
中国水稻科学 ›› 2020, Vol. 34 ›› Issue (5): 383-396.DOI: 10.16819/j.1001-7216.2020.0302
• 研究报告 • 下一篇
唐小涵1, 刘世家1, 刘喜1, 田云录1, 王云龙1, 滕烜1, 段二超1, 张元燕1, 江玲1, 张文伟1, 王益华1,*( ), 万建民1,2
), 万建民1,2
收稿日期:2020-03-01
修回日期:2020-03-18
出版日期:2020-09-10
发布日期:2020-09-10
通讯作者:
王益华
基金资助:
Xiaohan TANG1, Shijia LIU1, Xi LIU1, Yunlu TIAN1, Yunlong WANG1, Xuan TENG1, Erchao DUAN1, Yuanyan ZHANG1, Ling JIANG1, Wenwei ZHANG1, Yihua WANG1,*( ), Jianmin WAN1,2
), Jianmin WAN1,2
Received:2020-03-01
Revised:2020-03-18
Online:2020-09-10
Published:2020-09-10
Contact:
Yihua WANG
摘要:
【目的】氨酰-tRNA合成酶(aminoacyl-tRNA synthetases, aaRSs)与遗传信息传递密切相关,已发现植物中aaRSs家族蛋白在维持翻译功能之余,还参与配子发生与胚发育、质体的早期发育以及免疫信号的感知与病害防御等生物学过程。本研究利用水稻胚乳发育缺陷突变体,分析水稻色氨酰-tRNA合成酶(WRS1)在胚乳发育中的作用,证明WRS1基因编码一个影响水稻胚乳发育的关键因子。【方法】本研究通过甲烷磺酸乙酯(ethyl methane sulfonate, EMS)诱变籼稻(Oryza sativa subsp. indica)品种N22,筛选到一个稳定遗传的水稻粉质胚乳突变体(wrs1),图位克隆获得目标基因。对wrs1成熟种子进行形态学观察以及淀粉相关理化性质测定,利用细胞学切片分析wrs1发育中胚乳的结构,利用实时荧光定量PCR(quantitative real-time PCR, qRT-PCR)和GUS活性染色分析基因表达模式,通过qRT-PCR比较野生型与突变体花后12 d胚乳中淀粉合成相关基因表达情况,免疫印迹检测野生型与突变体成熟种子中淀粉合成酶蛋白积累情况,使用全自动氨基酸分析仪测定游离氨基酸含量。【结果】 wrs1突变体幼苗表现出明显的发育滞后且逐渐蔫萎死亡,从杂合突变体(WRS1wrs1)中分离到的粉质籽粒呈现明显的腹部皱缩,粒厚、千粒重下降,同时总淀粉含量下降,糊化淀粉的峰值黏度和崩解值均低于野生型。wrs1突变体发育胚乳中复合淀粉颗粒变小,排列疏松。WRS1定位于第12染色体长臂约183 kb的区间内,测序发现编码色氨酰-tRNA合成酶(tryptophanyl-tRNA synthetase, WRS)基因的第6外显子上发生单碱基替换,导致一个保守位置上的甲硫氨酸被替换。wrs1突变体中大部分淀粉合成相关基因表达量下调,且野生型与突变体间基因表达的变化与相应蛋白积累的差异存在不一致的趋势。wrs1突变体籽粒中蛋白质积累降低,而游离氨基酸含量显著升高。【结论】 WRS1编码色氨酰-tRNA合成酶,该基因突变后通过影响氨基酸稳态和蛋白质合成,造成淀粉合成相关基因异常表达从而影响淀粉的合成与积累,导致种子发育缺陷。
中图分类号:
唐小涵, 刘世家, 刘喜, 田云录, 王云龙, 滕烜, 段二超, 张元燕, 江玲, 张文伟, 王益华, 万建民. 色氨酰-tRNA合成酶基因WRS1调控水稻种子发育[J]. 中国水稻科学, 2020, 34(5): 383-396.
Xiaohan TANG, Shijia LIU, Xi LIU, Yunlu TIAN, Yunlong WANG, Xuan TENG, Erchao DUAN, Yuanyan ZHANG, Ling JIANG, Wenwei ZHANG, Yihua WANG, Jianmin WAN. Tryptophanyl-tRNA Synthetase Gene WRS1 Regulates Rice Seed Development[J]. Chinese Journal OF Rice Science, 2020, 34(5): 383-396.
| 引物名称 Primer name | 正向引物序列 Forward (5′-3′) | 反向引物序列 Reserve (5′-3′) | 产物大小 Product size / bp |
|---|---|---|---|
| N4 | TCCCTGATTCCATTTCTC | TTATGCCTCCATTGTTGA | 404 |
| N10 | GGCCATCTCATCCCCTTCAT | AATTGAGTCCCGGCCACC | 633 |
| Hyg | CTATTTCTTTGCCCTCGGAC | ATGCCTGAACTCACCGCGAC | 1083 |
表1 wrs1转基因家系鉴定所用引物
Table 1 Markers used for identification in wrs1 transgenic lines.
