全基因组关联分析(GWAS)鉴定水稻氮素利用效率候选基因
Identification of Candidate Genes for Rice Nitrogen Use Efficiency by Genome-wide Association Analysis
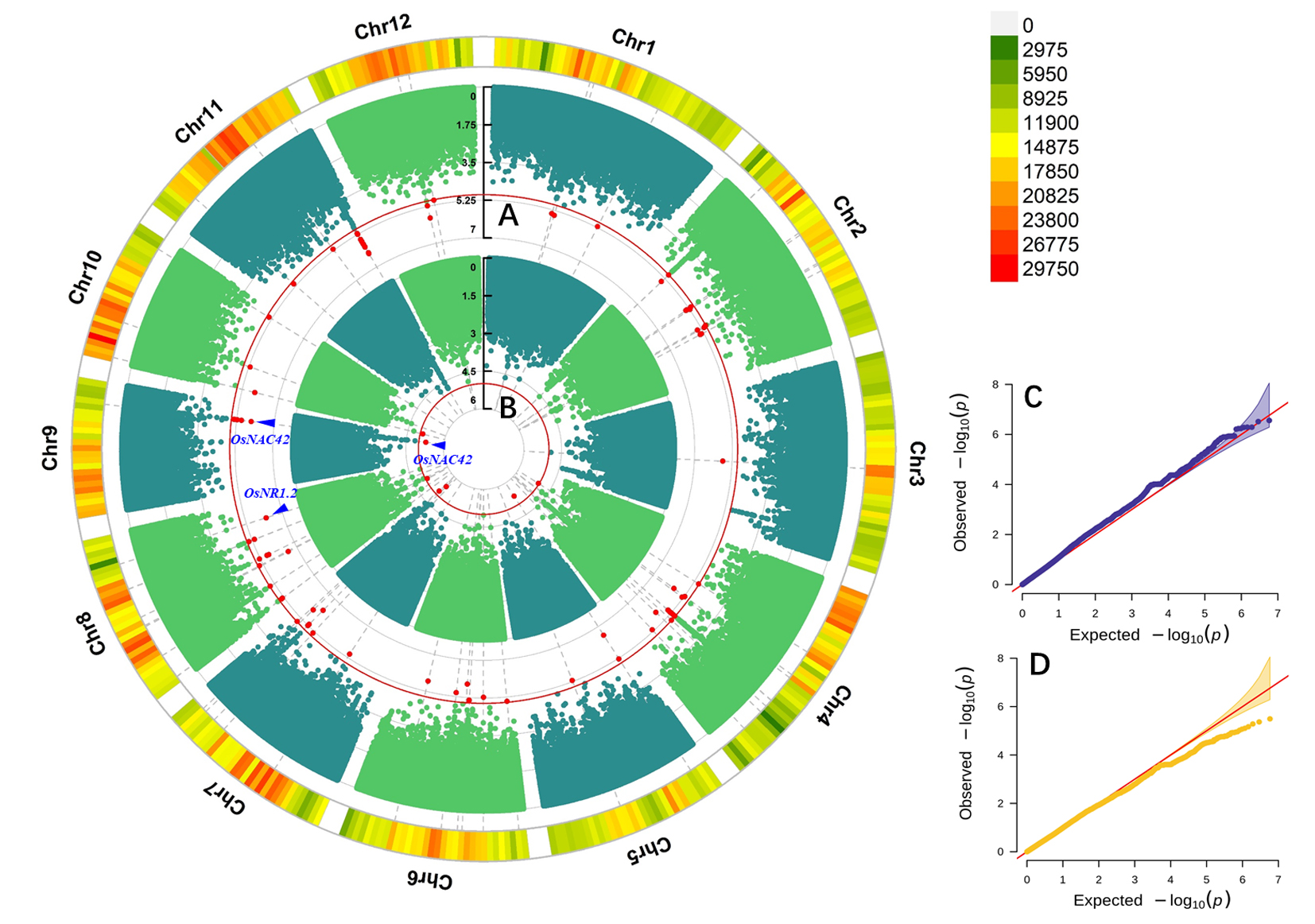
A和C分别为使用FarmCPU模型进行GWAS的曼哈顿图和QQ图;B和D分别为使用MLM模型进行GWAS的曼哈顿图和QQ图;曼哈顿图的图例为SNP密度。
A and C are Manhattan plot and QQ plot of GWAS using FarmCPU model respectively; B and D are Manhattan plot and QQ plot of GWAS using MLM model respectively; The legend of Manhattan plot is SNP density.