo of Xanthomonas oryzae pv. oryzae regulates bacterial virulence and production of diffusible signal factor[J]. Molecular Plant Pathology, 2011, 12(3): 227-237. |
| [15] | White F F, Potnis N, Jones J B, Koebnik R.The type III effectors of Xanthomonas[J]. Molecular Plant Pathology, 2009, 10(6): 749-766. |
| [16] | Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U.Breaking the code of DNA binding specificity of TAL-type III effectors[J]. Science, 2009, 326: 1509-1512. |
| [17] | Moscou M J, Bogdanove A J.A simple cipher governs DNA recognition by TAL effectors[J]. Science, 2009, 326: 1501. |
| [18] | Oliva R, Ji C H, Atienza-Grande1 G, Huguet-Tapia J C, Perez-Quintero A, Li T, Eom J S, Li C H, Nguyen H, Liu B, Auguy F, Sciallano C, Luu V T, Dossa G S, Cunnac S, Schmidt S M, Slamet-Loedin I H, Cruz C V, Szurek B, Frommer W B, White F F, Yang B. Broad-spectrum resistance to bacterial blight in rice using genome editing[J]. Nature Biotechnology, 2019, 37: 1344-1350. |
| [19] | Richter A, Streubel J, Blücher C, Szurek B, Reschke M, Grau J, Boch J.A TAL effector repeat architecture for frameshift binding[J]. Nature Communcations, 2014, 5: 3447. |
| [20] | Kim J G, Stork W, Mudgett M B.Xanthomonas type Ⅲ effector XopD desumoylates tomato transcription factor SlERF4 to suppress ethylene responses and promote pathogen growth[J]. Cell Host Microbe, 2013, 13(2): 143-154. |
| [21] | Kim J G, Taylor K W, Hotson A, Keegan M, Schmelz E A, Mudgett M B.XopD SUMO protease affects host transcription, promotes pathogen growth, and delays symptom development in Xanthomonas-infected tomato leaves[J]. Plant Cell, 2008, 20: 1915-1929. |
| [22] | Long J, Song C, Yan F, Zhou J, Zhou H, Yang B.Non-TAL effectors from Xanthomonas oryzae pv. oryzae suppress peptidoglycan-triggered MAPK activation in rice[J]. Frontiers in Plant Science, 2018, 12(9): 1857. |
| [23] | Medina C A, Reyes P A, Trujillo C A, Gonzalez J L, Bejarano D A, Montenegro N A, Jacobs J M, Joe A, Restrepo S, Alfano J R, Bernal A.The role of type Ⅲ effectors from Xanthomonas axonopodis pv. manihotis in virulence and suppression of plant immunity[J]. Molecular Plant Pathology, 2018, 19(3): 593-606. |
| [24] | Sinha D, Gupta M K, Patel H K, Ranjan A, Sonti R V.Cell wall degrading enzyme induced rice innate immune responses are suppressed by the type 3 secretion system effectors XopN, XopQ, XopX and XopZ of Xanthomonas oryzae pv. oryzae[J/OL]. PLoS ONE, 2013, 8(9): e75867. |
| [25] | Taylor K W, Kim J G, Su X B, Aakre C D, Roden J A, Adams C M, Mudgett M B.Tomato TFT1 is required for PAMP-triggered immunity and mutations that prevent T3S effector XopN from binding to TFT1 attenuate Xanthomonas virulence[J/OL]. PLoS Pathogens, 2012, 8(6): e1002768. |
| [26] | Li N, Han X, Feng D, Yuan D, Huang L J.Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: Do we understand what they are whispering[J]? International Journal of Molecular Sciences, 2019, 20(3): 671. |
| [27] | Roden J A, Belt B, Ross J B, Tachibana T, Vargas J, Mudgett M B.A genetic screen to isolate type III effectors translocated into pepper cells during Xanthomonas infection[J]. Proceedings of the National Academy of Sciences of the United States of America, 2004, 101(47): 16 624-16 629. |
| [28] | Jiang B L, He Y Q, Cen W J, Wei H Y, Jiang G F, Jiang W, Hang X H, Feng J X, Lu G T, Tang D J, Tang J L.The type III secretion effector XopXccN of Xanthomonas campestris pv. campestris is requried for full virulence[J]. Research in Microbiology, 2008, 159(3): 216-220. |
| [29] | Liao Z X, Li J Y, Mo X Y, Ni Z, Jiang W, He Y Q, Huang S.Type III effectors xopN and avrBS2 contribute to the virulence of Xanthomonas oryzae pv. oryzicola strain GX01[J]. Research in Microbiology, 2019: 10.002. |
| [30] | Cheong H, Kim C Y, Jeon J S, Lee B M, Sun Moon J, Hwang I.Xanthomonas oryzae pv. oryzae type III effector XopN targets OsVOZ2 and a putative thiamine synthase as a virulence factor in rice[J/OL]. PLoS One, 2013, 8(9): e73346. |
| [31] | 孙荣华. 水稻白叶枯菌JXOV中tal17.5基因的克隆与功能研究[D]. 南京:南京农业大学, 2016. |
| Sun R H.Cloning and functional characterization of tal 17.5 gene from Xanthomonas oryzae pv. oryzae JXOV strain[D]. Nanjing: Nanjing Agricultural University, 2016. | |
| [32] | Jiang N, Yan J, Liang Y, Shi Y, He Z, Wu Y, Zeng Q, Liu X, Peng J.Resistance genes and their interactions with bacterial blight/leaf streak pathogens (Xanthomonas oryzae) in rice (Oryza sativa L.): An updated review[J]. Rice (NY), 2020, 13(1): 3. |
| [33] | Ji Z, Wang C, Zhao K.Rice routes of counting Xanthomonas oryzae[J]. International Journal of Molecular Sciences, 2018, 19: 3008. |