| 引物名称 Primer name | 正向引物序列 Forward (5′-3′) | 反向引物序列 Reserve (5′-3′) | 产物大小 Product size / bp |
|---|---|---|---|
| N4 | TCCCTGATTCCATTTCTC | TTATGCCTCCATTGTTGA | 404 |
| N10 | GGCCATCTCATCCCCTTCAT | AATTGAGTCCCGGCCACC | 633 |
| Hyg | CTATTTCTTTGCCCTCGGAC | ATGCCTGAACTCACCGCGAC | 1083 |
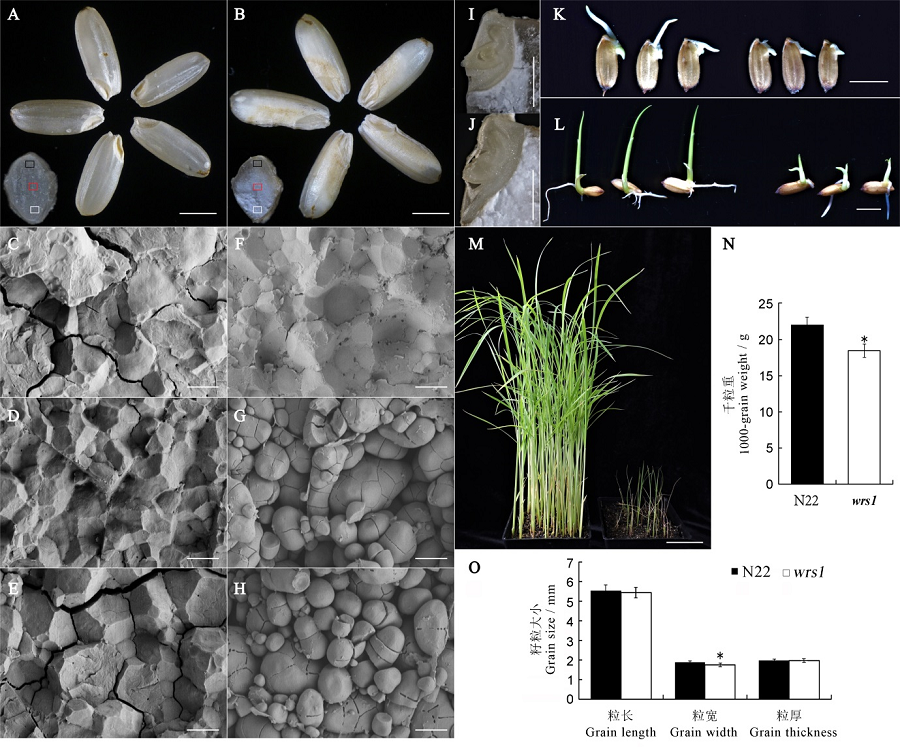
图1 野生型与wrs1突变体表型分析 A, B—野生型(A)和wrs1突变体(B)成熟种子外观比较,标尺为2 mm;C~H—野生型(C, D, E)和wrs1突变体(F, G, H)成熟种子横断面的扫描电镜观察(局部),由上到下依次对应野生型和wrs1突变体中的黑框、红框、白框,标尺为5 μm;I, J—野生型(I)和wrs1(J)突变体种子吸胀后的胚(30℃下吸胀10 h),标尺为1 mm;K~M—野生型和突变体不同时期的发育情况,分别为30℃催芽24 h(K)、发芽后3 d(L),发芽后20 d(M)的种子/幼苗发育状况,K, L标尺为5 mm,M标尺为5 cm;N—千粒重比较,n = 3;O—籽粒大小比较,n = 20。所有数值为平均值±标准差,t检验,*表示在P<0.05水平上差异显著。
Fig. 1. Phenotypic analysis of wild-type and the wrs1 mutant. A, B, Comparison of mature seeds of wild-type (A) and the wrs1 mutant (B). Bars = 2 mm. C~H, Scanning electron microscopic (SEM) analysis of cross-sections of wild-type (C-E) and wrs1 mutant (F-H) mature seeds. C-E and F-H correspond to the black frame, red frame, and white frames in wild-type and wrs1, respectively. Bars = 5 μm. I, J, Embryos of wild-type (I) and wrs1 mutant (J) seeds after imbibition (30℃, 10 h). Bars = 1 mm. K~M, Developmental status of wild-type and the wrs1 mutant. 24 hours after seed germination (K), 3 days after germination (L), and 20 days after germination (M). Bars = 5 mm in K and L, 5 cm in M. N, 1000-grain weight, n = 3. O, Grain length, width and thickness, n = 20. All values are means ± SD. *indicates significant difference at P<0.05 level(Student’s t-test).
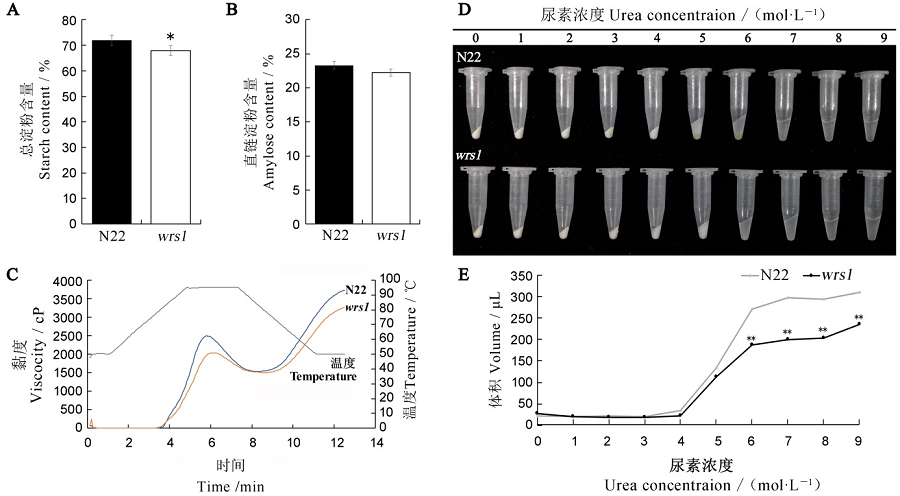
图2 野生型和wrs1突变体种子淀粉含量和理化特性分析 A, B—N22和wrs1突变体胚乳中总淀粉(A)与直链淀粉(B)含量,n = 3;C—野生型和wrs1突变体黏度特性比较;D—不同尿素浓度下N22和wrs1突变体米粉的膨胀情况;E—野生型和wrs1突变体米粉膨胀体积比较,n = 3。所有数值取平均值±标准差,t检验,*和**分别表示在0.05和0.01水平上差异显著。
Fig. 2. Content and physicochemical characteristics of starches in mature seeds of wild-type and the wrs1 mutant. A, B, Contents of total starch (A) and amylose (B) in wild-type and the wrs1 mutant. n = 3. C, Analysis of RVA characteristic values of starch in wild-type and the wrs1 mutant. D, Swollen volume of the wild-type and wrs1 starch in urea solution. E, Comparation of the swollen volume in the wild-type and wrs1. n = 3. Values are means ± SD. * and **indicate significant difference at 0.05 and 0.01 level, respectively(Student’s t-test).
| 材料 Material | 峰值黏度 Peak viscosity / cP | 热浆黏度 Hot paste viscosity / cP | 崩解值 Breakdown viscosity / cP | 冷胶黏度 Cool paste viscosity / cP | 消减值 Setback viscosity / cP | 糊化温度 Pasting temperature /℃ | 峰值时间 Peak time / min |
|---|---|---|---|---|---|---|---|
| N22 | 2494 | 1536 | 958 | 3727 | 1233 | 81.40 | 5.80 |
| wrs1 | 2041 | 1506 | 535 | 3262 | 1221 | 80.65 | 6.13 |
表2 野生型wrs1突变体淀粉RVA谱特征分析
Table 2 Analysis of RVA characteristic values of starch in wild-type and the wrs1 mutant.
| 材料 Material | 峰值黏度 Peak viscosity / cP | 热浆黏度 Hot paste viscosity / cP | 崩解值 Breakdown viscosity / cP | 冷胶黏度 Cool paste viscosity / cP | 消减值 Setback viscosity / cP | 糊化温度 Pasting temperature /℃ | 峰值时间 Peak time / min |
|---|---|---|---|---|---|---|---|
| N22 | 2494 | 1536 | 958 | 3727 | 1233 | 81.40 | 5.80 |
| wrs1 | 2041 | 1506 | 535 | 3262 | 1221 | 80.65 | 6.13 |
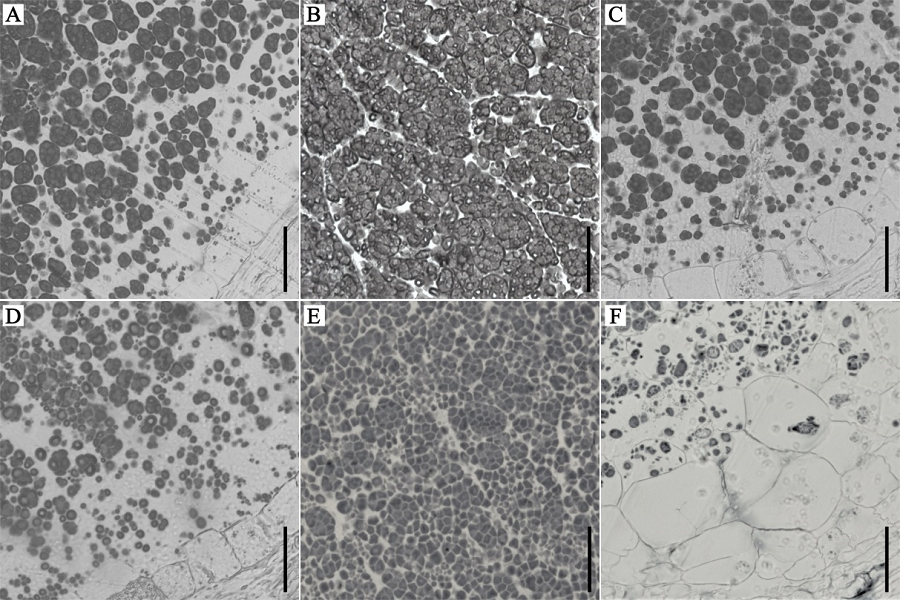
图3 野生型和wrs1突变体发育中的胚乳半薄切片观察 A, D—半薄切片分析野生型(A)与wrs1突变体(D)开花后第8天背部胚乳中的复合淀粉颗粒结构;B, E—半薄切片分析野生型(B)和wrs1突变体(E)开花后第8天中间部位胚乳中的复合淀粉颗粒结构;C, F—半薄切片分析野生型(C)和wrs1突变体(F)开花后第8天腹部胚乳中的复合淀粉颗粒结构;标尺为50 μm。
Fig. 3. Semi-thin section analysis of developing endosperm of wild-type and the wrs1 mutant. A, D—Semi-thin section analysis of the compound starch granules of back area of wild-type (A) and wrs1 (D) mutant endosperm at 8 DAF. B, E—Semi-thin section analysis of compound starch granules of central part of wild-type (B) and wrs1 mutant (E) endosperm at 8 DAF. C, F—Semi-thin section analysis of compound starch granules of the ventral side of wild-type (C) and wrs1 mutant (F) endosperm at 8 DAF. Bars = 50 μm.