| [34] | Yang S Q, Liu S Y, Zhao S, Yu Y H, Li R B, Duan C J, Tang J L, Feng J X.Molecular and pathogenic characterization of new Xanthomonas oryzae pv. oryzae strains from the coastline region of Fangchenggang city in China[J]. World Journal of Microbiology & Biotechnology, 2013, 29(4): 713-720. |
| [35] | Yang B, Sugio A, White F F.Os8N3 is a host disease-susceptibility gene for bacterial blight of rice[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103: 10 503-10 508. |
| [36] | 胡立. 水稻白叶枯病菌效应蛋白XopN病理功能与核黄素对拟南芥开花调控的影响[D]. 南京:南京农业大学, 2016. |
| Hu L.Pathological roles of Xanthomonas oryzae pv. oryzae effector XopN and Arabidopsis flowering regulation by Riboflavin content[D]. Nanjing: Nanjing Agricultural University, 2016. | |
| [37] | Chen L, Zhang S J, Zhang S S, Qu S P, Ren X Y, Long J Y, Yin Q, Qian J, Sun F, Zhang C L, Wang L X, Wu X L, Wu T Q, Zhang Z K, Chen Z Q, Hayes M, Beer S V, Dong H S.A fragment of the Xanthomonas oryzae pv. oryzicola harpin HpaGXooc reduces disease and increases yield of rice in extensive grower plantings[J]. Phytopathology, 2008, 98(7): 792-802. |
| [38] | 时立文. SPSS19.0统计分析从入门到精通[M]. 北京: 清华大学出版社, 2012. |
| Shi L W.SPSS19.0 statistical analysis from entry to the master[M]. Beijing: Tsinghua University Press, 2012. | |
| [39] | Zhu Y, Chen H, Fan J, Wang Y, Li Y, Chen J, Fan J, Yang S, Hu L, Leung H, Mew T W, Teng P S, Wang Z, Mundt C C.Genetic diversity and disease control in rice[J]. Nature, 2000, 406(6797): 718-722. |
| [40] | Li C, Li W, Zhou Z, Vhen H, Xie C, Lin Y.A new rice breeding method: CRISPR/Cas9 system editing of the Xa13 promoter to cultivate transgene-free bacterial blight-resistant rice[J]. Plant Biotechnology Journal, 2020, 18: 313-315. |
| [41] | Yuan M, Chu Z, Li X, Xu, C, Wang S. Pathogen-induced expression loss of function is the key factor in race-specific bacterial resistance conferred by a recessive R gene xa13 in rice[J]. Plant & Cell Physiology, 2009, 50(5): 947-955. |
| [42] | Song C, Yang B.Mutagenesis of 18 type III effectors reveals virulence function of XopZPXO99 in Xanthomonas oryzae pv. oryzae[J]. Molecular Plant-Microbe Interactions, 2010, 23: 893-902. |
| [43] | Zhao S, Mo W L, Wu F, Tang W, Tang J L, Szurek B, Verdier V, Koebnik R, Feng J X.Identification of non-TAL effectors in Xanthomonas oryzae pv. oryzae Chinese strain 13751 and analysis of their role in the bacterial virulence[J]. World Journal of Microbiology & Biotechnology, 2013, 29: 733-744. |
| [44] | 董汉松, 陈蕾, 邹珅珅. 植物分子免疫学[M]. 北京: 科学出版社, 2020 (待印). |
| Dong H S, Chen L, Zhou S S.Plant molecular immunology[M]. Beijing: Science Press, 2020 | |
| (To be printed). | |
| [45] | Akimoto-Tomiyama C, Furutani A, Tsuge S, Washington E J, Nishizawa Y, Minami E, Ochiai H.XopR, a type III effector secreted by Xanthomonas oryzae pv. oryzae, suppresses microbe-associated molecular pattern triggered immunity in Arabidopsis thaliana[J]. Molecular Plant-Microbe Interactions, 2012, 25: 505-514. |
| [46] | Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen J L, Zhang Q, Wang S.Promoter mutations of an essential gene for pollen development result in disease resistance in rice[J]. Genes & Development, 2006, 20(10): 1250-1255. |
| [47] | Sobhani A, Khanlarkhani N, Baazm M, Mohammadzadeh F, Najafi A, Mehdinejadiani S, Sargolzaei Aval F.Multipotent stem cell and current application[J]. Acta Medica Iranica, 2017, 55(1): 6-23. |
| [48] | Kim J G, Li X, Roden J A, Taylor K W, Aakre C D, Su B, Lalonde S, Kirik A, Chen Y, Baranage G, McLane H, Martin G B, Mudgett M B. Xanthomonas T3S effector XopN suppresses PAMP-triggered immunity and interacts with a tomato atypical receptor-like kinase and TFT1[J]. Plant Cell, 2009, 21: 1305-1323. |
| [49] | Ji H, Dong H.Key steps in type III secretion system (T3SS) towards translocon assembly with potential sensor at plant plasma membrane[J]. Molecular Plant Pathology, 2015, 16(7): 762-773. |
| [50] | Kvitko B H, Ramos A R, Morello J E, Oh H S, Collmer A.Identification of harpins in Pseudomonas syringae pv. tomato DC3000, which are functionally similar to HrpK1 in promoting translocation of type III secretion system effectors[J]. Journal of Bacteriology, 2007, 189: 8059-8072. |
| [51] | Römer P, Recht S, Strauß T, Elsaesser J, Schornack S, Boch J, Wang S, Lahaye T.Promoter elements of rice susceptibility genes are bound and activated by specific TAL effectors from the bacterial blight pathogen, Xanthomonas oryzae pv. oryzae[J]. New Phytology, 2010, 187: 1048-1057. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||