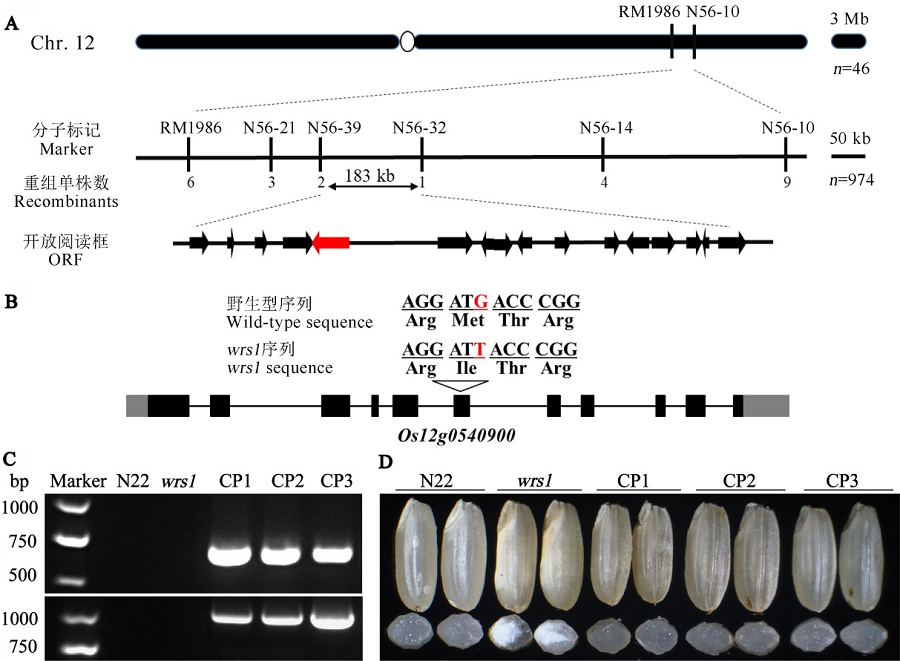
图4 WRS1的图位克隆 A—WRS1位点的精细定位,WRS1定位在第12染色体长臂标记N56-39与N56-32之间约183 kb区域内,包含16个预测基因;B—Os12g0540900基因结构和突变位点,序列中存在1处单碱基替换(红色),导致蛋白质中第256位氨基酸由Met变成Ile;C—转基因家系CP1、CP2和CP3的PCR鉴定结果,上下图分别为转基因阳性鉴定引物N10与Hyg的扩增情况;D—N22、wrs1突变体以及突变体背景的转基因阳性家系(CP1、CP2和CP3)的籽粒表型,上侧为各转基因阳性家系成熟种子外观,下侧为对应家系的种子横断面。
Fig. 4. Map-based cloning of WRS1. A, Fine mapping of the WRS1 locus. The WRS1 locus was mapped to a 183 kb region with markers N56-39 and N56-32 on the long arm of chromosome 12, which contains 16 predicted open reading frames (ORFs). B, The structure of Os12g0540900 and the mutation site. One nucleotide substitution (red) in the sequence is indicated, which results in a substitution of Met-256 to Ile. C, PCR analyses of wrs1 complemented lines (CP1, CP2 and CP3). The upper and lower panels in turn show the amplification of the transgenic positive identification primers N10 and Hyg. D, Mature seed appearance of wild-type, the wrs1 mutant and three positive transgenic lines (CP1, CP2 and CP3). The upside shows the mature seed phenotype of each positive transgenic line, and the downside shows cross-section of the seeds in corresponding transgenic line.
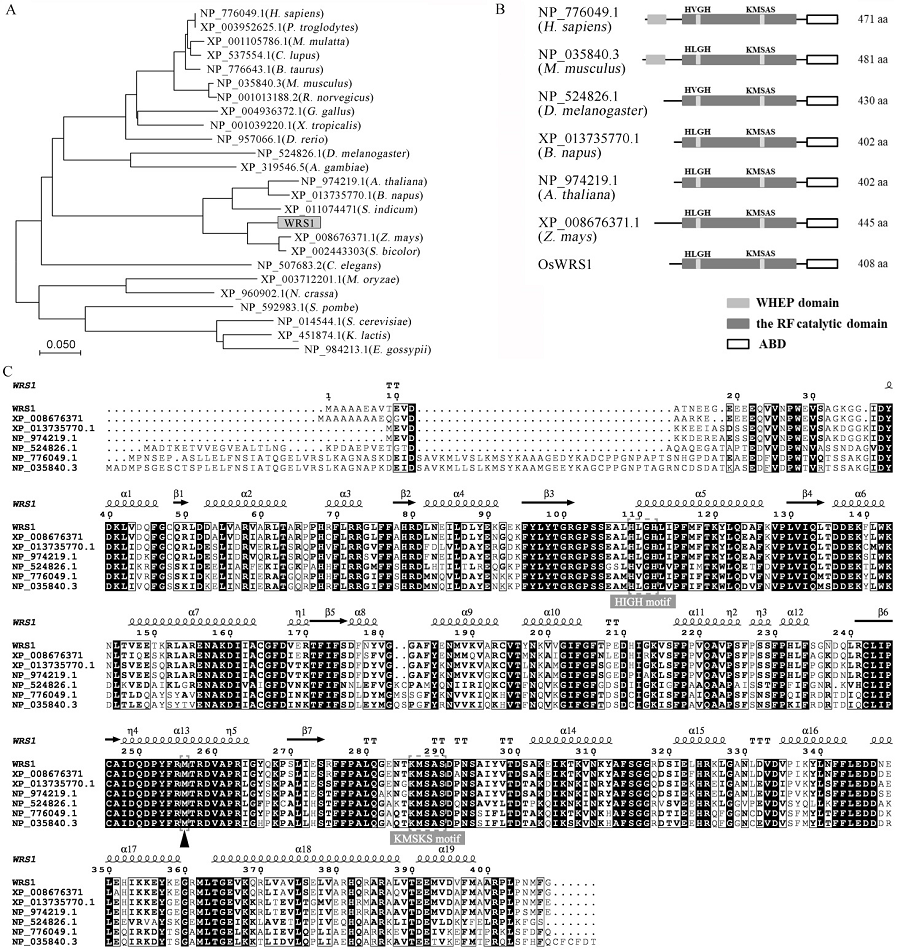
图5 WRS1与其同源蛋白结构及进化树分析 A—WRS1及其同源蛋白的进化树分析。进化树用MEGA7软件构建,采用邻接法,步长检验1000次重复。蛋白名字采用相应的NCBI编号。B—WRS1及其同源蛋白结构。灰色方框代表在动物中保守的WHEP结构域,黑色方框代表罗斯曼折叠催化核心结构域,白色方框代表反密码子结合结构域,aa代表氨基酸。C—WRS1及其同源蛋白序列比对,两个灰色方框区域代表Ⅰ类氨酰-tRNA合成酶特有的保守基序,箭头位置代表突变位点;二级结构从PHYRE2网站获得,蛋白预测基于同源建模法(normal mode),以已知的酵母色氨酰-tRNA合成酶晶体模型c3kt3D为模板;氨基酸序列采用ESPript3.0网站进行比对(默认参数)[37]。
Fig. 5. Structure and phylogenetic analysis of WRS1 protein and its homologs. A, A neighbor-joining tree of WRS1 and its homologs. The tree was constructed using MEGA7 and bootstrapped with 1000 replicates. The proteins are named after their NCBI accession numbers. B, Domain analysis of WRS1 and its homologous proteins. The gray boxes represent the WHEP domain that is conserved in animals, the black boxes represent the Rossmann fold catalytic domain, the white boxes represent the anti-codon binding domain, aa represents the amino acid. C, Protein sequence alignment of WRS1 and its homologs. Two gray boxes indicate the catalytic site of Class I aaRSs; The arrowhead indicates the mutation site. The secondary structure is obtained from the PHYRE2 website by homology modeling (normal mode) with a known yeast tryptophanyl-tRNA synthetase crystal model c3kt3D as the pattern; the amino acid sequences are compared by the ESPript 3.0 website (default).
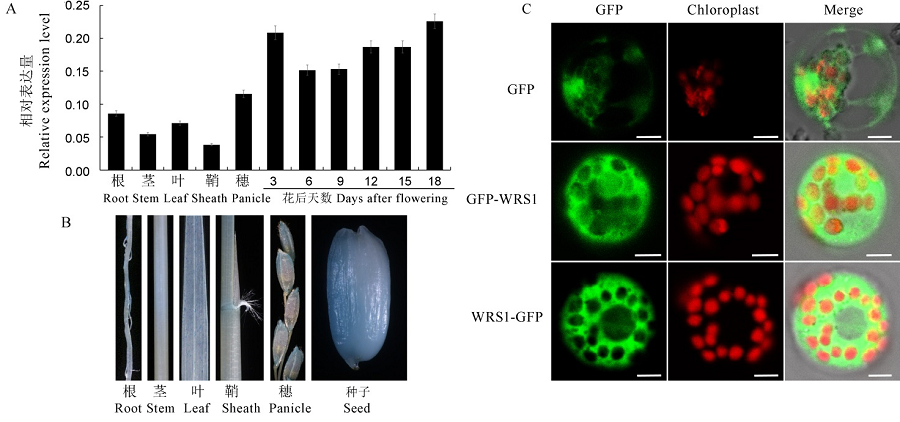
图6 WRS1的表达模式和亚细胞定位 A—WRS1在野生型的根、茎、叶、鞘、穗和不同发育天数的胚乳中的相对表达量;B—WRS1启动子驱动GUS基因表达的植株中根、茎、叶、鞘、幼穗和发育种子的GUS染色;C—35S启动子驱动下GFP-WRS1和WRS1-GFP融合表达载体在水稻原生质体中GFP荧光表达,对照为pAN580-GFP空载体,标尺为5 μm。
Fig. 6. Expression pattern and subcellular localization of WRS1. A, Relative expression levels of WRS1 in root, stem, leaf, sheath, panicle and developing seeds of wild-type. B, GUS staining patterns in root, stem, leaf, sheath, panicle and seed in transgenic plants. C, Fluorescence micrographs of the WRS1-GFP and GFP-WRS1 fusion constructs in rice protoplasts, with the empty pAN580-GFP served as a control. Bars, 5 μm.
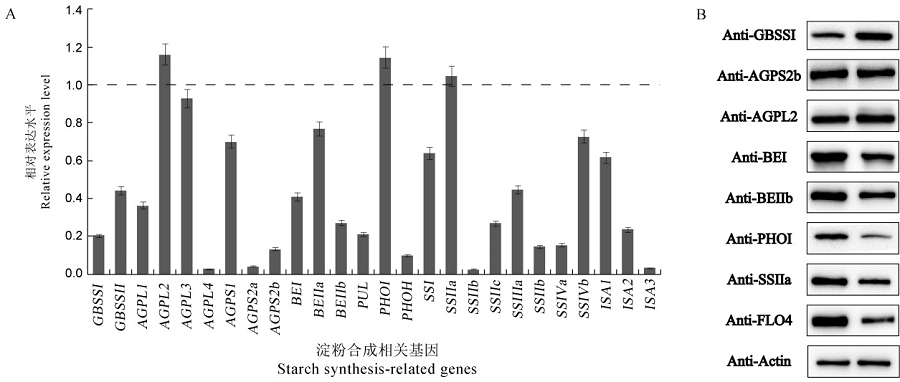
图7 淀粉合成相关基因的表达及淀粉合成酶蛋白的积累 A—淀粉合成相关基因表达分析,结果为wrs1突变体中的基因以N22中对应基因为基准的相对表达水平,提取花后12 d种子的总RNA,用Ubiquitin作为内参基因。所有数值为平均值±标准差(n = 3),t检验,*P<0.05,**P<0.01。B—Western blotting检测野生型和突变体成熟种子中的淀粉合成相关酶蛋白含量。GBSSⅠ—颗粒结合淀粉合酶Ⅰ;AGPS2b―腺苷葡萄糖焦磷酸化酶小亚基2b;AGPL2―腺苷葡萄糖焦磷酸化酶大亚基2;BEⅠ―淀粉分支酶Ⅰ;BEⅡb―淀粉分支酶Ⅱb;PHOⅠ―质体磷酸化酶Ⅰ;SSⅡa―淀粉合酶Ⅱa;FLO4—胞质丙酮酸磷酸双激酶B,用Actin作为上样内参。
Fig. 7. Expression and immunoblotting analyses of starch synthesis-related genes. A, Expression analysis of starch synthesis-related genes. The relative expression levels of the genes in the wrs1 mutant were based on the corresponding genes in N22. Total RNA was extracted from seeds at 12 days after flowering. Ubiquitin was used as control. n = 3. All values are means ± SD. Student’s t-test, *P<0.05, **P<0.01. B, Western blotting analysis of starch synthesis related proteins in wild-type and mutant mature seeds. GBSSⅠ, Granule bound starch synthaseⅠ; AGPS2b, ADP-glucose pyrophosphorylase small subunit 2b; AGPL2, ADP-glucose pyrophosphorylase large subunit 2; BEⅠ, Starch-branching enzyme; BEⅡb, Starch-branching enzymeⅡb; PHOⅠ, Plastid phosphorylaseⅠ; SSⅡa, Starch synthase Ⅱa; FLO4(FLOURY ENDOSPERM4), cytosolic pyruvate orthophosphate dikinase B; Actin was used as control.

图8 wrs1突变体的蛋白含量与游离氨基酸含量 A—野生型N22和wrs1突变体籽粒的蛋白含量;B—野生型N22与wrs1突变体籽粒中游离氨基酸含量;C—野生型与wrs1籽粒中游离氨基酸总量;D—野生型与wrs1突变体米粉中游离氨基酸的相对含量。所有数值为平均值±标准差(n = 3),t检验,*P<0.05,**P<0.01。
Fig. 8. Contents of protein and free amino acids in wrs1. A, Protein contents in wild-type and wrs1 grains. B, Free amino acid contents in the seeds of wild-type and the wrs1 mutant. C, Total free amino acids contents in the seeds of wild-type and the wrs1 mutant; D, Relative contents of free amino acids in mature seeds of wild-type and the wrs1 mutant. n = 3. All values are means ± SD. Student’s t-test, *P<0.05, **P<0.01.
| [1] | Fitzgerald M A, McCouch S R, Hall R D. Not just a grain of rice: The quest for quality[J]. Trends in Plant Science, 2009, 14(3): 133-139. |
| [2] | Crofts N, Nakamura Y, Fujita N.Critical and speculative review of the roles of multi-protein complexes in starch biosynthesis in cereals[J]. Plant Science, 2017, 262: 1-8. |
| [3] | Feike D, Seung D, Graf A, Bischof S, Ellick T, Coiro M, Soyk S, Eicke S, Mettler-Altmann T, Lu K J, Trick M, Zeeman S C, Smith A M.The starch granule-associated protein EARLY STARVATION1 is required for the control of starch degradation in Arabidopsis thaliana leaves[J]. Plant Cell, 2016, 28(6): 1472-1489. |
| [4] | Silver D M, Kotting O, Moorhead G B.Phosphoglucan phosphatase function sheds light on starch degradation[J]. Trends in Plant Science, 2014, 19(7): 471-478. |
| [5] | Hejazi M, Mahlow S, Fettke J.The glucan phosphorylation mediated by α-glucan, water dikinase (GWD) is also essential in the light phase for a functional transitory starch turn-over[J/OL].Plant Signaling & Behavior, 2014, 9(7): e28892. |
| [6] | Hwang S K, Nishi A, Satoh H, Okita T W.Rice endosperm-specific plastidial alpha-glucan phosphorylase is important for synthesis of short-chain malto- oligosaccharides[J]. Archives of Biochemistry and Biophysics, 2010, 495(1): 82-92. |
| [7] | Higgins J E, Kosar-Hashemi B, Li Z, Howitt C A, Larroque O, Flanagan B, Morell M K, Rahman S.Characterization of starch phosphorylases in barley grains[J]. Journal of the Science of Food and Agriculture, 2013, 93(9): 2137-2145. |
| [8] | Peng C, Wang Y, Liu F, Ren Y, Zhou K, Lv J, Zheng M, Zhao S, Zhang L, Wang C, Jiang L, Zhang X, Guo X, Bao Y, Wan J.FLOURY ENDOSPERM6 encodes a CBM48 domain-containing protein involved in compound granule formation and starch synthesis in rice endosperm[J]. Plant Journal, 2014, 77(6): 917-930. |
| [9] | Seung D, Soyk S, Coiro M, Maier B A, Eicke S, Zeeman S C.PROTEIN TARGETING TO STARCH is required for localizing GRANULE-BOUND STARCH SYNTHASE to starch granules and for normal amylose synthesis in Arabidopsis[J/OL].PLoS Biology, 2015, 13(2): e1002080. |
| [10] | Pfister B, Zeeman S C.Formation of starch in plant cells[J]. Cellular and Molecular Life Sciences, 2016, 73(14): 2781-2807. |
| [11] | Hunter B G, Beatty M K, Singletary G W, Hamaker B R, Dilkes B P, Larkins B A, Jung R.Maize opaque endosperm mutations create extensive changes in patterns of gene expression[J]. Plant Cell, 2002, 14(10): 2591-2612. |
| [12] | Wakasa Y, Yasuda H, Oono Y, Kawakatsu T, Hirose S, Takahashi H, Hayashi S, Yang L, Takaiwa F.Expression of ER quality control-related genes in response to changes in BiP1 levels in developing rice endosperm[J]. Plant Journal, 2011, 65(5): 675-689. |
| [13] | Han X, Wang Y, Liu X, Jiang L, Ren Y, Liu F, Peng C, Li J, Jin X, Wu F, Wang J, Guo X, Zhang X, Cheng Z, Wan J.The failure to express a protein disulphide isomerase-like protein results in a floury endosperm and an endoplasmic reticulum stress response in rice[J]. Journal of Experiment Botany, 2012, 63(1): 121-130. |
| [14] | Wang Y, Ren Y, Liu X, Jiang L, Chen L, Han X, Jin M, Liu S, Liu F, Lv J, Zhou K, Su N, Bao Y, Wan J.OsRab5a regulates endomembrane organization and storage protein trafficking in rice endosperm cells[J]. Plant Journal, 2010, 64(5): 812-824. |
| [15] | Liu F, Ren Y, Wang Y, Peng C, Zhou K, Lv J, Guo X, Zhang X, Zhong M, Zhao S, Jiang L, Wang H, Bao Y, Wan J.OsVPS9A functions cooperatively with OsRAB5A to regulate post-Golgi dense vesicle-mediated storage protein trafficking to the protein storage vacuole in rice endosperm cells[J]. Molecular Plant, 2013, 6(6): 1918-1932. |
| [16] | Ren Y, Wang Y, Liu F, Zhou K, Ding Y, Zhou F, Wang Y, Liu K, Gan L, Ma W, Han X, Zhang X, Guo X, Wu F, Cheng Z, Wang J, Lei C, Lin Q, Jiang L, Wu C, Bao Y, Wang H, Wan J.GLUTELIN PRECURSOR ACCUMULATION3 encodes a regulator of post-Golgi vesicular traffic essential for vacuolar protein sorting in rice endosperm[J]. Plant Cell, 2014, 26(1): 410-425. |
| [17] | Wang Y, Liu F, Ren Y, Wang Y, Liu X, Long W, Wang D, Zhu J, Zhu X, Jing R, Wu M, Hao Y, Jiang L, Wang C, Wang H, Bao Y, Wan J.GOLGI TRANSPORT 1B regulates protein export from the endoplasmic reticulum in rice endosperm cells[J]. Plant Cell, 2016, 28(11): 2850-2865. |
| [18] | Zhu J, Ren Y, Wang Y, Liu F, Teng X, Zhang Y, Duan E, Wu M, Zhong M, Hao Y, Zhu X, Lei J, Wang Y, Yu Y, Pan T, Bao Y, Wang Y, Wan J.OsNHX5-mediated pH homeostasis is required for post-Golgi trafficking of seed storage proteins in rice endosperm cells[J/OL].BMC Plant Biology, 2019, 19(1): 295. |
| [19] | Perona J J, Gruic-Sovulj I.Synthetic and Editing Mechanisms of Aminoacyl-tRNA Synthetases[M]//Kim S. Topics in Current Chemistry-Series, 2014: 1-41. |
| [20] | Guo M, Yang X L, Schimmel P.New functions of aminoacyl-tRNA synthetases beyond translation[J]. Nature Reviews Molecular Cell Biology, 2010, 11(9): 668-674. |
| [21] | Park S G, Ewalt K L, Kim S.Functional expansion of aminoacyl-tRNA synthetases and their interacting factors: New perspectives on housekeepers[J]. Trends in Biochemical Sciences, 2005, 30(10): 569-574. |
| [22] | Duchêne A, Giritch A, Hoffmann B, Cognat V, Lancelin D, Peeters N M, Zaepfel M, Maréchal-Drouard L, Small I D.Dual targeting is the rule for organellar aminoacyl-tRNA synthetases in Arabidopsis thaliana[J]. Proceedings of the National Academy of Sciences of the United States of America, 2005, 102(45): 16484-16489. |
| [23] | Steiner R E, Ibba M. Regulation of tRNA-dependent translational quality control[J/OL]. IUBMB Life, 2019, 69: DOI: 10.1002/iub.2080. |
| [24] | Jin M. Unique roles of tryptophanyl-tRNA synthetase in immune control and its therapeutic implications[J].Experimental & Molecular Medicine, 2019, 51(1): . |
| [25] | Yakobov N, Debard S, Fischer F, Senger B, Becker H D.Cytosolic aminoacyl-tRNA synthetases: Unanticipated relocations for unexpected functions[J]. Biochimica et Biophysica Acta, 2018, 1861(4): 387-400. |
| [26] | Ray P, Arif A, Fox P.Macromolecular complexes as depots for releasable regulatory proteins[J]. Trends in Biochemical Sciences, 2007, 32(4): 158-164. |
| [27] | Berg M, Rogers R, Muralla R, Meinke D.Requirement of aminoacyl-tRNA synthetases for gametogenesis and embryo development in Arabidopsis[J]. Plant Journal, 2005, 44(5): 866-878. |
| [28] | Yang X, Li G, Tian Y, Song Y, Liang W, Zhang D.A rice glutamyl-tRNA synthetase modulates early anther cell division and patterning[J]. Plant Physiology, 2018, 177(2): 728-744. |
| [29] | Kagi C, Baumann N, Nielsen N, Stierhof Y D, Gross-Hardt R.The gametic central cell of Arabidopsis determines the lifespan of adjacent accessory cells[J]. Proceedings of the National Academy of Sciences of the United States of America, 2010, 107(51): 22 350-22 355. |
| [30] | Kim Y K, Lee J Y, Cho H S, Lee S S, Ha H J, Kim S, Choi D, Pai H S.Inactivation of organellar glutamyl- and seryl-tRNA synthetases leads to developmental arrest of chloroplasts and mitochondria in higher plants[J]. Journal of Biological Chemistry, 2005, 280(44): 37098-37106. |
| [31] | Wang Y, Wang C, Zheng M, Lyu J, Xu Y, Li X, Niu M, Long W, Wang D, Wang H, Terzaghi W, Wang Y, Wan J.WHITE PANICLE1, a val-tRNA synthetase regulating chloroplast ribosome biogenesis in rice, is essential for early chloroplast development[J]. Plant Physiology, 2016, 170(4): 2110-2123. |
| [32] | Zhang Y, Hao Y, Wang Y, Wang C, Wang Y, Long W, Wang D, Liu X, Jiang L, Wan J.Lethal albinic seedling, encoding a threonyl-tRNA synthetase, is involved in development of plastid protein synthesis system in rice[J]. Plant Cell Reports, 2017, 36(7): 1053-1064. |
| [33] | Kitagawa M, Balkunde R, Bui H, Jackson D.An aminoacyl tRNA Synthetase, OKI1, is required for proper shoot meristem size in Arabidopsis[J]. Plant Cell Physiology, 2019, 60(11): 2597-2608. |
| [34] | Zuo D, Yi S, Liu R, Qu B, Huang T, He W, Li C, Li H, Liao Y.A deoxynivalenol-activated methionyl-tRNA synthetase gene from wheat encodes a nuclear localized protein and protects plants against fusarium pathogens and mycotoxins[J]. Phytopathology, 2016, 106(6): 614-623. |
| [35] | Luna E, van Hulten M, Zhang Y, Berkowitz O, López A, Pétriacq P, Sellwood M A, Chen B, Burrell M, van de Meene A, Pieterse C M J, Flors V, Ton J. Plant perception of β-aminobutyric acid is mediated by an aspartyl-tRNA synthetase[J]. Nature Chemical Biology, 2014, 10(6): 450-456. |
| [36] | Long W, Wang Y, Zhu S, Jing W, Wang Y, Ren Y, Tian Y, Liu S, Liu X, Chen L, Wang D, Zhong M, Zhang Y, Hu T, Zhu J, Hao Y, Zhu X, Zhang W, Wang C, Zhang W, Wan J.FLOURY SHRUNKEN ENDOSPERM1 connects phospholipid metabolism and amyloplast development in rice[J]. Plant Physiology, 2018, 177(2): 698-712. |
| [37] | Robert X, Gouet P.Deciphering key features in protein structures with the new ENDscript server[J]. Nucleic Acids Research, 2014, 42: W320-W324. |
| [38] | Ries F, Herkt C, Willmund F.Co-translational protein folding and sorting in chloroplasts[J]. Plants (Basel), 2020, 9(2): 214. doi: 10.3390/plants9020214. |
| [39] | Zheng M, Wang Y, Liu X, Sun J, Wang Y, Xu Y, Lv J, Long W, Zhu X, Guo X, Jiang L, Wang C, Wan J.The RICE MINUTE-LIKE1 (RML1) gene, encoding a ribosomal large subunit protein L3B, regulates leaf morphology and plant architecture in rice[J]. Journal of Experimental Botany, 2016, 67(11): 3457-3469. |
| [40] | Wang S, Lei C, Wang J, Ma J, Tang S, Wang C, Zhao K, Tian P, Zhang H, Qi C, Cheng Z, Zhang X, Guo X, Liu L, Wu C, Wan J.SPL33, encoding an eEF1A-like protein, negatively regulates cell death and defense responses in rice[J]. Journal of Experimental Botany, 2017, 68(5): 899-913. |
| [41] | Jia J, Arif A, Ray P S, Fox P L.WHEP domains direct noncanonical function of glutamyl-prolyl tRNA synthetase in translational control of gene expression[J]. Molecular Cell, 2008, 29(6): 679-690. |
| [42] | Sajish M, Zhou Q, Kishi S, Valdez D J, Kapoor M, Guo M, Lee S, Kim S, Yang X L, Schimmel P.Trp-tRNA synthetase bridges DNA-PKcs to PARP-1 to link IFN-gamma and p53 signaling[J]. Nature Chemical Biology, 2012, 8(6): 547-554. |
| [1] | 杜溢墨 潘天 田云录 刘世家 刘喜 江玲 张文伟 王益华* 万建民. 水稻粉质皱缩胚乳突变体fse4的表型分析与基因克隆[J]. 中国水稻科学, 2019, 33(6): 499-512. |
| [2] | 方鹏飞,李三峰,焦桂爱,谢黎虹,胡培松,魏祥进*,唐绍清*. 水稻粉质胚乳突变体flo7的理化性质及基因定位[J]. 中国水稻科学, 2014, 28(5): 447-457. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||